Abstract
Single-cell genetics were used to interrogate clonal complexity and the sequence of mutational events in STIL-TAL1+ T-ALL. Single-cell multicolour FISH was used to demonstrate that the earliest detectable leukaemia subclone contained the STIL-TAL1 fusion and copy number loss of 9p21.3 (CDKN2A/CDKN2B locus), with other copy number alterations including loss of PTEN occurring as secondary subclonal events. In three cases, multiplex qPCR and phylogenetic analysis were used to produce branching evolutionary trees recapitulating the snapshot history of T-ALL evolution in this leukaemia subtype, which confirmed that mutations in key T-ALL drivers, including NOTCH1 and PTEN, were subclonal and reiterative in distinct subclones. Xenografting confirmed that self-renewing or propagating cells were genetically diverse. These data suggest that the STIL-TAL1 fusion is a likely founder or truncal event. Therapies targeting the TAL1 auto-regulatory complex are worthy of further investigation in T-ALL.
Similar content being viewed by others
Introduction
Single-cell genetics in haematopoietic [1,2,3] and other cancers [4, 5] have revealed substantial intraclonal complexity. In general, this diversity reflects evolutionary phylogenies with derivative subclones branching off from founder precursors [6]. Architectural population diversity in cancer has important implications for reservoirs of cells involved in progression of disease and drug resistance therapy. Bioinformatic derivations of evolutionary trees can reveal the most likely sequence of genetic events and distinguish mutations that are present in all cancer cells, as truncal or founder events, versus those that are secondary and subclonally distributed [7,8,9]. This in turn carries implications for minimal residual disease (MRD) monitoring and targeted therapy.
Few such studies have been performed to date in T-ALL, although comparative genetic profiling of diagnostic, xenograft and relapse samples confirms clonal complexity [10, 11]. T-ALL is biologically diverse reflecting levels of differentiation arrest within the thymus and distinctive genetic lesions [12]. We elected to study a single, common subtype of T-ALL, namely those with STIL-TAL1 fusion. We used multicolour FISH and single-cell multiplex quantitative-PCR (qPCR) to determine the phylogenetic architecture of diagnostic samples and to infer the order of genetic events comparing STIL-TAL1 fusion, which we postulated as a founder lesion, with other common genetic lesions including CDKN2A loss, PTEN mutation or loss and NOTCH1 mutation. In selected cases, we compared the clonal architecture of xenotransplanted samples with that observed in the diagnostic sample. This enabled us to infer the subclonal origins and genetic diversity of cells with propagating or stem cell activity.
Materials and methods
Patient samples
Diagnostic DNA of 19 T-ALL cases aged 1–24 years and 1 cell line (RPMI 8402) known to have the STIL-TAL1 rearrangement were available. The study was conducted in accordance with the Declaration of Helsinki and appropriate consent and ethical approval for the study was obtained (Ethics approval numbers CCR2285 and 16/SE/0219).
T-ALL molecular screening and cloning
Diagnostic DNA from all STIL-TAL1 cases was analysed for mutations in known T-ALL mutational hotspots in NOTCH1 (exons 26, 27 and 34), FBXW7 (exons 9 and 10), PTEN (exon 7) and IL7R (exon 6) using previously published methods [13,14,15,16]. All diagnostic samples were analysed by SNP-array to identify genomic losses and gains using the Affymetrix SNP 6.0 platform. Genotyping and generation of QC data were performed in Genotyping ConsoleTM v4.1.4 software (Affymetrix). CNAG version 3.3.0.1 beta was used to normalise output to a self-reference (patient remission DNA) or via a batch pairwise analysis using sex-matched control samples. The STIL-TAL1 patient-specific gene fusion was sequenced for the three cases that underwent single-cell genotyping analyses using previously published methods [17]. The TA Cloning Kit® (Invitrogen by Life TechnologiesTM) was used for cloning experiments according to the manufacturer’s instructions.
Next-generation sequencing (NGS)
Whole exome sequencing (WES) was undertaken by Oxford Gene Technology. See Supplementary Methods for details and bioinformatics. Any genomic ‘drivers’ included in the single-cell genotyping experiments were validated with Sanger sequencing using custom primers designed using Primer Blast (Table S1).
Fluorescence in situ hybridisation
Fixed cytospins were prepared from archived viable cells and interphase FISH was performed with patient-specific FISH probes for the various copy number losses using in-house FISH probes (Table S2) and previously described methods [18]. See Supplementary Methods for details.
Bioinformatic assessment of RAG recombinase activity at PTEN indel breakpoints
MEME Suite 4.11.4 [19] was used to conduct an agnostic search for the RAG recombinase consensus heptamer and nonamer and also for the tetramer sequence (CACA) identified [20] as being recurrently present at RAG-mediated breakpoint sequences. We also used a weighted matrix algorithm (code availability—script kindly provided by the laboratory of Dr Papaemmanuil) to generate RAG recombination signal sequences (RSS) scores for each deletion breakpoint of interest. Essentially, this ascribed a likelihood score or weight to each base pair in the putative heptamer-spacer-nonamer sequence of interest according to the likelihood of deviation from consensus based on what the base pair is for the heptamer/nonamer and the number of bases rather than base choice per se for the spacer (score details outlined in ref. [21]).
Single-cell genotyping and single-cell Sanger sequencing
Single-cell genetic analysis was performed using stored viable cells for cases CUL76, 6116 and 6030 and paired xenograft material. Our previously established multiplex qPCR approach was used [2] with minor modifications. See Supplementary Methods for details.
Xenograft material
Limited archived xenograft DNA and single-cell material prepared from xenograft bone marrow was available on samples CUL76, 6030 and 6116. Material generated using NOG (NOG-ShiSCID-IL2gamma null) mice was available for samples 6030 and 6116 and using NRG (NOD-Rag1nullIL2rgnull, NOD rag gamma) mice for sample CUL76. In all cases, 1 × 106 cells were injected and experiments were performed on material stored from primary passage bone marrow.
Results
Molecular screening on STIL-TAL1+ T-ALL samples for recurrent drivers
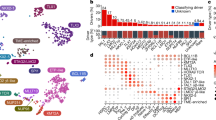
Nineteen STIL-TAL1+ cases and one cell line were screened for common genetic rearrangements using copy number profiling and Sanger sequencing. Data on Brazilian samples BR75, BR74, S1 and S2 have been previously published [22, 23]. Four cases also underwent WES to identify both known driver targets for inclusion in single-cell experiments and novel drivers. Results of molecular screening are summarised in Table 1 (further details available from Tables S8, S9 and S10). Mean target coverage across the samples (and paired remission samples) that underwent WES was 95×–126× (average 109×). Total of 7–10 protein altering SNVs and 3–5 protein altering indels (detectable at a read depth >20) were detected per sample (Tables S11 and S12). All drivers incorporated in to single-cell experiments were validated with qPCR or Sanger sequencing.
In keeping with previous work highlighting that cases from the TAL/LMO gene expression subgroup tend to have a higher incidence of PTEN mutations and a lower incidence of NOTCH1 mutations [24], within our cohort of 20 patients (combining PTEN inactivation due to exon 7 mutation or copy number loss) we detected a frequency of PTEN inactivation of at least 40% (around double the frequency of PTEN inactivation (22%) detected in a recent study of 145 T-ALL cases using heteroduplex analysis, mutations and SNP arrays) [25]. Additionally in the cases that underwent WES, mutations in PTEN outside the hotspot of exon 7 were detected (exon 8 in sample 6030 and exon 5 in CF10), suggesting that estimates based on copy number analysis and PTEN exon 7 sequencing may be an underestimate. A high frequency of copy number losses of 9p (95%, incorporating CDKN2A/CDKN2B locus) and 6q (30%) was also observed and these were identified as key drivers to include in single-cell genotyping studies.
Exome sequencing of four cases defined relevant known and potential T-ALL drivers to include in single-cell analysis experiments. Novel T-ALL potential drivers were detected: mutations in BMPR1A, FREM2 and PIK3CD. BMPR1A (bone morphogenetic protein receptor, type 1A) is a polyposis associated gene—a type 1A transmembrane serine/threonine kinase listed in the cosmic cancer gene census of genes functionally linked to cancer and PIK3CD is a subunit of phosphatidylinositol-4,5-bisphosphate 3-kinase a key member of the PTEN-PI3 kinase pathway. Both 6030 and 6116 contained one mutation per sample in FREM2 and one of these mutations (found re-iteratively in sample 6116) is found in the cosmic database as a somatic mutation in a GI carcinoma (COSM287123). Additionally, this gene was recurrently but non-significantly mutated in three T-ALL samples in a large exome T-ALL sequencing study [26]. The gene encodes an integral membrane protein containing many chondroitin sulphate proteoglycan element repeats and calx-beta domains (although the identified mutations lie outside these domains, Figure S3). In view of the possible association with T-ALL, and the emerging role of the micro-environment/extracellular matrix in leukaemia and cancer, we included FREM2 mutations in our single-cell studies.
Reiterative inactivation of PTEN
Sanger sequencing and NGS data for samples 6030 and CF5 suggested the presence of multiple low-level PTEN exon 7 indels running in parallel (Fig. 1a, b). Parallel work by a collaborating group using HPLC wave technology (heteroduplex analysis) [25] also led to similar conclusions. Cloning experiments validated at least four independent indels for sample 6030. Sample CF10 also had two PTEN indels in exons 5 and 7 (detected by WES and Sanger sequencing). The multiple PTEN indels resulted functionally in the generation of stop codons in all but one of the mutations analysed (10/11; Table S13). Given the striking reiterative inactivation of PTEN, we hypothesised that these small structural alterations could be RAG-mediated given the involvement of aberrant RAG in the formation of the STIL-TAL1 fusion and CDKN2A deletions and the observation that RSS elsewhere in PTEN have been implicated in the formation of small microdeletions [27]. However, we did not find bioinformatic evidence to support this explanation with maximum RSS scores of 6.75 (Table S14) compared to >8.55 in B cell precursor (BCP)-ALL samples using the same weighted matrix algorithm [20]. Only one sample (CF10) had the RAG-associated CACA tetramer identified in Papaemmanuil et al. (2014) close to the mutation breakpoint.
Reiterative bi-allelic deletion of PTEN. a Multiplex qPCR data show that PTEN indels 2 and 3 are localised within the same single cells. The coloured boxes at the junction between the vertical qPCR assays (run in duplicates) and the horizontal individual cells represent mutation-specific amplification with the colour of each box correlating with the level of amplification (raw Ct value). Black represents no amplification. b Reiterative PTEN exon 7 indel mutations are seen within the raw data reads of whole exome sequencing data for sample 6030. c Cloning of PTEN exon 7 from representative single cells demonstrated that these indels are present on separate alleles as one single-cell clone contains indel 2 but is wild-type (WT) for indel 3, whereas another single-cell clone contains indel 3 but is WT for indel 2 demonstrating bi-allelic inactivation of PTEN. Cell number 17 refers to experimental single-cell DNA plate position
Single-cell studies in STIL-TAL1+ T-ALL
Multicolour FISH
We investigated the order of acquisition of CNAs with reference to the STIL-TAL1 fusion in samples 6030, CF5, CF6 and HK328. In the majority of cases, the earliest ancestor subclone contained the STIL-TAL1 fusion in combination with bi-allelic loss of 9p21.3 (containing CDKN2A locus) as shown for samples CF5 and CF6 (Fig. 2). In contrast with BCP-ALL [9] in the majority of cases 9p21.3 could not be separated in time from the presumed founder gene fusion/translocation. Additional copy number losses including 6q and PTEN occurred in a secondary and subclonal fashion. However, in sample 6030 the earliest detectable subclone contained STIL-TAL1+ cells with 1 copy of 9p21.3. This observation was validated using two independent FISH probes and the clone was also detected in xenograft bone marrow derived from sample 6030 (Figure S1a).
Multicolour FISH in STIL-TAL1 T-ALL. Examples for evolutionary trees for samples CF5 and CF6 are shown. Percentages represent the frequency of each subclone as assessed by single-cell multicolour FISH with circle size representing relative frequencies. Photos shown are from sample CF5 using the following FISH probes: STIL-TAL1 (labelled spectrum red-spectrum green); p16.1 as a marker of CDKN2A loss (labelled Cy3, coloured blue in diagram); PTEN (labelled Biotin-Cy5, coloured pink in diagram). Note the STIL-TAL1 FISH probe is a subdeletion probe. Normal cells have two co-localised red-green signals. Cells with the STIL-TAL1 fusion due to 1p33 deletion lose one red signal while retaining the control green signal. ST F: STIL-TAL1 Fusion. Numbers (0, 1, 2) next to gene name (PTEN or CDKN2A) represent copy number results by FISH, e.g., 2 PTEN, two copies of PTEN, 0 CDKN2A, zero copies of CDKN2A
Single-cell multiplex qPCR
Although multicolour FISH provides proof-of-principle evidence for clonal heterogeneity in T-ALL, the number of driver genetic events that can be examined is limited and mutations in key T-ALL signalling pathways cannot be incorporated. We undertook a detailed single-cell genotyping study on three samples that represented all the key STIL-TAL1 genetic driver events identified in this study and which had archived paired xenograft material available. Archived diagnostic leukaemia cells from cases 6030, 6116 and CUL76 underwent high-throughput single-cell multiplex qPCR analysis allowing simultaneous investigation of the patient-specific STIL-TAL1 gene fusion in each case along with indels, SNVs and CNA designated driver status for each leukaemia sample. Quality control assessments and analysis were performed as previously described [2].
Phylogenetic trees for these three diagnostic cases constructed from single-cell data using the principle of maximum parsimony are shown in Fig. 3a–c. In all cases the root of the tree contains a common ancestor cell containing the patient-specific STIL-TAL1 fusion and copy number loss of CDKN2A. Other key drivers are subclonal but in some cases re-iterative in keeping with observations made in prior studies of B cell leukaemias and solid tumours [2, 9, 28,29,30]. The STIL-TAL1 assays used were specific for the patient-specific fusions (Table S15) and tested against both normal DNA and other STIL-TAL1 positive material to confirm assay specificity. This study provides the first evidence that a single STIL-TAL1 fusion occurs per case (previous studies assessing stability through xenograft passage have used chromosome 1p copy number as a surrogate for the gene fusion but this would not differentiate the presence of multiple overlapping fusions with relatively conserved mutation breakpoint region). NOTCH1 mutation and PTEN mutation are secondary and subclonal events in the case studies. The multiple PTEN mutations noted in sample 6030 were tracked at single-cell level and single-cell cloning and Sanger sequencing were used to demonstrate unequivocally that in clones with >1 mutation, mutations occurred on independent alleles suggesting a selective pressure for bi-allelic inactivation of PTEN (Fig. 1c).
a–c Single-cell multiplex qPCR evolutionary analysis samples 6030, 6116 and CUL76. Evolutionary trees generated by bioinformatics analysis of single-cell data are shown. Gene names of drivers tracked are shown next to the subclones. Presence of a gene name indicates gene mutation present in the case of SNVs or indels. Where the driver is a deletion number of copies of gene present (0 or 1) is shown. Yellow circles represent leukaemic subclones and a black circle and the N label indicates the normal state. The size of the circle is proportional to the number of cells in each subclone and the detected genetic markers are listed below each circle. Red T = subclones that read-out in xenotransplants (detailed xenograft single-cell data in Supplementary Table 16). Grey boxes represent inferred subclones; these are groups of cells, which have died out, been outcompeted or if still present, exist at low frequencies below the level of reliable detection using this approach. Tree branch lengths are directly proportional to the number of evolutionary changes inferred and the points at which the branches diverge (nodes) represent the ancestor state of a clonal clade; a monophyletic group, which includes all descendants of the ancestor. The number in grey colour at each node indicates the bootstrap value. The phylogeny shows how the clonal expansion has evolved from a common ancestor toward the observed states. Note that for case CUL76 as a limited number of drivers were tracked based on mutation screening and copy number data another known mutation present in this sample (SORC3, passenger mutation) was included to aid assessment of clonal structure. a Case 6116 (N = 308 cells). The root of the tree harbours the STIL-TAL1 fusion (F) and a homozygous CDKN2A deletion. Two equally parsimonious trees (I and II) are generated which differ as to whether the FREM2 or PIK3CD mutated clone gives rise to the latest detectable C4 clone. The NOTCH1 mutation is a subclonal event. b CUL76 (N = 151 cells). The root of the tree harbours the STIL-TAL1 F and a homozygous CDKN2A deletion and a LMO2 deletion in this case. PTEN inactivation, through copy number loss, and NOTCH1 mutation are subclonal. The NOTCH1 TAD domain exon 34 mutation was only detected in clone C2 (NOTCH1 mutation status determined by single-cell Sanger sequencing of 44 single cells). The pie chart shows the proportions of C2 subclone cells that are either wild-type (WT) or NOTCH1 mutated. c 6030 (N = 216 cells). The root of the tree harbours the STIL-TAL1 F and a homozygous CDKN2A deletion. PTEN mutations are subclonal and re-iterative (3 PTEN indels in exon 7—labelled PTEN indels 1, 2 and 3 plus one PTEN indel in exon 8). The C6 subclone contains two independent PTEN exon 7 mutations. PTEN exon 8 mutation was only detected in clone C7 (PTEN exon 8 mutation status determined by single-cell Sanger sequencing of 37 single cells). The pie chart shows the proportions of C7 subclone cells that are WT, heterozygous PTEN exon mutated or homozygous PTEN exon 8 mutated
The use of xenograft models can be used to assess clonal heterogeneity and evolution in the leukaemia initiating or so called ‘cancer stem-cell’ compartment providing insight in to genetic driver stability and evolutionary pressures on clonal selection. Single-cell assessment of xenograft bone marrow in the three cases examined demonstrated contrasting models of clonal evolution, although it is acknowledged that limited conclusions can be drawn from the single xenograft per case experiments performed. Clones that read-out in the xenograft are designated ‘T’ status in the evolutionary tree diagrams (Fig. 3a–c). In cases 6116 and CUL76, multiple clones read out in the xenograft confirming the position of NOTCH1 as a subclonal driver and demonstrating multiple competing subclones in the leukaemia initiating cell compartment. However, in case 6030 a single dominant clone (C7) predominated containing the STIL-TAL1 fusion, bi-allelic 9p21.3 deletion and a PTEN exon 8 mutation, which was present in both heterozygous and homozygous form. The proportion of cells within this subclone with a homozygous (as opposed to heterozygous) PTEN exon 8 mutation was higher in the xenograft than the diagnostic sample (88% versus 47%) and all xenograft single cells had either the heterozygous or homozygous mutation (i.e., no PTEN exon 8 wild-type cells detected based on Sanger sequencing of 78 single cells). The clones with PTEN exon 7 mutations were not detected in the xenograft. This validated bulk DNA sequencing data from the xenograft material which demonstrated the homozygous PTEN exon 8 mutation and wild-type PTEN exon 7 analysis (data not shown).
Comparison of 6030 xenograft single-cell FISH and multiplex qPCR data initially appeared to generate conflicting results with FISH data suggesting the presence of a clone with 1 copy of 9p21.3 in both diagnostic and xenograft material but single-cell data demonstrating bi-allelic loss in all subclones. Examination of SNP copy number data resolved this discrepancy, which was due to one of the 9p21.3 deletions being smaller than the size of the FISH probe so that the FISH subclone containing ‘1 copy’ of 9p21.3, 2 copies of 4p/6q (Figure S1a) actually corresponded to clone C7 in multiplex qPCR data (Fig. 3c), i.e., CDKN2A actually demonstrated bi-allelic deletion when small qPCR assays were used to assess copy number status. Examination of paired diagnosis/xenograft copy number data for case 6030 also demonstrated that CDKN2A loss is a subclonal event as with regard to the loss of the second allele of CDKN2A at least two independent clones had emerged with distinct breakpoints (Figure S1b). Since multiplex qPCR data for CDKN2A was based on copy number assays, it is possible that multiple CDKN2A deletions may exist in any one sample, which would not be detected using the analysis method used.
Discussion
These single-cell genetic analyses allow us to infer phylogenetic trees describing clonal evolution. Technical limitations mean that interpretation of these data carries the caveat that we will have under-estimated clonal complexity. Our genetic markers are also limited. More genetically distinct subclones will exist than we currently detect. For case 6116 due to the limited genetic markers used we were unable to determine whether the re-iterative mutation event was PIK3CD or FREM2 mutation but did validate the bioinformatic analysis with single-cell Sanger sequencing analysis of FREM2 and PIK3CD mutation in single cells of the diagnostic and xenograft samples to confirm that re-iterative mutation had occurred (Figure S2).
Despite these caveats, several informative conclusions can be drawn. A consistent feature is that STIL-TAL1 fusion and CDKN2A loss are both early or truncal events, in contrast to other recurrent genetic changes including NOTCH1 and PTEN mutation that are secondary and subclonal. These observations have implications for selection of mutations for minimal residual disease tracking and as targets for therapy. Given the position of the STIL-TAL1 fusion in leukaemia evolution, therapies targeting the TAL1 regulatory complex [31] are worthy of further investigation.
It is difficult to discern from our study whether STIL-TAL1 or CDKN2A loss is an initiating event, or which comes first. The most ancestral cell in the phylogenetic structure has STIL-TAL1 fusion plus loss of both CDKN2A alleles. However, the latter are distinctive and presumed independent events, but only one clone-specific STIL-TAL1 fusion exists. This finding suggests that at least one of the CDKN2A allele deletions occurs subsequent to STIL-TAL1 fusion. This is in keeping with previously published work demonstrating reiterative CDKN2A deletions in STIL-TAL1 cases when breakpoints of 9p21.3 deletions in paired diagnostic/xenograft/relapse material were examined [11]. We note that both STIL-TAL1 fusion and CDKN2A loss are likely to involve ‘off target’ RAG-dependent mutational mechanisms.
The clonal architectures in STIL-TAL1+ ALL cases (Fig. 3) for which we had genome sequencing data showed branching structure as previously described in B cell precursor ALL [9]. To some extent, this is driven by reiterative mutations of the same driver genes and resultant parallel clonal evolution. This is evident with several driver genes but most clearly with PTEN in patient 6030 (Fig. 3c). Cloning these multiple mutations from single cells confirmed their uniqueness (Fig. 1c). In BCP-ALL, reiterative copy number changes (e.g., in ETV6, PAX5, CDKN2A or BTG1) are the consequence of RAG-mediated mutation, followed by selection [9, 20, 30]. We found no bioinformatics support for RAG involvement in the PTEN mutations we identified. We note, however, that RSS for RAGs have been previously implicated in PTEN small, microdeletions [27]. Irrespective of the mutational mechanisms involved in these PTEN mutations, we conclude that in STIL-TAL1 ALL there is a strong selective pressure for these genetic lesions, most likely related to epistasis or a strong functional complementarity between PTEN inactivation/loss, STIL-TAL1 fusion and CDKN2A loss.
The objective, in our limited xenograft studies, was to determine the subclonal origin and genetic diversity of propagating or self-renewing cells in STIL-TAL1+ ALL. The data indicate that multiple subclones read out in the mice reflecting, we suggest, the existence of genetically diverse stem cells. In patient 6116 (Fig. 3a), all four diagnostic subclones read out in the transplants and in patient CUL76 (Fig. 3b) all three subclones. Patient 6030 gave a different result however. One dominant subclone (subclone C7; Fig. 3c) read-out of the seven that existed in the diagnostic sample. However, the diagnostic subclone C7 was heterogeneous with respect to PTEN exon 8 status and this directly was also reflected in the transplant read-outs, indicating that two small subclones of clone C7 had propagated in the mice, providing further evidence for selection favouring PTEN-mutated clones. Genetically diverse propagating or stem cells were similarly demonstrated in B cell precursor ALL [9, 32] and in glioblastoma [33] and are likely to be a common feature in cancer [34, 35]. The implication of this observation is that multiple subclones harbour the proliferative potential to fuel progression of disease, relapse and drug resistance.
References
Paguirigan AL, Smith J, Meshinchi S, Carroll M, Maley C, Radich JP. Single-cell genotyping demonstrates complex clonal diversity in acute myeloid leukemia. Sci Transl Med. 2015;7:281re282.
Potter NE, Ermini L, Papaemmanuil E, Cazzaniga G, Vijayaraghavan G, Titley I, et al. Single-cell mutational profiling and clonal phylogeny in cancer. Genome Res. 2013;23:2115–25.
Gawad C, Koh W, Quake SR. Dissecting the clonal origins of childhood acute lymphoblastic leukemia by single-cell genomics. Proc Natl Acad Sci USA. 2014;111:17947–52.
Muller S, Liu SJ, Di Lullo E, Malatesta M, Pollen AA, Nowakowski TJ, et al. Single-cell sequencing maps gene expression to mutational phylogenies in PDGF- and EGF-driven gliomas. Mol Syst Biol. 2016;12:889.
Leung ML, Davis A, Gao R, Casasent A, Wang Y, Sei E, et al. Single-cell DNA sequencing reveals a late-dissemination model in metastatic colorectal cancer. Genome Res. 2017;27:1287–1299.
Greaves M, Maley CC. Clonal evolution in cancer. Nature. 2012;481:306–13.
Hiley C, de Bruin EC, McGranahan N, Swanton C. Deciphering intratumor heterogeneity and temporal acquisition of driver events to refine precision medicine. Genome Biol. 2014;15:453.
Brastianos PK, Carter SL, Santagata S, Cahill DP, Taylor-Weiner A, Jones RT, et al. Genomic characterization of brain metastases reveals branched evolution and potential therapeutic targets. Cancer Discov. 2015;5:1164–77.
Anderson K, Lutz C, van Delft FW, Bateman CM, Guo Y, Colman SM, et al. Genetic variegation of clonal architecture and propagating cells in leukaemia. Nature. 2011;469:356–61.
Mullighan CG, Phillips LA, Su X, Ma J, Miller CB, Shurtleff SA, et al. Genomic analysis of the clonal origins of relapsed acute lymphoblastic leukemia. Science. 2008;322:1377–80.
Clappier E, Gerby B, Sigaux F, Delord M, Touzri F, Hernandez L, et al. Clonal selection in xenografted human T cell acute lymphoblastic leukemia recapitulates gain of malignancy at relapse. J Exp Med. 2011;208:653–61.
Belver L, Ferrando A. The genetics and mechanisms of T cell acute lymphoblastic leukaemia. Nat Rev Cancer. 2016;16:494–507.
Weng AP, Ferrando AA, Lee W, Morris JPT, Silverman LB, Sanchez-Irizarry C, et al. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science. 2004;306:269–71.
Kox C, Zimmermann M, Stanulla M, Leible S, Schrappe M, Ludwig WD, et al. The favorable effect of activating NOTCH1 receptor mutations on long-term outcome in T-ALL patients treated on the ALL-BFM 2000 protocol can be separated from FBXW7 loss of function. Leukemia. 2010;24:2005–13.
Jotta PY, Ganazza MA, Silva A, Viana MB, da Silva MJ, Zambaldi LJ, et al. Negative prognostic impact of PTEN mutation in pediatric T-cell acute lymphoblastic leukemia. Leukemia. 2010;24:239–42.
Zenatti PP, Ribeiro D, Li W, Zuurbier L, Silva MC, Paganin M, et al. Oncogenic IL7R gain-of-function mutations in childhood T-cell acute lymphoblastic leukemia. Nat Genet. 2011;43:932–9.
Breit TM, Mol EJ, Wolvers-Tettero IL, Ludwig WD, van Wering ER, van Dongen JJ. Site-specific deletions involving the tal-1 and sil genes are restricted to cells of the T cell receptor alpha/beta lineage: T cell receptor delta gene deletion mechanism affects multiple genes. J Exp Med. 1993;177:965–77.
Kearney LACS. Specialised fluorescence in situ hybridization (FISH) techniques for leukaemia research. In: Kearney L, Colman S (eds). Leukaemia methods and protocols, vol. 1. New York: Springer; 2009. p. 57–70.
McLaren W, Gil L, Hunt SE, Riat HS, Ritchie GR, Thormann A, et al. The ensembl variant effect predictor. Genome Biol. 2016;17:122.
Papaemmanuil E, Rapado I, Li Y, Potter NE, Wedge DC, Tubio J, et al. RAG-mediated recombination is the predominant driver of oncogenic rearrangement in ETV6-RUNX1 acute lymphoblastic leukemia. Nat Genet. 2014;46:116–25.
Hesse JE, Lieber MR, Mizuuchi K, Gellert M. V(D)J recombination: a functional definition of the joining signals. Genes Dev. 1989;3:1053–61.
Mansur MB, Ford AM, van Delft FW, Gonzalez D, Emerenciano M, Maia RC, et al. Occurrence of identical NOTCH1 mutation in non-twinned sisters with T-cell acute lymphoblastic leukemia. Leukemia. 2011;25:1368–70.
Mansur MB, Hassan R, Barbosa TC, Splendore A, Jotta PY, Yunes JA, et al. Impact of complex NOTCH1 mutations on survival in paediatric T-cell leukaemia. BMC Cancer. 2012;12:9.
Zuurbier L, Petricoin EF 3rd, Vuerhard MJ, Calvert V, Kooi C, Buijs-Gladdines JG, et al. The significance of PTEN and AKT aberrations in pediatric T-cell acute lymphoblastic leukemia. Haematologica. 2012;97:1405–13.
Jenkinson S, Kirkwood AA, Goulden N, Vora A, Linch DC, Gale RE. Impact of PTEN abnormalities on outcome in pediatric patients with T-cell acute lymphoblastic leukemia treated on the MRC UKALL2003 trial. Leukemia. 2016;30:39–47.
De Keersmaecker K, Atak ZK, Li N, Vicente C, Patchett S, Girardi T, et al. Exome sequencing identifies mutation in CNOT3 and ribosomal genes RPL5 and RPL10 in T-cell acute lymphoblastic leukemia. Nat Genet. 2013;45:186–90.
Mendes RD, Sarmento LM, Cante-Barrett K, Zuurbier L, Buijs-Gladdines JG, Povoa V, et al. PTEN microdeletions in T-cell acute lymphoblastic leukemia are caused by illegitimate RAG-mediated recombination events. Blood. 2014;124:567–78.
Gerlinger M, Horswell S, Larkin J, Rowan AJ, Salm MP, Varela I, et al. Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat Genet. 2014;46:225–33.
Shlush LI, Zandi S, Mitchell A, Chen WC, Brandwein JM, Gupta V, et al. Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature. 2014;506:328–33.
Waanders E, Scheijen B, van der Meer LT, van Reijmersdal SV, van Emst L, Kroeze Y, et al. The origin and nature of tightly clustered BTG1 deletions in precursor B-cell acute lymphoblastic leukemia support a model of multiclonal evolution. PLoS Genet. 2012;8:e1002533.
Sanda T, Lawton LN, Barrasa MI, Fan ZP, Kohlhammer H, Gutierrez A, et al. Core transcriptional regulatory circuit controlled by the TAL1 complex in human T cell acute lymphoblastic leukemia. Cancer Cell. 2012;22:209–21.
Notta F, Mullighan CG, Wang JC, Poeppl A, Doulatov S, Phillips LA, et al. Evolution of human BCR-ABL1 lymphoblastic leukaemia-initiating cells. Nature. 2011;469:362–7.
Piccirillo SG, Colman S, Potter NE, van Delft FW, Lillis S, Carnicer MJ, et al. Genetic and functional diversity of propagating cells in glioblastoma. Stem Cell Rep. 2015;4:7–15.
Greaves M. Evolutionary determinants of cancer. Cancer Discov. 2015;5:806–20.
Kreso A, Dick JE. Evolution of the cancer stem cell model. Cell Stem Cell. 2014;14:275–91.
Acknowledgements
We thank Mrs Susan M Colman for assistance with FISH protocols; Dr Claire Schwab for identification of STIL-TAL1 samples and Ms Tracey Perry for assistance with in vivo experiments. We acknowledge the ECOG group for the use of the sample 21922 and the Bloodwise Childhood Leukaemia Cell Bank for provision of primary haematological malignancy samples used in this study. We thank Dr Elli Papaemmanuil for the use of the weighted matrix algorithm for assessment of RAG recombinase activity.
CLF was supported by a clinical research training fellowship from Bloodwise (formerly, Leukaemia & Lymphoma Research) (grant number 11035). MBM was supported by the Partner Fellowship (European Haematology Association #2011/01); by the International Award for Research in Leukaemia (Lady Tata Memorial Trust) and by the Ministry of Health (INCA-Brazil). MSM was supported by Rally Foundation fellowship. AAF was supported by NIH (grant R35 CA210065) and Leukemia and Lymphoma Society (R0749-14). MG is supported by a Wellcome Trust award [105104/Z/14/Z] to the Centre for Evolution and Cancer.
Author contributions
CLF co-designed the study, conducted, supervised and analysed all the experiments and wrote the paper; MBM performed molecular investigations and analysed data; AMF, SJ and RG assisted with molecular investigations and analyses; FWvD assisted with SNP procedures and analyses; VJW and PK conducted in vivo experiments; MSM performed characterisation and selection of samples; AAF provided stored xenograft material for sample CUL76; LE performed the bioinformatics analyses; CJH, MSPO, AAF and PK provided clinical samples and immunophenotypic/cytogenetic and/or clinical data; IT supervised all single-cell sorting; NEP supervised all single-cell experiments and phylogenetic analysis; MG co-designed and supervised the study and co-wrote the paper. All authors critically reviewed and approved the final draft of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Furness, C.L., Mansur, M.B., Weston, V.J. et al. The subclonal complexity of STIL-TAL1+ T-cell acute lymphoblastic leukaemia. Leukemia 32, 1984–1993 (2018). https://doi.org/10.1038/s41375-018-0046-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-018-0046-8
This article is cited by
-
Unlocking the Complexity: Exploration of Acute Lymphoblastic Leukemia at the Single Cell Level
Molecular Diagnosis & Therapy (2024)
-
Generating in vitro models of NTRK-fusion mesenchymal neoplasia as tools for investigating kinase oncogenic activation and response to targeted therapy
Oncogenesis (2023)
-
Transitioning single-cell genomics into the clinic
Nature Reviews Genetics (2023)
-
Targeting the NOTCH1-MYC-CD44 axis in leukemia-initiating cells in T-ALL
Leukemia (2022)
-
Phosphoproteomic profiling of T cell acute lymphoblastic leukemia reveals targetable kinases and combination treatment strategies
Nature Communications (2022)