Abstract
Background
Several studies have reported DNA methylation in blood to be associated with body mass index (BMI), but few have investigated causal aspects of the association. We used a twin family design to assess this association at two life points and applied a novel analytical approach to appraise the evidence for causality.
Methods
The methylation profile of DNA from peripheral blood was measured for 479 Australian women from 130 twin families. Linear regression was used to estimate the associations of DNA methylation at ~410,000 cytosine-guanine dinucleotides (CpGs), and of the average DNA methylation at ~20,000 genes, with current BMI, BMI at age 18–21 years, and the change between the two (BMI change). A novel regression-based methodology for twins, Inference about Causation through Examination of Familial Confounding (ICE FALCON), was used to assess causation.
Results
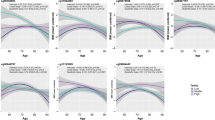
At a 5% false discovery rate, nine, six and 12 CpGs at 24 loci were associated with current BMI, BMI at age 18–21 years and BMI change, respectively. The average DNA methylation of the BHLHE40 and SOCS3 loci was associated with current BMI, and of the PHGDH locus with BMI change. From the ICE FALCON analyses with BMI as the predictor and DNA methylation as the outcome, a woman’s DNA methylation level was associated with her co-twin’s BMI, and the association disappeared after conditioning on her own BMI, consistent with BMI causing DNA methylation. To the contrary, using DNA methylation as the predictor and BMI as the outcome, a woman’s BMI was not associated with her co-twin’s DNA methylation level, consistent with DNA methylation not causing BMI.
Conclusion
For middle-aged women, peripheral blood DNA methylation at several genomic locations is associated with current BMI, BMI at age 18–21 years and BMI change. Our study suggests that BMI has a causal effect on peripheral blood DNA methylation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Petronis A. Epigenetics as a unifying principle in the aetiology of complex traits and diseases. Nature. 2010;465:721–7.
Esteller M. Epigenetics in cancer. N Engl J Med. 2008;358:1148–59.
Wang X, Zhu H, Snieder H, Su S, Munn D, Harshfield G, et al. Obesity related methylation changes in DNA of peripheral blood leukocytes. BMC Med. 2010;8:87.
Kuehnen P, Mischke M, Wiegand S, Sers C, Horsthemke B, Lau S, et al. An Alu element-associated hypermethylation variant of the POMC gene is associated with childhood obesity. PLoS Genet. 2012;8:e1002543.
Ding X, Zheng D, Fan C, Liu Z, Dong H, Lu Y, et al. Genome-wide screen of DNA methylation identifies novel markers in childhood obesity. Gene. 2015;566:74–83.
Almen MS, Nilsson EK, Jacobsson JA, Kalnina I, Klovins J, Fredriksson R, et al. Genome-wide analysis reveals DNA methylation markers that vary with both age and obesity. Gene. 2014;548:61–67.
Su S, Zhu H, Xu X, Wang X, Dong Y, Kapuku G, et al. DNA methylation of the LY86 gene is associated with obesity, insulin resistance, and inflammation. Twin Res Hum Genet. 2014;17:183–91.
Dick KJ, Nelson CP, Tsaprouni L, Sandling JK, Aissi D, Wahl S, et al. DNA methylation and body-mass index: a genome-wide analysis. Lancet. 2014;383:1990–8.
Aslibekyan S, Demerath EW, Mendelson M, Zhi D, Guan W, Liang L, et al. Epigenome-wide study identifies novel methylation loci associated with body mass index and waist circumference. Obesity. 2015;23:1493–501.
Demerath EW, Guan W, Grove ML, Aslibekyan S, Mendelson M, Zhou YH, et al. Epigenome-wide association study (EWAS) of BMI, BMI change and waist circumference in African American adults identifies multiple replicated loci. Hum Mol Genet. 2015;24:4464–79.
Al Muftah WA, Al-Shafai M, Zaghlool SB, Visconti A, Tsai PC, Kumar P, et al. Epigenetic associations of type 2 diabetes and BMI in an Arab population. Clin Epigenetics. 2016;8:13.
Ali O, Cerjak D, Kent JW Jr., James R, Blangero J, Carless MA, et al. Methylation of SOCS3 is inversely associated with metabolic syndrome in an epigenome-wide association study of obesity. Epigenetics. 2016;11:699–707.
Mendelson MM, Marioni RE, Joehanes R, Liu C, Hedman AK, Aslibekyan S, et al. Association of body mass index with DNA methylation and gene expression in blood cells and relations to cardiometabolic disease: a mendelian randomization approach. PLoS Med. 2017;14:e1002215.
Wahl S, Drong A, Lehne B, Loh M, Scott WR, Kunze S, et al. Epigenome-wide association study of body mass index, and the adverse outcomes of adiposity. Nature. 2017;541:81–86.
Wilson LE, Harlid S, Xu Z, Sandler DP, Taylor JA. An epigenome-wide study of body mass index and DNA methylation in blood using participants from the Sister Study cohort. Int J Obes. 2017;41:194–9.
Geurts YM, Dugue PA, Joo JE, Makalic E, Jung CH, Guan W et al. Novel associations between blood DNA methylation and body mass index in middle-aged and older adults. Int J Obes. 2017; e-pub ahead of print16 Oct 2017; https://doi.org/10.1038/ijo.2017.269.
Chambers JC, Loh M, Lehne B, Drong A, Kriebel J, Motta V, et al. Epigenome-wide association of DNA methylation markers in peripheral blood from Indian Asians and Europeans with incident type 2 diabetes: a nested case-control study. Lancet Diabetes Endocrinol. 2015;3:526–34.
Li S, Wong EM, Southey MC, Hopper JL. Association between DNA methylation at SOCS3 gene and body mass index might be due to familial confounding. Int J Obes. 2017;41:995–6.
Relton CL, Davey Smith G. Two-step epigenetic Mendelian randomization: a strategy for establishing the causal role of epigenetic processes in pathways to disease. Int J Epidemiol. 2012;41:161–76.
VanderWeele TJ, Tchetgen Tchetgen EJ, Cornelis M, Kraft P. Methodological challenges in mendelian randomization. Epidemiology. 2014;25:427–35.
Odefrey F, Stone J, Gurrin LC, Byrnes GB, Apicella C, Dite GS, et al. Common genetic variants associated with breast cancer and mammographic density measures that predict disease. Cancer Res. 2010;70:1449–58.
Li S, Wong EM, Joo JE, Jung CH, Chung J, Apicella C, et al. Genetic and environmental causes of variation in the difference between biological age based on DNA methylation and chronological age for middle-aged women. Twin Res Hum Genet. 2015;18:720–6.
Joo JE, Wong EM, Baglietto L, Jung CH, Tsimiklis H, Park DJ, et al. The use of DNA from archival dried blood spots with the Infinium HumanMethylation450 array. BMC Biotechnol. 2013;13:23.
Aryee MJ, Jaffe AE, Corrada-Bravo H, Ladd-Acosta C, Feinberg AP, Hansen KD, et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics. 2014;30:1363–9.
Maksimovic J, Gordon L, Oshlack A. SWAN: subset-quantile within array normalization for illumina infinium HumanMethylation450 BeadChips. Genome Biol. 2012;13:R44.
Johnson WE, Li C, Rabinovic A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics. 2007;8:118–27.
Price ME, Cotton AM, Lam LL, Farre P, Emberly E, Brown CJ, et al. Additional annotation enhances potential for biologically-relevant analysis of the Illumina Infinium HumanMethylation450 BeadChip array. Epigenetics Chromatin. 2013;6:4.
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinformatics. 2012;13:86.
Benjamini Y, Hochberg Y. Controlling the false discovery rate - a practical and powerful approach to multiple testing. J R Stat Soc B Methodol. 1995;57:289–300.
Hopper JL, Bui QM, Erbas B, Matheson MC, Gurrin LC, Burgess JA, et al. Does eczema in infancy cause hay fever, asthma, or both in childhood? Insights from a novel regression model of sibling data. J Allergy Clin Immunol. 2012;130:1117-22.e11.
Stone J, Dite GS, Giles GG, Cawson J, English DR, Hopper JL. Inference about causation from examination of familial confounding: application to longitudinal twin data on mammographic density measures that predict breast cancer risk. Cancer Epidemiol Biomark Prev. 2012;21:1149–55.
Bui M, Bjornerem A, Ghasem-Zadeh A, Dite GS, Hopper JL, Seeman E. Architecture of cortical bone determines in part its remodelling and structural decay. Bone. 2013;55:353–8.
Dite GS, Gurrin LC, Byrnes GB, Stone J, Gunasekara A, McCredie MR, et al. Predictors of mammographic density: insights gained from a novel regression analysis of a twin study. Cancer Epidemiol Biomark Prev. 2008;17:3474–81.
Davey CG, Lopez-Sola C, Bui M, Hopper JL, Pantelis C, Fontenelle LF, et al. The effects of stress-tension on depression and anxiety symptoms: evidence from a novel twin modelling analysis. Psychol Med. 2016;46:3213–8.
Greenberg AS, Obin MS. Obesity and the role of adipose tissue in inflammation and metabolism. Am J Clin Nutr. 2006;83:461S–465S.
Gregor MF, Hotamisligil GS. Inflammatory mechanisms in obesity. Annu Rev Immunol. 2011;29:415–45.
Monteiro R, Azevedo I. Chronic inflammation in obesity and the metabolic syndrome. Mediat Inflamm. 2010;2010:289645.
Sintes J, Cuenca M, Romero X, Bastos R, Terhorst C, Angulo A, et al. Cutting edge: Ly9 (CD229), a SLAM family receptor, negatively regulates the development of thymic innate memory-like CD8+T and invariant NKT cells. J Immunol. 2013;190:21–26.
Chatterjee M, Rauen T, Kis-Toth K, Kyttaris VC, Hedrich CM, Terhorst C, et al. Increased expression of SLAM receptors SLAMF3 and SLAMF6 in systemic lupus erythematosus T lymphocytes promotes Th17 differentiation. J Immunol. 2012;188:1206–12.
Ikeda A, Hasegawa K, Masaki M, Moriguchi T, Nishida E, Kozutsumi Y, et al. Mixed lineage kinase LZK forms a functional signaling complex with JIP-1, a scaffold protein of the c-Jun NH(2)-terminal kinase pathway. J Biochem. 2001;130:773–81.
Masaki M, Ikeda A, Shiraki E, Oka S, Kawasaki T. Mixed lineage kinase LZK and antioxidant protein-1 activate NF-kappaB synergistically. Eur J Biochem. 2003;270:76–83.
Hsu YM, Zhang Y, You Y, Wang D, Li H, Duramad O, et al. The adaptor protein CARD9 is required for innate immune responses to intracellular pathogens. Nat Immunol. 2007;8:198–205.
Netea MG, Azam T, Ferwerda G, Girardin SE, Walsh M, Park JS, et al. IL-32 synergizes with nucleotide oligomerization domain (NOD) 1 and NOD2 ligands for IL-1beta and IL-6 production through a caspase 1-dependent mechanism. Proc Natl Acad Sci USA. 2005;102:16309–14.
Davey MP, Martin TM, Planck SR, Lee J, Zamora D, Rosenbaum JT. Human endothelial cells express NOD2/CARD15 and increase IL-6 secretion in response to muramyl dipeptide. Microvasc Res. 2006;71:103–7.
van Beelen AJ, Zelinkova Z, Taanman-Kueter EW, Muller FJ, Hommes DW, Zaat SA, et al. Stimulation of the intracellular bacterial sensor NOD2 programs dendritic cells to promote interleukin-17 production in human memory T cells. Immunity. 2007;27:660–9.
McDonald C, Chen FF, Ollendorff V, Ogura Y, Marchetto S, Lecine P, et al. A role for Erbin in the regulation of Nod2-dependent NF-kappaB signaling. J Biol Chem. 2005;280:40301–9.
Zhao Y, Alonso C, Ballester I, Song JH, Chang SY, Guleng B, et al. Control of NOD2 and Rip2-dependent innate immune activation by GEF-H1. Inflamm Bowel Dis. 2012;18:603–12.
Richmond RC, Sharp GC, Ward ME, Fraser A, Lyttleton O, McArdle WL, et al. DNA methylation and BMI: investigating identified methylation sites at HIF3A in a causal framework. Diabetes. 2016;65:1231–44.
Xu X, Su S, Barnes V, Snieder H, Wang X. Abstract MP55: DNA methylation mediates the effects of obesity on insulin resistance in african american youth and young adults. Circulation 2014; 129(Suppl 1): AMP55.
Richmond RC, Hemani G, Tilling K, Davey Smith G, Relton CL. Challenges and novel approaches for investigating molecular mediation. Hum Mol Genet. 2016;25:R149–R156.
Acknowledgements
We thank all women participating in this study. The data analysis was facilitated by Spartan, the High Performance Computer and Cloud hybrid system of the University of Melbourne. This research was facilitated through access to Twins Research Australia, a national resource supported by a Centre of Research Excellence Grant (ID: 1079102), from the National Health and Medical Research Council (NHMRC). . The AMDTSS was supported by NHMRC (grant numbers 1050561 and 1079102), Cancer Australia and National Breast Cancer Foundation (grant number 509307). SL is supported by the Australian Government Research Training Program Scholarship from the University of Melbourne. TLN is supported by a NHMRC Post-Graduate Scholarship and the Richard Lovell Travelling Scholarship from the University of Melbourne. MCS is a NHMRC Senior Research Fellow. JLH is a NHMRC Senior Principal Research Fellow.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Li, S., Wong, E.M., Bui, M. et al. Inference about causation between body mass index and DNA methylation in blood from a twin family study. Int J Obes 43, 243–252 (2019). https://doi.org/10.1038/s41366-018-0103-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-018-0103-4
This article is cited by
-
Epigenome-wide association study in Chinese monozygotic twins identifies DNA methylation loci associated with blood pressure
Clinical Epigenetics (2023)
-
Identification of key DNA methylation changes on fasting plasma glucose: a genome-wide DNA methylation analysis in Chinese monozygotic twins
Diabetology & Metabolic Syndrome (2023)
-
Causal relationships between breast cancer risk factors based on mammographic features
Breast Cancer Research (2023)
-
Replicating associations between DNA methylation and body mass index in a longitudinal sample of older twins
International Journal of Obesity (2020)
-
Deciphering the DNA Methylome of Polycystic Ovary Syndrome
Molecular Diagnosis & Therapy (2020)