Abstract
CD8+ T cell immunosurveillance is crucial in solid tumors and T cell dysfunction leads to tumor progression. In contrast, the role of CD8+ T cells in the control of leukemia is less clear. We characterized the molecular signature of leukemia stem/progenitor cells (LSPCs) and paired CD8+ T cells in patients with acute myeloid leukemia (AML). Epigenetic alterations via histone deacetylation reduced the expression of immune-related genes in bone marrow (BM)-infiltrating CD8+ T cells. Surprisingly, a silenced gene expression pattern in CD8+ T cells significantly correlated with an improved prognosis. To define interactions between CD8+ T cells and LSPCs, we performed comprehensive correlative network modeling. This analysis indicated that CD8+ T cells contribute to the maintenance/expansion of LSPCs, particularly in favorable risk AML. Functionally, CD8+ T cells in favorable AML induced the expansion of LSPCs by stimulating the autocrine production of important hematopoietic cytokines such as interleukin (IL)-3. In contrast, LSPCs in aggressive AML were characterized by a higher activation of stemness/proliferation-related pathways and develop independent of BM CD8+ T cells. Overall, our study indicates that CD8+ T cells support and expand LSPCs in favorable risk AML whereas intermediate and adverse risk AML possess the intrinsic molecular abnormalities to develop independently.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is a heterogeneous group of hematologic malignant diseases, characterized by maturation arrest and increased proliferation of myeloid blasts [1, 2]. Based on the (cyto)genetic profile of AML blasts, patients can be classified into three different risk groups: favorable risk (40–45%), intermediate risk (25–35%), and adverse risk (25–30%) [3].
Leukemia stem cells (LSCs) represent a minor fraction of the bulk leukemia cell population that reconstitutes and propagates the disease. They are resistant to chemotherapy and irradiation and therefore are the main cause of relapse [4,5,6,7,8]. LSCs possess stem cell properties such as self-renewal and quiescence that are regulated by cell-intrinsic and cell-extrinsic mechanisms [9,10,11]. As cell-intrinsic drivers, LSCs display molecular abnormalities that lead to constitutive activation of the nuclear factor kB (NF-kB) [2], Wnt [12], and Notch signaling pathways [13]. In addition, similar to hematopoietic stem cells (HSCs), LSCs interact with cells of the bone marrow (BM) microenvironment, including endothelial cells, specialized BM stromal cells, and osteoblasts [14,15,16]. These cells constitute the HSC niche and regulate maintenance, self-renewal, and differentiation of HSCs and LSCs [17]. Immune cells are part of the BM microenvironment and interact with HSCs as well as leukemia stem and progenitor cells (LSPCs). For example, CD8+ T cells have the potential to support the maintenance of HSCs and facilitate their engraftment after allogeneic stem-cell transplantation [18, 19].
It is well documented that AML responds to immune-mediated therapies such as allogeneic hematopoietic stem cell transplantation and donor lymphocyte infusions [14, 20]. Similarly, AML blasts can be lysed by adoptively transferred gene-modified T cells in xenotransplantation experiments [21]. However, several studies indicate that LSCs escape elimination by CD8+ T cell and natural killer (NK) cells by down-regulating important molecules/pathways for immune recognition or by the expression of immune-inhibitory molecules [22, 23]. Experimental evidence indicates that cytokines secreted by immune cells and defined cell–cell interactions such as the CD70/CD27 interaction expand HSCs and LSCs and contribute to disease progression [24, 25]. However, the role of the adaptive immune system in the control of human AML and especially in the regulation of LSCs is poorly understood.
We performed a comprehensive transcriptomic profiling of BM-derived LSCs and leukemia progenitor cells together with paired CD8+ T cells of AML patients from different molecular risk groups. This analysis indicated that epigenetic mechanisms silence the gene expression of CD8+ T cells in AML. Importantly, a silenced gene expression pattern correlated with improved prognosis. Correlation network modeling revealed that CD8+ T cells regulate LSPC in favorable risk but not in adverse risk AML. Functionally, we show that CD8+ T cells induce the autocrine production of the hematopoietic cytokines such as IL-3 in favorable risk AML that expands LSPCs. The interaction of CD8+ T cells with LSPCs was gradually lost from favorable risk to intermediate and adverse risk AML. In contrast, LSPCs from patients with intermediate and adverse risk AML had a higher expression of genes related to stemness and cell proliferation. This study indicates that LSPCs in favorable risk AML are regulated by extrinsic signals such as BM-infiltrating CD8+ T cells, whereas mainly cell-intrinsic mechanisms drive LSPC expansion in aggressive AML.
Materials and methods
Patients
Blood and BM aspirates from patients diagnosed with AML at the Department of Medical Oncology, University Hospital Bern were prospectively collected. Thirty patients were selected from this repository based on the FACS immune-phenotype of the AML cells and the risk category. The clinical and molecular characteristics of the AML patients and controls are listed in Supplementary Table 1. This study was approved by the local ethical committee (Kantonale Ethikkommission Bern, KEK122/14).
Molecular profiling and correlation network modeling
Transcriptomic analysis was performed on 111 different well-defined FACS-purified samples of hematopoietic stem/progenitors and paired CD8+ T cells from 30 AML patients and 7 controls. Modeling of several tens of thousands of predictive correlation network was assessed to map potential links between genes expressed in stem/progenitor cells and paired CD8+ T cells in all AML patients of different risk groups and controls. Selection of investigated genes, biological signaling, and correlations were further functionally validated. For details, see Supplementary Methods.
Data availability
All transcriptomic data compiled for this study have been deposited in NCBI GEO under the accession code GSE117090. In addition, expression data from GEO public repository was assessed as a validation cohort (GSE6891).
Complete methods are included in the Supplementary Appendix.
Results
A silenced gene expression pattern in CD8+ T cells correlates with improved prognosis in AML
We first characterized the complete gene expression signature of FACS-purified CD34+CD38− AML LSCs, CD34+CD38+ AML progenitors and paired CD8+ T cells in the BM of AML patients at the time point of diagnosis (Supplementary Fig. 1, Supplementary Table 1). As expected, patients in the favorable risk group had a better overall survival in comparison to the other AML risk groups (Supplementary Fig. 2).
The expression of 258 genes was significantly changed in BM-infiltrating CD8+ T cells from AML patients compared to healthy donors (Supplemental Dataset 1). Interestingly, 222 genes were down-regulated and only 36 genes were up-regulated (Fig. 1a, b). The down-regulated genes are involved in signaling pathways related to T cell activation, differentiation, and function, such as NFkB, Wnt, FoxO, T cell receptor (TCR), and cytokine/chemokine signaling (Fig. 1c). The expression profile of key genes for T cell polarization in CD8+ T cells of AML patients revealed a TC1 phenotype in controls but a skewing towards a TC2 phenotype in AML patients (Fig. 1d). Out of the 222 down-regulated genes, we defined a 40-gene panel of genes that belong to the NFkB, Wnt, FoxO and Notch pathway, TCR and cytokine/chemokine signaling according to the results of the gene ontology enrichment and pathway analysis (Fig. 1c, Supplementary Table 2). Based on the calculated mean gene expression level of these 40 pre-defined genes, outcome-based cut-points were defined using X-Tile software [26]. AML patients with a mean gene expression below or above the defined cut-point were classified as patients with a “silenced gene expression (SGE) signature” or “non-silenced gene expression” (non-SGE) signature, respectively. Principal component analysis (PCA) revealed a closer similarity between the non-SGE group and healthy controls while patients with the SGE signature exhibited more distinct gene expression patterns (Fig. 1e). Importantly, patients with a SGE signature in BM CD8+ T cells had a significantly better overall survival compared to patients with a non-SGE signature (Fig. 1f). Although the analysis in the different risk groups is limited by a rather small sample size, the expression of a SGE signature correlated with longer survival only in the favorable risk but not in the intermediate or adverse risk group (Supplementary Fig. 3). Multivariate analysis for the SGE signature adjusted for AML risk group, patient age, percentage of blasts in BM and blood, leukocyte counts, and sex, confirmed the SGE signature as an independent prognostic marker for overall survival in our analyzed AML patient cohort (Fig. 1f).
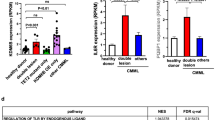
A silenced gene expression pattern in CD8+ T cells correlates with improved prognosis. a Volcano plot showing differentially expressed genes of leukemia CD8+ T cells (AML vs. Ctrl); y-axis: negative log of P-value; x-axis: log2-fold change; red dots: up-regulated genes; blue dots: down-regulated genes. b Heatmap illustrating differentially expressed genes in CD8+ T cells (AML vs. Ctrl). c Pathway enrichment analysis of 222 down-regulated genes. d Heatmap illustrating the expression profile of key genes for CD8+ T cell phenotype polarization. Silenced gene expression (SGE); non-silenced gene expression (non-SGE). e PCA indicating the similarities between CD8+ T cells with SGE or non-SGE signature and controls according to their gene expression profile. f Kaplan–Meier plots of overall survival (OS) for AML patients according to the 40-gene panel of CD8+ T cells. Multivariate analysis for SGE signature adjusted for AML risk groups, age, percentage of blasts in BM and blood, leukocyte counts, and sex. g Kaplan–Meier plots of overall survival (OS) for AML patients in the validation cohort according to the 40-gene signature of CD8+ T cells. Cox-regression analysis for SGE signature in the validation cohort. h Pathway enrichment analysis of 36 up-regulated genes. i Heatmap illustrating the expression profile of up-regulated histone organizer/regulator genes. j Karyogram panel shows significant enrichment of down-regulated genes to particular regions in the genome. Important selected genes are highlighted in blue. k Expression profile of selected genes from three AML patients with SGE signature (AML #15, favorable group; AML #29, intermediate group; AML #1, adverse group) and T cells from two AML patients with non-SGE signature (AML #7, favorable group and AML #28, adverse group). Analysis was performed immediately after FACS-purifying (untreated) and after 24 h treatment with VPA or vehicle. The fold differences of gene expression were calculated as the ratio of treated (VPA or vehicle) vs. untreated conditions. Statistics: f, g log-rank test; and multiple Cox-regression, k Student’s t-test (2-tailed). *P < 0.05, **P < 0.01, ***P < 0.001. CI confidence interval, OS overall survival. (See also Supplementary Fig. 2–3; Supplementary Table 2; Supplementary Dataset 1)
To validate the prognostic value of this 40-genes panel in a larger cohort of AML patients, we analyzed the expression of these genes in a publically available dataset comprising gene expression of non-fractionated BM cells of AML patients [27, 28]. Our defined 40-gene panel showed significantly better overall survival for patients with a SGE signature than for patients with a non-SGE signature. Cox-regression analysis further confirmed the SGE signature as a prognostic marker for overall survival (Fig. 1g).
In our analysis, we observed a systemic down-regulation of genes in AML CD8+ T cells compared to controls. Chromatin modification mainly via histone deacetylation is one of the key mechanisms for gene silencing [29]. Only 36 genes were up-regulated in AML CD8+ T cells (Fig. 1b). Pathway enrichment analysis revealed that these 36 up-regulated genes are primarily involved in the control of chromatin organization or negative regulation of transcription and gene expression (Fig. 1h). Nine of these 36 up-regulated genes are controlling chromatin organization/regulation (Fig. 1i). Chromosomal position-based gene-mapping analysis (karyogram) showed that down-regulated genes were not randomly distributed all over the genome but significantly enriched in some particular regions. These data suggest altered histone organization in particular chromosomal regions in leukemic CD8 T cells (Fig. 1j).
Histone deacetylation catalyzed by the histone deacetylase (HDAC) is a central switch of permissive to repressive chromatin domains leading to transcriptional silencing [30]. To functionally test the effect of histone deacetylation on the observed SGE signature, we treated FACS-purified CD8+ T cells isolated from three AML patients with SGE signature and from two AML patients with non-SGE signature with the HDAC inhibitor, valproic acid (VPA). VPA treatment significantly reversed the SGE phenotype and increased the expression of 10 selected key genes involved in the regulation of Wnt-, Notch-, MAPK-signaling, T cell differentiation, and in metabolism. In contrast, VPA did not change the expression of these genes in T cells with non-SGE signature (Fig. 1k). In addition, VPA treatment did not change the expression of IL3 in CD8+ T cells from different AML patients, indicating that IL3 gene expression in T cells is not regulated by histone deacetylation (Fig. 1k).
Overall, these data indicate that important genes involved in CD8+ T cell activation, differentiation, and function are down-regulated in AML due to pathologic epigenetic alterations mainly mediated via histone deacetylation. Down-regulation of these genes in CD8+ T cells correlates with an improved prognosis of AML patients.
LSPCs from AML patients display a dysregulated expression of genes involved in proliferation, stemness, and immune-recognition
We next investigated the gene expression signature of CD34+CD38− AML LSCs and CD34+CD38+ AML progenitors. HSPCs from healthy donors (Ctrl) had a similar gene expression profile and therefore clustered together in the PCA. In contrast, the gene expression of AML cells was more diverse and differed from healthy controls (Fig. 2a). In AML LSCs and progenitors, 403 and 1309 genes were differentially expressed compared to controls, respectively (Fig. 2a, Supplementary Dataset 1). A comprehensive pathway analysis of the differentially expressed genes revealed that mainly pathways related to stemness, cell proliferation, cell cycle, or immune cell signaling were changed in AML LSCs and progenitor cells (Fig. 2b). The number of 155 genes (58 up-regulated and 97 down-regulated genes) were similarly dysregulated in AML LSCs and progenitors when compared to controls (Fig. 2c). Pathway enrichment analysis revealed that up-regulated genes are mainly involved in cell proliferation, cell cycle, or immune-related signaling while down-regulated genes are primarily involved in signaling pathways mediating antigen-presentation and interaction with immune cells (Fig. 2c). Taken together, AML LSCs and progenitors had a higher expression of genes involved in stemness and cell proliferation compared to HSPCs whereas the down-regulated genes and pathways facilitate immune-escape.
Gene expression analysis of LSPCs from AML patients. a PCA of AML LSPCs and control HSPCs. Volcano plots representing differentially expressed genes of CD34+CD38− and CD34+CD38+ AML and control cells; y-axis: negative log of P-value; x-axis: log2-fold change; red dots: up-regulated genes; blue dots: down-regulated genes. b Pathway enrichment analysis of differentially expressed genes in each cell population. c Venn-diagram of significant up/down-regulated genes within leukemia stem/progenitor cells (AML vs. Ctrl). Heatmap indicating the profile of 155 differentially expressed intersection genes (58 up-regulated and 97 down-regulated genes). Pathway enrichment analysis of 58 up-regulated and 97 down-regulated intersection genes. (See also Supplementary Dataset 1)
Genes involved in immunosurveillance are down-regulated whereas stemness-related genes are up-regulated in adverse risk AML
We next compared the gene expression profiles of AML LSCs, progenitors, and paired CD8+ T cells between the three different risk categories (Fig. 3a). AML progenitors shared 294 dysregulated intersection genes across all AML risk categories. In contrast, only 67 intersection genes were found in AML LSCs. The gene expression profile of BM CD8+ T cells differed significantly between the different risk groups with only 39 intersection genes that were similarly altered in all three-risk categories (Fig. 3a). AML progenitors, which comprise the highly proliferative AML cells, showed the highest expression of genes related to cell cycling. In contrast, genes involved in stemness- and leukemogenesis-related pathways were expressed at higher levels in AML LSCs (e.g., Wnt, ErbB, and NF-kB). These stemness-related pathways were expressed at highest levels in LSCs from adverse risk AML and at lowest levels in LSCs from the favorable risk group. This finding is in line with previous studies indicating that a LSC-related gene signature is a negative prognostic marker in AML [31, 32]. Furthermore, genes involved in immune-related pathways such as cytokine–cytokine receptor interactions and cytokine/chemokine signaling were more expressed in LSCs from favorable risk AML but were not expressed in adverse risk AML (Fig. 3b).
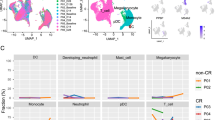
Gene expression signature of leukemia stem/progenitor cells and paired CD8+ T cells across AML risk groups. a Venn-diagrams showing differentially expressed genes in CD34+CD38− LSCs, CD34+CD38+ progenitors and CD8+ T cells across AML risk groups (favorable, intermediate, or adverse vs. Ctrl). b Pathway analysis of differentially expressed genes in LSPCs and CD8+ T cells across AML risk groups. c Gene set enrichment analysis (GSEA) representing the normalized enrichment score (NES) of gene sets linked to Wnt signaling for LSPCs and CD8+ T cells (favorable, intermediate, or adverse vs. Ctrl). Bar charts showing the number of up-regulated Wnt-related/target genes in AML patients from different risk categories vs. controls. Statistics: Student’s t-test (2-tailed). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. (See also Supplementary Dataset 1)
In line with these findings, genes involved in pathways mediating immune cell function and target cell recognition such as TCR and chemokine signaling as well as cytokine–cytokine receptor interaction were more expressed in paired CD8+ T cells from favorable risk AML but were down-regulated in the adverse risk group. Similarly, signaling pathways involved in T cell effector function, such as NF-kB and Notch signaling, were significantly down-regulated in CD8+ T cells isolated from intermediate and adverse risk AML patients [33]. The Wnt pathway, which is crucial for the differentiation into memory T cells [33, 34], was inactivated in CD8+ T cells from patients with adverse risk AML (Fig. 3b). These results suggest that T cells in intermediate and high risk AML are less functional than in favorable risk AML and might be exhausted. However, the analysis of markers defining exhausted CD8+ T cells on gene level did not reveal differences between the AML risk groups (Supplementary Fig. 4). Importantly, the frequency of BM-infiltrating CD8+ T cells did not differ between the AML risk groups or healthy controls (Supplementary Fig. 5). This indicates that functional but not numerical differences lead to the CD8+ T cell-mediated expansion of LSPCs in favorable risk AML.
Wnt signaling is fundamental for stemness and effector function of LSCs and CD8+ T cells, respectively [12]. Gene set enrichment analysis (GSEA) in our AML cohort revealed a significant enrichment for genes involved in Wnt pathway activation in LSPCs from intermediate and adverse risk group AML patients, but not in LSPCs from the favorable risk group. In contrast, Wnt-related genes were down-regulated in paired CD8+ T cells from AML patients of the intermediate and adverse risk group (Fig. 3c). These results suggest that LSPCs of adverse risk AML patients develop and expand largely independently of CD8+ T cells, whereas CD8+ T cells are mainly active in favorable risk AML patients.
CD8+ T cells regulate LSPCs in favorable but not adverse risk AML
To define possible interactions of genes/pathways in CD8+ T cells with genes/pathways in AML LSPCs, comprehensive correlation networks were constructed. In each network, a node was defined as a gene expressed either in CD8+ T cells or in LSPCs. A node (gene) in one cell that correlates significantly with more than 15 nodes in the other cell type was considered as a hub (high-degree correlated node) (Fig. 4a). The assumption was that a gene expressed in CD8+ T cells communicates or coordinates other genes in LSPCs or vice versa. Three types of correlations were detected: (1) “appear”, a correlation present in AML but not in controls, (2) “disappear”, a correlation present in the controls and absent in AML, and (3) “flip”, where the direction of the correlation changes.
Correlation network modeling between leukemia stem/progenitor cells and paired CD8+ T cells. a Schematic view of correlation networks for AML samples (“disease”). The red box represents hubs with a high degree of correlations and black box indicates low degree correlated nodes. b Quantification of hub and node genes in LSPCs and CD8+ T cells. c Schematic view of correlation networks that only appear in one or more of the AML groups but not in the healthy controls (“appear”); networks which are only detectable in control samples but not in any of the AML risk groups (“disappear”); networks that are significant in both the control group and patient groups but have opposite signs (“flip”). d Bar graph representing the number of node gene in the appear, disappear, and flip networks, in different AML risk categories. e Quantification of hub genes in the appear network in AML risk categories. f Venn-diagram of CD8+ T cells’ hubs genes in different AML risk categories. g Pathway analysis of node genes within LSPCs that show significant correlation to CD8+ T cell hubs. (See also Supplementary Fig. 6–7; Supplementary Table 3; Supplementary Dataset 2)
The highest number of nodes was detected in AML progenitor cells and in CD8+ T cells, whereas few nodes were detected in AML LSCs (Fig. 4a, b). Interestingly, the highest number of hubs were detected in CD8+ T cells, whereas fewer hubs were found in LSPCs (Fig. 4a, b, Supplementary Dataset 2). We next analyzed these networks in the different AML risk categories (Fig. 4c, d). The number of nodes in the “appear network” in AML LSPCs and CD8+ T cells gradually decreased from the favorable risk to the adverse risk group (Fig. 4d). In contrast, the number of nodes in “disappear” and “flip networks” did not change across AML risk groups (Fig. 4c, d). Importantly, the majority of hubs in CD8+ T cells were present in the favorable risk group and their number gradually decreased in the intermediate and adverse risk groups (Fig. 4e). No CD8+ T cells hubs were detected in intermediate and adverse risk groups that were not present in the favorable risk group (Fig. 4e, f). Only few hubs were identified in CD34+CD38+ progenitors. However, similar to CD8+ T cells, reduced number of hubs were identified with increased risk category (Fig. 4e, Supplementary Fig. 6).
The majority of nodes within AML LSPCs, which were connected to CD8+ hubs, were up-regulated compared to controls (Supplementary Fig. 7). The number of nodes in LSPCs connected to given hubs in CD8+ T cells was gradually decreased from favorable to intermediate and adverse risk AML (Supplementary Table 3).
In order to identify the signaling pathways in AML LSPCs, which are modulated by the identified CD8+ T cell hubs, we performed a pathway analysis. The significantly altered pathways in LSPCs included stemness, cell proliferation and survival, immune-related signaling, gene expression regulation, growth factor signaling, and metabolism which were most activated in favorable risk AML (Fig. 4g, Supplementary Fig. 7).
Taken together, our correlation network modeling indicated that CD8+ T cells regulate LSPCs in favorable risk AML. This interaction is reduced or absent in intermediate and adverse risk AML.
CD8+ T cells induce the expansion and maintenance of LSPCs in favorable risk AML
To functionally analyze the potential influence of CD8+ T cells on LSPCs, we FACS-purified both LSCs or AML progenitor cells together with paired CD8+ T cells from BM of nine different AML patients (Fig. 5a, Supplementary Table 1). The co-culture of CD8+ T cells with LSCs resulted in an up to 2-fold increase in colony-formation compared to LSC mono-culture in the favorable risk group. In contrast, the increase in colony formation in intermediate and adverse risk was less pronounced and only observed at higher T cell:LSC ratios (Fig. 5b). In addition, the number of cells per colonies was not significantly changed, suggesting that differentiation was not affected by co-culture of LSPCs with CD8+ T cells (Fig. 5c). Further re-plating of cells derived from primary colony assays showed a significant difference in the colony-formation capacity of co-cultured LSCs with a higher ratio of CD8+ T cells compared to LSC mono-culture. This indicates that LSCs with unimpaired self-renewal capacity are expanded by CD8+ T cells (Supplementary Fig. 8).
CD8+ T cells regulate LSPCs in favorable risk AML. a Experimental setup. b, c Fold differences in the numbers of colonies or cells from co-cultured conditions (LSCs with CD8+ T cells) vs. mono-cultured (LSCs only) condition (three patients per each AML subgroup; #3, #22, and #28 from adverse risk group; #5, #17, and #21 from intermediate risk group; #14, #15, and #26 from favorable risk group). d An example of a correlation network in favorable group: MGAT5 hub gene in CD8+ T cells is predicted as a coordinator for different genes in LSPCs. e Correlation analysis of MGAT5 hub in CD8+ T cells vs. IL3 in CD34+CD38+ cells in different AML risk categories. f IL3 gene expression in re-purified LSPCs from mono-culture or co-culture with CD8+ T cells; (three AML patients from favorable risk category). IL-3 protein in culture supernatants of LSPCs monocultures and co-cultures with CD8+ T cells. g MGAT5 and IL3 gene expression quantification upon siRNA-mediated gene silencing of MGAT5. The fold differences of gene expression were calculated as the ratio of siMGAT5 treated condition vs. siCtrl in re-purified cell populations (three AML patients from favorable risk category). h Correlation analysis of MGAT5 expression in CD8+ T cells vs. IL3 in LSPCs after MGAT5 gene knockdown. i IL3 gene expression in FACS-purified favorable risk AML CD34+ LSPCs from mono-culture or co-culture with CD8+ T cells in the presence of neutralizing antibodies to EGFR, IL-2, or IFNγ (n = 3; mean ± SD). j Fold differences in the numbers of colonies from LSC monocultures- and co-cultures with CD8+ T cells upon neutralization with αIL-3 antibody (three patients from favorable risk AML). k Heatmap illustrating expression profile of five selected genes in re-purified LSPCs after mono-culture or co-culture with CD8+ T cells (three AML patients from favorable risk category). l Graphical scheme of the interaction between CD8+ T cells and LSPCs. Statistics: Student’s t-test (2-tailed). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. (See also Supplementary Fig. 8–11; Supplementary Dataset 2)
MGAT5 was identified as a hub in CD8+ T cells of all risk groups. However, while MGAT5 positively correlated with the expression of IL3 and other important genes in CD34+CD38+ progenitors in favorable risk AML patients, this correlation is lost in intermediate and adverse risk AML (Fig. 5d, e, Supplementary Fig. 9, Supplementary Dataset 2).
Our results suggest that CD8+ T cells regulate LSPCs in favorable AML by inducing the production of important hematopoietic cytokines such as IL-3. To address this possibility, we co-cultured FACS-purified CD8+ T cells with paired LSPCs derived from the same patients of favorable risk AML and analyzed the expression of IL3 mRNA and protein. LSPCs of all risk groups expressed the IL-3 receptor alpha (IL-3Ra). While, CD8+ T cells did not express IL3Ra on mRNA and protein levels (Supplementary Fig. 10). Co-culture with CD8+ T cells resulted in an up to 3-fold increase in IL3 mRNA expression in LSPCs and an up to 2-fold increase in IL-3 protein expression in culture supernatants compared to LSPCs mono-culture from favorable risk AML (Fig. 5f). In healthy controls, the expression of MGAT5 in CD8+ T cells did not correlate with IL3 expression in progenitors (Supplementary Fig. 11a). In addition, co-culture of CD8+ T cells with CD34+ HSPCs derived from healthy donors did not increase the level of IL3 mRNA expression (Supplementary Fig. 11b). However, co-culture of pooled CD8+ T cells derived from three good risk AML patients significantly increased the colony formation capacity of both, AML LSCs and control HSCs (Supplementary Fig. 11c). Interestingly, colony formation in LSCs was increased significantly more than in HSCs. This suggests that AML LSCs have a higher proliferative capacity in response to stimulation by CD8+ T cells than normal HSCs. Therefore, favorable risk AML LSPCs respond to similar interactions with CD8+ T cells as normal HSPCs. However, only BM-infiltrating CD8+ T cells of AML patients but not of healthy donors induce IL-3 production in LSCs and HSCs.
Knockdown of MGAT5 gene in CD8+ T cells using a siRNA before initiation of the co-culture revealed an up to 2-fold decrease in IL3 mRNA expression in LSPCs (Fig. 5g). The level of MGAT5 gene expression after siRNA treatment of CD8 T cells significantly correlated with the IL3 expression in co-cultured LSPCs (Fig. 5h). In silico pathway analysis predicted that MGAT5 expressed in CD8+ T cells triggers the expression of IL3 in LSPCs via EGF/EGFR, IL-2 or IFNγ signaling (Supplementary Fig. 9a). Correlation analysis confirmed a significant positive correlation between IL3 expression in CD34+CD38+ progenitors and EGF/EGFR or IL2 expression in CD8+ T cells in favorable risk AML but not in intermediate or adverse risk AML (Supplementary Fig. 9b–c). We tested this concept experimentally by co-culturing FACS-purified CD34+ LSPCs with paired CD8+ T cells in the presence of neutralizing antibodies to these specific cytokines and growth factor receptor. Blocking EGFR and IL-2, but not IFNγ reduced IL3 mRNA expression in FACS-purified CD34+ LSPCs (Fig. 5i). This finding indicates that CD8+ T cells induce IL-3 production in LSPCs mainly by IL-2 and EGF/EGFR signaling.
Neutralization of IL-3 in the co-culture of CD8+ T cells with LSCs from favorable risk AML patients significantly decreased the colony-formation capacity (Fig. 5j). In addition, co-culture of CD34+ LSPCs with CD8+ T cells resulted in the up-regulation of selected genes that are regulated by hubs with documented function in hematopoiesis, leukemia development, cell proliferation, or immune tolerance (CD47, CD58, CD63, CD99, and IL17b) (Fig. 5k, Supplementary Table 3) [35,36,37,38,39].
Taken together, these findings indicated that BM CD8+ T cells induce pathways in favorable risk LSPCs that promote leukemia development, such as the autocrine production of hematopoietic cytokines. In contrast, LSPCs in more aggressive forms of AML develop largely independent of CD8+ T cells (Fig. 5l).
Discussion
It is well documented that the immune system contributes to the control of solid tumors, a process called “tumor immunosurveillance”. However, cancer cells evade immune recognition and elimination by cytotoxic CD8+ lymphocytes via various mechanisms, including loss of antigen and the expression of immune inhibitory molecules [40, 41]. Thus, tumor-infiltrating lymphocytes (TILs) are often dysfunctional or “exhausted” due to the interaction with immune inhibitory ligands expressed by tumor cells [40]. The presence of dysfunctional CD8+ T cells in TILs is a negative prognostic and predictive factor for the response to treatment with immune-checkpoint inhibitors [42, 43].
In the present study, we analyzed the gene expression signature of BM CD8+ T cells together with paired AML LSCs and progenitor cells. The majority of differentially expressed genes were down-regulated in CD8+ T cells derived from AML patients. The down-regulated genes are involved in key functions of T cell activation and differentiation including NFkB, Notch, Wnt, FoxO, TCR, and cytokine/chemokine signaling. This indicates that similar to TILs in solid tumors, CD8+ T cells in the AML BM are dysfunctional. Surprisingly, patients with dysfunctional CD8+ T cells as indicated by a SGE signature had a significantly better survival compared to patients with non-SGE signature. This difference may be explained by the fact that TILs comprise a population of activated tumor-specific T cells whereas the BM is a secondary lymphoid organ that contains mainly memory T cells with different specificities [44].
The up-regulated genes in CD8+ T cells of AML patients included main drivers of histone deacetylation and epigenetic regulation, which correlated with the SGE signature in our study population. HDAC enzymes regulate key functions in T cells, such as maturation, migration, and TCR signaling [45]. Recently, it was reported that the transcription factors Tcf1 and Lef1 favor CD8+ T cell development by HDAC mediated suppression of lineage-inappropriate genes [46]. In addition, HDAC inhibitors enhance anti-tumor activity of antigen-specific cytotoxic T cells against solid tumors and multiple myeloma [47]. Similarly, treatment of AML and myelodysplasia patients with azacitidine and VPA induces a CD8+ T-cell response to the MAGE cancer testis antigen [48]. This suggests that a SGE signature of T cells might be reversed by treatment with demethylating agents.
To define possible interactions of CD8+ T cells with human AML LSPCs, we performed a comprehensive correlation network analysis. The goal of this analysis was to identify genes in CD8+ T cells that regulate pathways in LSPCs and vice versa (hubs). Interestingly, most hub genes were identified in CD8+ T cells whereas only few hubs were present in LSPCs. The number of CD8+ T cell hubs was the highest in favorable risk AML and then gradually lower from intermediate to adverse risk AML. In addition, a given CD8+ T cell hub correlated with the highest number of genes (nodes) in LSPCs from favorable risk group and this number was gradually reduced in intermediate to adverse risk AML. In addition, AML progenitors expressed more node genes than AML LSCs, indicating that CD8+ T cells predominantly interact with progenitor cells. Our finding suggests that CD8+ T cells regulate LSPCs in favorable risk AML while more aggressive forms of AML develop independently of BM-infiltrating CD8+ T cells.
According to our comprehensive network modeling, hubs in CD8+ T cells modulate stemness, cell-proliferation and cell-survival, immune-related signaling, and growth factor signaling mainly in favorable risk AML. CD8+ T cells’ hub genes in favorable risk AML, correlate with an increased gene expression of cytokine/chemokines such as IL3 and IL17B, and cell surface molecules (e.g., CD47, CD58, CD63, and CD99) that have been associated with hematopoiesis, leukemia development, cell proliferation, or immune tolerance [35,36,37,38,39, 49].
Our results indicated that CD8+ T cells induce IL3 production in AML LSPCs via IL2 and EGF signaling leading to their expansion. This is in agreement with reports documenting that IL-3 increases proliferation and expansion of CML stem cells and AML blasts [50, 51]. Interestingly, the level of IL-3Ra expression on AML blasts is associated with increased cellularity, enhanced proliferation, and with poor prognosis [52].
CD8+ T cells regulate LSPCs mainly in favorable risk AML, while more aggressive forms of AML develop independently of BM-infiltrating CD8+ T cells. The frequency of AML-specific T cells is rather low [53, 54]. Therefore, the majority of the analyzed CD8+ T cells in the BM may be BM-resident memory CD8+ T cells that are not leukemia-specific. We document that in adverse risk AML, the positive correlation between IL2 or EGF expressed in CD8+ T cells to IL3 in AML progenitors was lost (Supplementary Fig. 9c). We did not detect differences in the level of gene silencing in CD8+ T cells in different risk categories (Fig. 1, Supplementary Dataset 1). In addition, we also tested the expression of markers used to characterize exhausted CD8+ T cells (CD244 (2B4), CD160, TIGIT, HAVCR2, LAG3, PDCD1 (PD-1)). This analysis did not reveal a preferential expression of exhaustion markers in T cells of intermediate or adverse risk AML (Supplementary Fig. 4). In addition, genes involved in immune-related pathways such as cytokine–cytokine receptor interactions and cytokine/chemokine signaling were expressed in LSCs from favorable risk AML but were not expressed in adverse risk AML (Fig. 3b). Therefore, we suggest that the lack of interaction between CD8+ T cells and LSPCs in intermediate and adverse risk AML is not due to alterations in T cells but due to molecular changes in LSPCs in intermediate and high risk AML that render them independent in regard of proliferation and expansion. AML LSCs and progenitors in aggressive AML had a higher expression of genes involved in stemness and cell proliferation (as an intrinsic driver of leukemia) compared to LSPCs in favorable risk AML, whereas the down-regulated genes and pathways facilitated immune-escape and immune-tolerance. Stemness signatures in blasts are an important negative prognostic marker [12, 24]. The canonical Wnt pathway, which is central for HSC maintenance and development, is constitutively active in myeloid leukemia and of crucial importance for LSCs [24, 55, 56]. Fusion proteins such as AML1-ETO, MLL-ENL, PLZF-RARα, and PML-RARα induce Wnt signaling via activation of γ-catenin [57,58,59]. In addition, activating mutations in FLT3 are associated with high β-catenin levels and correlate with poor overall survival in AML patients [59, 60]. We document that stemness-related pathways, especially the Wnt pathway, were more active in adverse risk than in intermediate or favorable risk AML.
AML is a very heterogeneous disease. It is therefore not surprising that the interaction of CD8+ T cells with LSPCs also varies in different molecular subtypes of AML. Interestingly, although the prognostic risk groups still comprise a molecular very heterogeneous group of diseases, we found a clear difference in the regulation of LSPCs. Favorable risk AML have less intrinsic molecular abnormalities that drive proliferation and stemness. The disease development depends on external cues from the niche, e.g., from CD8+ T cells. In contrast, more aggressive AML is propagated mainly by cell-intrinsic mechanisms and develop independent of immune cells.
References
Estey E, Dohner H. Acute myeloid leukaemia. Lancet. 2006;368:1894–907.
Hoesel B, Schmid JA. The complexity of NF-κB signaling in inflammation and cancer. Mol Cancer. 2013;12:86.
Zeisig BB, Kulasekararaj AG, Mufti GJ, So CW. SnapShot: acute myeloid leukemia. Cancer Cell. 2012;22:698.
Lapidot T, Sirard C, Vormoor J, Murdoch B, Hoang T, Caceres-Cortes J, et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature. 1994;367:645–8.
Hope KJ, Jin L, Dick JE. Acute myeloid leukemia originates from a hierarchy of leukemic stem cell classes that differ in self-renewal capacity. Nat Immunol. 2004;5:738–43.
Guzman ML, Allan JN. Concise review: leukemia stem cells in personalized medicine. Stem Cells. 2014;32:844–51.
Murone M, Radpour R, Attinger A, Chessex AV, Huguenin AL, Schurch CM, et al. The multi-kinase inhibitor Debio 0617B reduces maintenance and self-renewal of primary human AML CD34(+) stem/progenitor cells. Mol Cancer Ther. 2017;16:1497–510.
Radpour R, Forouharkhou F. Single-cell analysis of tumors: creating new value for molecular biomarker discovery of cancer stem cells and tumor-infiltrating immune cells. World J Stem Cells. 2018;10:160–71.
Rice KN, Jamieson CH. Molecular pathways to CML stem cells. Int J Hematol. 2010;91:748–52.
Passegue E, Weisman IL. Leukemic stem cells: where do they come from? Stem Cell Rev. 2005;1:181–8.
Radpour R. Tracing and targeting cancer stem cells: new venture for personalized molecular cancer therapy. World J Stem Cells. 2017;9:169–78.
Staal FJ, Famili F, Garcia Perez L, Pike-Overzet K. Aberrant Wnt signaling in leukemia. Cancers. 2016;8:78.
Tohda S. NOTCH signaling roles in acute myeloid leukemia cell growth and interaction with other stemness-related signals. Anticancer Res. 2014;34:6259–64.
Sanchez-Aguilera A, Mendez-Ferrer S. The hematopoietic stem-cell niche in health and leukemia. Cell Mol Life Sci. 2017;74:579–90.
Doron B, Handu M, Kurre P. Concise review: adaptation of the bone marrow stroma in hematopoietic malignancies: current concepts and models. Stem Cells. 2018;36:304–12.
Riether C, Schurch CM, Ochsenbein AF. Regulation of hematopoietic and leukemic stem cells by the immune system. Cell Death Differ. 2015;22:187–98.
Shiozawa Y, Havens AM, Pienta KJ, Taichman RS. The bone marrow niche: habitat to hematopoietic and mesenchymal stem cells, and unwitting host to molecular parasites. Leukemia. 2008;22:941–50.
Geerman S, Brasser G, Bhushal S, Salerno F, Kragten NA, Hoogenboezem M, et al. Memory CD8(+) T cells support the maintenance of hematopoietic stem cells in the bone marrow. Haematologica. 2018;103:e230–3.
Gandy KL, Domen J, Aguila H, Weissman IL. CD8+TCR+ and CD8+TCR− cells in whole bone marrow facilitate the engraftment of hematopoietic stem cells across allogeneic barriers. Immunity. 1999;11:579–90.
Hale G, Waldmann H. Control of graft-versus-host disease and graft rejection by T cell depletion of donor and recipient with Campath-1 antibodies. Results of matched sibling transplants for malignant diseases. Bone Marrow Transplant. 1994;13:597–611.
Tasian SK, Kenderian SS, Shen F, Ruella M, Shestova O, Kozlowski M, et al. Optimized depletion of chimeric antigen receptor T cells in murine xenograft models of human acute myeloid leukemia. Blood. 2017;129:2395–407.
Austin R, Smyth MJ, Lane SW. Harnessing the immune system in acute myeloid leukaemia. Crit Rev Oncol Hematol. 2016;103:62–77.
Masarova L, Kantarjian H, Garcia-Mannero G, Ravandi F, Sharma P, Daver N. Harnessing the immune system against leukemia: monoclonal antibodies and checkpoint strategies for AML. Adv Exp Med Biol. 2017;995:73–95.
Riether C, Schurch CM, Buhrer ED, Hinterbrandner M, Huguenin AL, Hoepner S, et al. CD70/CD27 signaling promotes blast stemness and is a viable therapeutic target in acute myeloid leukemia. J Exp Med. 2017;214:359–80.
Riether C, Schurch CM, Flury C, Hinterbrandner M, Druck L, Huguenin AL, et al. Tyrosine kinase inhibitor-induced CD70 expression mediates drug resistance in leukemia stem cells by activating Wnt signaling. Sci Transl Med. 2015;7:298ra119.
Camp RL, Dolled-Filhart M, Rimm DL. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin Cancer Res. 2004;10:7252–9.
Verhaak RG, Wouters BJ, Erpelinck CA, Abbas S, Beverloo HB, Lugthart S, et al. Prediction of molecular subtypes in acute myeloid leukemia based on gene expression profiling. Haematologica. 2009;94:131–4.
de Jonge HJ, Valk PJ, Veeger NJ, ter Elst A, den Boer ML, Cloos J, et al. High VEGFC expression is associated with unique gene expression profiles and predicts adverse prognosis in pediatric and adult acute myeloid leukemia. Blood. 2010;116:1747–54.
Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705.
Verdone L, Caserta M, Di Mauro E. Role of histone acetylation in the control of gene expression. Biochem Cell Biol. 2005;83:344–53.
Gentles AJ, Plevritis SK, Majeti R, Alizadeh AA. Association of a leukemic stem cell gene expression signature with clinical outcomes in acute myeloid leukemia. J Am Med Assoc. 2010;304:2706–15.
Ng SW, Mitchell A, Kennedy JA, Chen WC, McLeod J, Ibrahimova N, et al. A 17-gene stemness score for rapid determination of risk in acute leukaemia. Nature. 2016;540:433–7.
Backer RA, Hombrink P, Helbig C, Amsen D. The fate choice between effector and memory T cell lineages: asymmetry, signal integration, and feedback to create bistability. Adv Immunol. 2018;137:43–82.
Sorcini D, Bruscoli S, Frammartino T, Cimino M, Mazzon E, Galuppo M, et al. Wnt/β-catenin signaling induces integrin α4β1 in T cells and promotes a progressive neuroinflammatory disease in mice. J Immunol. 2017;199:3031–41.
Galli S, Zlobec I, Schurch C, Perren A, Ochsenbein AF, Banz Y. CD47 protein expression in acute myeloid leukemia: a tissue microarray-based analysis. Leuk Res. 2015;39:749–56.
Kong F, Gao F, Li H, Liu H, Zhang Y, Zheng R, et al. CD47: a potential immunotherapy target for eliminating cancer cells. Clin Transl Oncol. 2016;18:1051–5.
McArdel SL, Terhorst C, Sharpe AH. Roles of CD48 in regulating immunity and tolerance. Clin Immunol. 2016;164:10–20.
Seubert B, Cui H, Simonavicius N, Honert K, Schafer S, Reuning U, et al. Tetraspanin CD63 acts as a pro-metastatic factor via beta-catenin stabilization. Int J Cancer. 2015;136:2304–15.
Yamaguchi Y, Fujio K, Shoda H, Okamoto A, Tsuno NH, Takahashi K, et al. IL-17B and IL-17C are associated with TNF-alpha production and contribute to the exacerbation of inflammatory arthritis. J Immunol. 2007;179:7128–36.
Baitsch L, Baumgaertner P, Devevre E, Raghav SK, Legat A, Barba L, et al. Exhaustion of tumor-specific CD8(+) T cells in metastases from melanoma patients. J Clin Invest. 2011;121:2350–60.
Gorenshteyn D, Zaslavsky E, Fribourg M, Park CY, Wong AK, Tadych A, et al. Interactive big data resource to elucidate human immune pathways and diseases. Immunity. 2015;43:605–14.
Fridman WH, Pages F, Sautes-Fridman C, Galon J. The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer. 2012;12:298–306.
Ji RR, Chasalow SD, Wang L, Hamid O, Schmidt H, Cogswell J, et al. An immune-active tumor microenvironment favors clinical response to ipilimumab. Cancer Immunol Immunother. 2012;61:1019–31.
Di Rosa F, Gebhardt T. Bone marrow T cells and the integrated functions of recirculating and tissue-resident memory T cells. Front Immunol. 2016;7:51
Haery L, Thompson RC, Gilmore TD. Histone acetyltransferases and histone deacetylases in B- and T-cell development, physiology and malignancy. Genes Cancer. 2015;6:184–213.
Xing S, Li F, Zeng Z, Zhao Y, Yu S, Shan Q, et al. Tcf1 and Lef1 transcription factors establish CD8(+) T cell identity through intrinsic HDAC activity. Nat Immunol. 2016;17:695–703.
Bae J, Hideshima T, Tai YT, Song Y, Richardson P, Raje N, et al. Histone deacetylase (HDAC) inhibitor ACY241 enhances anti-tumor activities of antigen-specific central memory cytotoxic T lymphocytes against multiple myeloma and solid tumors. Leukemia. 2018;32:1932–47.
Goodyear O, Agathanggelou A, Novitzky-Basso I, Siddique S, McSkeane T, Ryan G, et al. Induction of a CD8+ T-cell response to the MAGE cancer testis antigen by combined treatment with azacitidine and sodium valproate in patients with acute myeloid leukemia and myelodysplasia. Blood. 2010;116:1908–18.
Chung SS, Eng WS, Hu W, Khalaj M, Garrett-Bakelman FE, Tavakkoli M, et al. CD99 is a therapeutic target on disease stem cells in myeloid malignancies. Sci Transl Med. 2017;9:eaaj2025.
Nievergall E, Ramshaw HS, Yong AS, Biondo M, Busfield SJ, Vairo G, et al. Monoclonal antibody targeting of IL-3 receptor alpha with CSL362 effectively depletes CML progenitor and stem cells. Blood. 2014;123:1218–28.
Onetto-Pothier N, Aumont N, Haman A, Park L, Clark SC, De Lean A, et al. IL-3 inhibits the binding of GM-CSF to AML blasts, but the two cytokines act synergistically in supporting blast proliferation. Leukemia. 1990;4:329–36.
Testa U, Riccioni R, Militi S, Coccia E, Stellacci E, Samoggia P, et al. Elevated expression of IL-3Ralpha in acute myelogenous leukemia is associated with enhanced blast proliferation, increased cellularity, and poor prognosis. Blood. 2002;100:2980–8.
Le Dieu R, Taussig DC, Ramsay AG, Mitter R, Miraki-Moud F, Fatah R, et al. Peripheral blood T cells in acute myeloid leukemia (AML) patients at diagnosis have abnormal phenotype and genotype and form defective immune synapses with AML blasts. Blood. 2009;114:3909–16.
Narita M, Takahashi M, Liu A, Nikkuni K, Furukawa T, Toba K, et al. Leukemia blast-induced T-cell anergy demonstrated by leukemia-derived dendritic cells in acute myelogenous leukemia. Exp Hematol. 2001;29:709–19.
Staal FJ, Clevers HC. WNT signalling and haematopoiesis: a WNT–WNT situation. Nat Rev Immunol. 2005;5:21–30.
Wang Y, Krivtsov AV, Sinha AU, North TE, Goessling W, Feng Z, et al. The Wnt/beta-catenin pathway is required for the development of leukemia stem cells in AML. Science. 2010;327:1650–3.
Yeung J, Esposito MT, Gandillet A, Zeisig BB, Griessinger E, Bonnet D, et al. β-Catenin mediates the establishment and drug resistance of MLL leukemic stem cells. Cancer Cell. 2010;18:606–18.
Zheng X, Beissert T, Kukoc-Zivojnov N, Puccetti E, Altschmied J, Strolz C, et al. Gamma-catenin contributes to leukemogenesis induced by AML-associated translocation products by increasing the self-renewal of very primitive progenitor cells. Blood. 2004;103:3535–43.
Xu J, Suzuki M, Niwa Y, Hiraga J, Nagasaka T, Ito M, et al. Clinical significance of nuclear non-phosphorylated beta-catenin in acute myeloid leukaemia and myelodysplastic syndrome. Br J Haematol. 2008;140:394–401.
Ysebaert L, Chicanne G, Demur C, De Toni F, Prade-Houdellier N, Ruidavets JB, et al. Expression of beta-catenin by acute myeloid leukemia cells predicts enhanced clonogenic capacities and poor prognosis. Leukemia. 2006;20:1211–6.
Acknowledgements
This work was supported by grants from the Werner und Hedy Berger-Janser Stiftung, Swiss National Science Foundation, and the Stiftung für klinisch-experimentelle Tumorforschung (Bern). The authors thank Ursina Lüthi for excellent technical support. The authors thank the staff of the FACSLab (Department of BioMedical Research (DBMR), University of Bern, Switzerland) for providing technical assistance.
Author information
Authors and Affiliations
Contributions
RR designed and performed experiments, analyzed and interpreted data, and wrote the manuscript. CR designed experiments and interpreted data. CS and RB analyzed the data. SH performed experiments. AFO designed experiments, wrote the manuscript, and supervised the project.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Radpour, R., Riether, C., Simillion, C. et al. CD8+ T cells expand stem and progenitor cells in favorable but not adverse risk acute myeloid leukemia. Leukemia 33, 2379–2392 (2019). https://doi.org/10.1038/s41375-019-0441-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-019-0441-9
This article is cited by
-
Circulating miRNA panels as a novel non-invasive diagnostic, prognostic, and potential predictive biomarkers in non-small cell lung cancer (NSCLC)
British Journal of Cancer (2024)
-
Aged hematopoietic stem cells entrap regulatory T cells to create a prosurvival microenvironment
Cellular & Molecular Immunology (2023)
-
Characteristics of circulating adaptive immune cells in patients with colorectal cancer
Scientific Reports (2022)
-
Azacitidine-induced reconstitution of the bone marrow T cell repertoire is associated with superior survival in AML patients
Blood Cancer Journal (2022)
-
Alterations of T-cell-mediated immunity in acute myeloid leukemia
Oncogene (2020)