Abstract
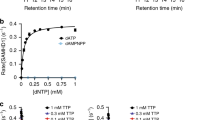
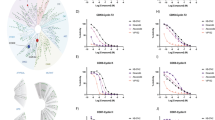
Activating mutations in NT5C2, a gene encoding cytosolic purine 5′-nucleotidase (cN-II), confer chemoresistance in relapsed acute lymphoblastic leukemia. Here we show that all mutants became independent of allosteric effects of ATP and thus constitutively active. Structural mapping of mutations described in patients demonstrates that 90% of leukemia-specific allelles directly affect two regulatory hotspots within the cN-II molecule—the helix A region: residues 355–365, and the intersubunit interface: helix B (232–242) and flexible interhelical loop L (400–418). Furthermore, analysis of hetero-oligomeric complexes combining wild-type (WT) and mutant subunits showed that the activation is transmitted from the mutated to the WT subunit. This intersubunit interaction forms structural basis of hyperactive NT5C2 in drug-resistant leukemia in which heterozygous NT5C2 mutation gave rise to hetero-tetramer mutant and WT proteins. This enabled us to define criteria to aid the prediction of NT5C2 drug resistance mutations in leukemia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Li BS, Li H, Bai Y, Kirschner-Schwabe R, Yang JJ, Chen Y, et al. Negative feedback-defective PRPS1 mutants drive thiopurine resistance in relapsed childhood ALL. Nat Med. 2015;21:563–71.
Tzoneva G, Perez-Garcia A, Carpenter Z, Khiabanian H, Tosello V, Allegretta M, et al. Activating mutations in the NT5C2 nucleotidase gene drive chemotherapy resistance in relapsed ALL. Nat Med. 2013;19:368–71.
Meyer JA, Wang J, Hogan LE, Yang JJ, Dandekar S, Patel JP, et al. Relapse-specific mutations in NT5C2 in childhood acute lymphoblastic leukemia. Nat Genet. 2013;45:290–4.
Fatmi MQ, Chang CE. The role of oligomerization and cooperative regulation in protein function: the case of tryptophan synthase. PLoS Comput Biol. 2010;6:e1000994.
Hnizda A, Skerlova J, Fabry M, Pachl P, Sinalova M, Vrzal L, et al. Oligomeric interface modulation causes misregulation of purine 5 -nucleotidase in relapsed leukemia. BMC Biol. 2016;14:91.
Spychala J, Madrid-Marina V, Fox IH. High Km soluble 5’-nucleotidase from human placenta. Properties and allosteric regulation by IMP and ATP. J Biol Chem. 1988;263:18759–65.
Wallden K, Nordlund P. Structural basis for the allosteric regulation and substrate recognition of human cytosolic 5’-nucleotidase II. J Mol Biol. 2011;408:684–96.
Wallden K, Stenmark P, Nyman T, Flodin S, Graslund S, Loppnau P, et al. Crystal structure of human cytosolic 5’-nucleotidase II: insights into allosteric regulation and substrate recognition. J Biol Chem. 2007;282:17828–36.
Mueller U, Darowski N, Fuchs MR, Forster R, Hellmig M, Paithankar KS, et al. Facilities for macromolecular crystallography at the Helmholtz-Zentrum Berlin. J Synchrotron Radiat. 2012;19:442–9.
Fiser A, Sali A. ModLoop: automated modeling of loops in protein structures. Bioinformatics. 2003;19:2500–1.
Jubb HC, Higueruelo AP, Ochoa-Montano B, Pitt WR, Ascher DB, Blundell TL. Arpeggio: a web server for calculating and visualising interatomic interactions in protein structures. J Mol Biol. 2017;429:365–71.
Pires DE, Ascher DB, Blundell TL. DUET: a server for predicting effects of mutations on protein stability using an integrated computational approach. Nucleic Acids Res. 2014;42(Web Server issue):W314–319.
Pires DE, Ascher DB, Blundell TL. mCSM: predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics. 2014;30:335–42.
Rozbesky D, Sovova Z, Marcoux J, Man P, Ettrich R, Robinson CV, et al. Structural model of lymphocyte receptor NKR-P1C revealed by mass spectrometry and molecular modeling. Anal Chem. 2013;85:1597–604.
Zaliova M, Kotrova M, Bresolin S, Stuchly J, Stary J, Hrusak O, et al. ETV6/RUNX1-like acute lymphoblastic leukemia: a novel B-cell precursor leukemia subtype associated with the CD27/CD44 immunophenotype. Genes Chromosomes Cancer. 2017;56:608–16.
Kotrova M, Musilova A, Stuchly J, Fiser K, Starkova J, Mejstrikova E, et al. Distinct bilineal leukemia immunophenotypes are not genetically determined. Blood. 2016;128:2263–6.
Zikanova M, Skopova V, Hnizda A, Krijt J, Kmoch S. Biochemical and structural analysis of 14 mutant adsl enzyme complexes and correlation to phenotypic heterogeneity of adenylosuccinate lyase deficiency. Hum Mutat. 2010;31:445–55.
Yu B, Howell PL. Intragenic complementation and the structure and function of argininosuccinate lyase. Cell Mol Life Sci. 2000;57:1637–51.
Baker RP, Urban S. Cytosolic extensions directly regulate a rhomboid protease by modulating substrate gating. Nature. 2015;523:101–5.
Marton Z, Guillon R, Krimm I, Preeti, Rahimova R, Egron D, et al. Identification of noncompetitive inhibitors of cytosolic 5 ‘-nucleotidase II using a fragment-based approach. J Med Chem. 2015;58:9680–96.
Ma X, Edmonson M, Yergeau D, Muzny DM, Hampton OA, Rusch M, et al. Rise and fall of subclones from diagnosis to relapse in pediatric B-acute lymphoblastic leukaemia. Nat Commun. 2015;6:6604.
Kunz JB, Rausch T, Bandapalli OR, Eilers J, Pechanska P, Schuessele S, et al. Pediatric T-cell lymphoblastic leukemia evolves into relapse by clonal selection, acquisition of mutations and promoter hypomethylation. Haematologica. 2015;100:1442–50.
Richter-Pechanska P, Kunz JB, Hof J, Zimmermann M, Rausch T, Bandapalli OR, et al. Identification of a genetically defined ultra-high-risk group in relapsed pediatric T-lymphoblastic leukemia. Blood Cancer J. 2017;7:e523.
Ding LW, Sun QY, Mayakonda A, Tan KT, Chien W, Lin DC, et al. Mutational profiling of acute lymphoblastic leukemia with testicular relapse. J Hematol Oncol. 2017;10:65.
Reitman ZJ, Yan H. Isocitrate dehydrogenase 1 and 2 mutations in cancer: alterations at a crossroads of cellular metabolism. J Natl Cancer Inst. 2010;102:932–41.
Doerr A. Single-particle cryo-electron microscopy. Nat Methods. 2016;13:23.
Aster JC, DeAngelo DJ. Resistance revealed in acute lymphoblastic leukemia. Nat Med. 2013;19:264–5.
Tzoneva G, Dieck CL, Oshima K, Ambesi-Impiombato A, Sanchez-Martin M, Madubata CJ, et al. Clonal evolution mechanisms in NT5C2 mutant-relapsed acute lymphoblastic leukaemia. Nature. 2018;553:511–4.
Bricard G, Cadassou O, Cassagnes LE, Cros-Perrial E, Payen-Gay L, Puy JY, et al. The cytosolic 5’-nucleotidase cN-II lowers the adaptability to glucose deprivation in human breast cancer cells. Oncotarget. 2017;8:67380–93.
Acknowledgements
This work was supported by grant 15-06582S from the Czech Science Foundation and in part by the Ministry of Education of the Czech Republic (program “NPU I” LO1304 and program “InterBioMed” LO1302) and by European Regional Development Fund Project No. CZ.02.1.01/0.0/0.0/16_019/0000729. Institutional support was provided by projects RVO 61388963 and 68378050 of the Academy of Sciences of the Czech Republic. MZ and JT were funded by grant 15-30626A from Czech Health Research Council and grant Primus/MED/28 from Charles University. DBA was supported by a C. J. Martin Research Fellowship from the National Health and Medical Research Council of Australia (APP1072476) and the Jack Brockhoff Foundation (JBF 4186, 2016). We acknowledge beamline access at MX14.1 and MX14.3 of the BESSY (HZB Berlin, Germany) to collect crystal diffraction data. The authors would like to thank Irena Sieglova, MSc and Anna Soldanova, BSc for technical help during protein preparations and Katsiaryna Tratsiak, MSc for crystallization trials.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Hnízda, A., Fábry, M., Moriyama, T. et al. Relapsed acute lymphoblastic leukemia-specific mutations in NT5C2 cluster into hotspots driving intersubunit stimulation. Leukemia 32, 1393–1403 (2018). https://doi.org/10.1038/s41375-018-0073-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-018-0073-5