ABSTRACT
Background
Obesity was established as a relevant modifiable risk factor in the onset and progression of colorectal cancer (CRC). This relationship could be mediated by an epigenetic regulation.
Objectives
The current work aimed to explore the effects of excess body weight on the DNA methylation profile of CRC using a genome-wide DNA methylation approach and to identify an epigenetic signature of obesity-related CRC.
Methods
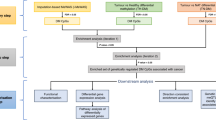
Fifty-six CRC-diagnosed patients (50 years) were included in the study and categorized according to their body mass index (BMI) as non-obese (BMI ≤ 25 kg/m2) or overweight/obese (BMI > 25 kg/m2). Data from Infinium 450k array-based methylomes of 28 CRC tumor samples were coupled with information on BMI categories. Additionally, DNA methylation results were validated in 28 CRC tumor samples.
Results
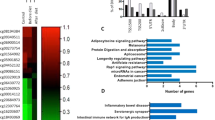
The analysis revealed statistically significant differences at 299 CpG sites, and they were mostly characterized as changes towards CpG hypermethylation occurring in the obese group. The 152 identified genes were involved in inflammatory and metabolic functional processes. Among these genes, novel genes were identified as epigenetically regulated in CRC depending on adiposity. ZNF397OS and ZNF543 represented the top scoring associated events that were further validated in an independent cohort and exhibited strong correlation with BMI and excellent and statistically significant efficiency in the discrimination of obese from non-obese CRC patients (area under the curve >0.80; p < 0.05).
Conclusions
The present study identifies a potential epigenome mark of obesity-related CRC that could be useful for precision medicine in the management of this disease taking into account adiposity as a relevant risk factor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Stevens GA, Singh GM, Lu Y, Danaei G, Lin JK, Finucane MM, et al. National, regional, and global trends in adult overweight and obesity prevalences. Popul Health Metr. 2012;10:22.
Calle EE, Rodriguez C, Walker-Thurmond K, Thun MJ. Overweight, obesity, and mortality from cancer in a prospectively studied cohort of U.S. adults. N Engl J Med. 2003;348:1625–38.
Kyrgiou M, Kalliala I, Markozannes G, Gunter MJ, Paraskevaidis E, Gabra H, et al. Adiposity and cancer at major anatomical sites: umbrella review of the literature. BMJ. 2017;356:j477.
Marmot M, et al. Food, nutrition, physical activity, and the prevention of cancer: a global perspective. World Cancer Research Fund/American Institute for Cancer Research; 2007. https://www.wcrf.org/sites/default/files/english.pdf.
Campbell PT, Newton CC, Dehal AN, Jacobs EJ, Patel AV, Gapstur SM. Impact of body mass index on survival after colorectal cancer diagnosis: the Cancer Prevention Study-II Nutrition Cohort. J Clin Oncol. 2012;30:42–52.
Freisling H, Arnold M, Soerjomataram I, O’Doherty MG, Ordonez-Mena JM, Bamia C, et al. Comparison of general obesity and measures of body fat distribution in older adults in relation to cancer risk: meta-analysis of individual participant data of seven prospective cohorts in Europe. Br J Cancer. 2017;116:1486–97.
Parekh N, Chandran U, Bandera EV. Obesity in cancer survival. Annu Rev Nutr. 2012;32:311–42.
Zheng J, Zhao M, Li J, Lou G, Yuan Y, Bu S, Xi Y, et al. Obesity-associated digestive cancers: A review of mechanisms and interventions. Tumour Biol. 2017;39:1010428317695020. https://doi.org/10.1177/1010428317695020. Review.
Crujeiras AB, Casanueva FF. Obesity and the reproductive system disorders: epigenetics as a potential bridge. Hum Reprod Update. 2015;21:249–61.
Crujeiras AB, Diaz-Lagares A, Stefansson OA, Macias-Gonzalez M, Sandoval J, Cueva J, et al. Obesity and menopause modify the epigenomic profile of breast cancer. Endocr Relat Cancer. 2017;24:331–43.
Crujeiras AB, Diaz-Lagares. A DNA methylation in obesity and associated diseases. In: Garcia-Gimenez JL, editor. Epigenetic biomarkers and diagnostics. Elsevier ed. Amsterdam: Elsevier; 2015. p. 313-29.
Bird A. DNA methylation patterns and epigenetic memory. Genes Dev. 2002;16:6–21.
Laird PW. The power and the promise of DNA methylation markers. Nat Rev Cancer. 2003;3:253–66.
Sharma S, Kelly TK, Jones PA. Epigenetics in cancer. Carcinogenesis. 2010;31:27–36.
Wong JJ, Hawkins NJ, Ward RL. Colorectal cancer: a model for epigenetic tumorigenesis. Gut. 2007;56:140–8.
Elliott EN, Kaestner KH. Epigenetic regulation of the intestinal epithelium. Cell Mol Life Sci. 2015;72:4139–56.
Hagland HR, Berg M, Jolma IW, Carlsen A, Soreide K. Molecular pathways and cellular metabolism in colorectal cancer. Dig Surg. 2013;30:12–25.
Crujeiras AB, Diaz-Lagares A, Moreno-Navarrete JM, Sandoval J, Hervas D, Gomez A. et al. Genome-wide DNA methylation pattern in visceral adipose tissue differentiates insulin-resistant from insulin-sensitive obese subjects. Transl Res. 2016;178:13–24, e5.
World Health Organization (WHO). Obesity: Preventing and Managing the Global Epidemic. Geneva: World Health Organization; 2000.
Sandoval J, Heyn H, Moran S, Serra-Musach J, Pujana MA, Bibikova M, et al. Validation of a DNA methylation microarray for 450,000 CpG sites in the human genome. Epigenetics. 2011;6:692–702.
Weisenberger DJ, Siegmund KD, Campan M, Young J, Long TI, Faasse MA, et al. CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet. 2006;38:787–93.
Weisenberger DJ, Levine AJ, Long TI, Buchanan DD, Walters R, Clendenning M, et al. Association of the colorectal CpG island methylator phenotype with molecular features, risk factors, and family history. Cancer Epidemiol Biomark Prev. 2015;24:512–9.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Gonzalez-Muniesa P, Martinez-Gonzalez MA, Hu FB, Despres JP, Matsuzawa Y, Loos RJF, et al. Obesity. Nat Rev Dis Prim. 2017;3:17034.
Crujeiras AB, Diaz-Lagares A, Carreira MC, Amil M, Casanueva FF. Oxidative stress associated to dysfunctional adipose tissue: a potential link between obesity, type 2 diabetes mellitus and breast cancer. Free Radic Res. 2013;47:243–56.
Howe LR, Subbaramaiah K, Hudis CA, Dannenberg AJ. Molecular pathways: adipose inflammation as a mediator of obesity-associated cancer. Clin Cancer Res. 2013;19:6074–83.
Funahashi Y, Yoshino Y, Yamazaki K, Mori Y, Mori T, Ozaki Y, et al. DNA methylation changes at SNCA intron 1 in patients with dementia with Lewy bodies. Psychiatry Clin Neurosci. 2017;71:28–35.
Gao Y, Li Z, Guo X, Liu Y, Zhang K. DLX4 as a prognostic marker for hepatocellular carcinoma. Neoplasma. 2014;61:318–23.
Hollington P, Neufing P, Kalionis B, Waring P, Bentel J, Wattchow D, et al. Expression and localization of homeodomain proteins DLX4, HB9 and HB24 in malignant and benign human colorectal tissues. Anticancer Res. 2004;24:955–62.
Kim JS, Park IS, Park HE, Kim SY, Yun JA, Jung CK, et al. alpha-Synuclein in the colon and premotor markers of Parkinson disease in neurologically normal subjects. Neurol Sci. 2017;38:171–9.
Li B, Li B, Sun H, Zhang H. The predicted target gene validation, function, and prognosis studies of miRNA-22 in colorectal cancer tissue. The journal Tumour Biol. 2017;39:1010428317692257.
Li Z, Liu Q, Piao J, Hua F, Wang J, Jin G, et al. Clinicopathological implications of Tiam1 overexpression in invasive ductal carcinoma of the breast. BMC Cancer. 2016;16:681.
Lind GE, Danielsen SA, Ahlquist T, Merok MA, Andresen K, Skotheim RI, et al. Identification of an epigenetic biomarker panel with high sensitivity and specificity for colorectal cancer and adenomas. Mol Cancer. 2011;10:85.
Minard ME, Ellis LM, Gallick GE. Tiam1 regulates cell adhesion, migration and apoptosis in colon tumor cells. Clin Exp Metastas-. 2006;23:301–13.
Prigent A, Lionnet A, Corbille AG, Derkinderen P. [Neuropathology and pathophysiology of Parkinson’s disease: Focus on alpha-synuclein]. Presse Med. 2017;46(Part 1):182–6.
Rotermund C, Truckenmuller FM, Schell H, Kahle PJ. Diet-induced obesity accelerates the onset of terminal phenotypes in alpha-synuclein transgenic mice. J Neurochem. 2014;131:848–58.
Rothman SM, Griffioen KJ, Fishbein KW, Spencer RG, Makrogiannis S, Cong WN, et al. Metabolic abnormalities and hypoleptinemia in alpha-synuclein A53T mutant mice. Neurobiol Aging. 2014;35:1153–61.
Tang H, Wei P, Duell EJ, Risch HA, Olson SH, Bueno-de-Mesquita HB, et al. Genes–environment interactions in obesity- and diabetes-associated pancreatic cancer: a GWAS data analysis. Cancer Epidemiol Biomark Prev. 2014;23:98–106.
Trinh B, Ko SY, Haria D, Barengo N, Naora H. The homeoprotein DLX4 controls inducible nitric oxide synthase-mediated angiogenesis in ovarian cancer. Mol Cancer. 2015;14:97.
Zhang TJ, Zhou JD, Yang DQ, Wang YX, Yao DM, Ma JC, et al. Hypermethylation of DLX4 predicts poor clinical outcome in patients with myelodysplastic syndrome. Clin Chem Lab Med. 2016;54:865–71.
Zhou JD, Zhang TJ, Wang YX, Yang DQ, Yang L, Ma JC, et al. DLX4 hypermethylation is a prognostically adverse indicator in de novo acute myeloid leukemia. Tumour Biol. 2016;37:8951–60.
Xu D, Qu L, Hu J, Li G, Lv P, Ma D, et al. Transmembrane protein 106A is silenced by promoter region hypermethylation and suppresses gastric cancer growth by inducing apoptosis. J Cell Mol Med. 2014;18:1655–66.
Diaz-Lagares A, Mendez-Gonzalez J, Hervas D, Saigi M, Pajares MJ, Garcia D, et al. A novel epigenetic signature for early diagnosis in lung cancer. Clin Cancer Res. 2016;22:3361–71.
Severson PL, Tokar EJ, Vrba L, Waalkes MP, Futscher BW. Coordinate H3K9 and DNA methylation silencing of ZNFs in toxicant-induced malignant transformation. Epigenetics. 2013;8:1080–8.
Stefansson OA, Moran S, Gomez A, Sayols S, Arribas-Jorba C, Sandoval J, et al. A DNA methylation-based definition of biologically distinct breast cancer subtypes. Mol Oncol. 2015;9:555–68.
Acknowledgements
We thank María Amil, Diana García, and Carles Arribas for their technical support, as well as Biobank of the Public Health System of Andalusia (BBSSPA) of the health counseling and the head of the Anatomical Pathology Department of the Hospital Complex of Specialties Virgen de la Victoria Malaga Spain.
Funding
This study was supported by “Centros de Investigacion Biomedica En Red” (CIBERobn) of the “Instituto de Salud Carlos III” (ISCIII), and grants from ISCIII (PI11/01661, PI14/01012, PI15/01114, PE13/00024) co-financed by the European Regional Development Fund (FEDER). DC-C was recipient of an FPU grant from Education Ministry, Madrid, Spain (13/04211). AD-L was funded by the ISCIII through a research contract "Rio Hortega" (CM14/00067) and "Juan Rodés" (JR17/00016), ABC and JS are “Miguel Servet” researchers (ISCIII, CP17/00088 and CP13/00055, respectively) and MM-G was recipient of the Nicolas Monarde program from the Servicio Andaluz de Salud, Junta de Andalucia, Spain (C-0029-2014).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Crujeiras, A.B., Morcillo, S., Diaz-Lagares, A. et al. Identification of an episignature of human colorectal cancer associated with obesity by genome-wide DNA methylation analysis. Int J Obes 43, 176–188 (2019). https://doi.org/10.1038/s41366-018-0065-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-018-0065-6
This article is cited by
-
Discovery of novel DNA methylation biomarker panels for the diagnosis and differentiation between common adenocarcinomas and their liver metastases
Scientific Reports (2024)
-
Effects of obesity, and of weight loss following bariatric surgery, on methylation of DNA from the rectal mucosa and in cell-free DNA from blood
International Journal of Obesity (2023)
-
Alterations of DNA methylation and expression of genes related to thyroid hormone metabolism in colon epithelium of obese patients
BMC Medical Genomics (2022)
-
Behavioral Risk Factors and Risk of Early-Onset Colorectal Cancer: Review of the Mechanistic and Observational Evidence
Current Colorectal Cancer Reports (2021)
-
Body mass index-associated molecular characteristics involved in tumor immune and metabolic pathways
Cancer & Metabolism (2020)