Abstract
Background
Obesity is an established risk factor for several common chronic diseases such as breast and colorectal cancer, metabolic and cardiovascular diseases; however, the biological basis for these relationships is not fully understood. To explore the association of obesity with these conditions, we investigated peripheral blood leucocyte (PBL) DNA methylation markers for adiposity and their contribution to risk of incident breast and colorectal cancer and myocardial infarction.
Methods
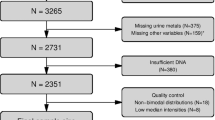
DNA methylation profiles (Illumina Infinium® HumanMethylation450 BeadChip) from 1941 individuals from four population-based European cohorts were analysed in relation to body mass index, waist circumference, waist-hip and waist-height ratio within a meta-analytical framework. In a subset of these individuals, data on genome-wide gene expression level, biomarkers of glucose and lipid metabolism were also available. Validation of methylation markers associated with all adiposity measures was performed in 358 individuals. Finally, we investigated the association of obesity-related methylation marks with breast, colorectal cancer and myocardial infarction within relevant subsets of the discovery population.
Results
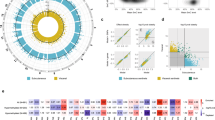
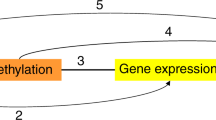
We identified 40 CpG loci with methylation levels associated with at least one adiposity measure. Of these, one CpG locus (cg06500161) in ABCG1 was associated with all four adiposity measures (P = 9.07×10−8 to 3.27×10−18) and lower transcriptional activity of the full-length isoform of ABCG1 (P = 6.00×10−7), higher triglyceride levels (P = 5.37×10−9) and higher triglycerides-to-HDL cholesterol ratio (P = 1.03×10−10). Of the 40 informative and obesity-related CpG loci, two (in IL2RB and FGF18) were significantly associated with colorectal cancer (inversely, P < 1.6×10−3) and one intergenic locus on chromosome 1 was inversely associated with myocardial infarction (P < 1.25×10−3), independently of obesity and established risk factors.
Conclusion
Our results suggest that epigenetic changes, in particular altered DNA methylation patterns, may be an intermediate biomarker at the intersection of obesity and obesity-related diseases, and could offer clues as to underlying biological mechanisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Stevens GA, Singh GM, Lu Y, Danaei G, Lin JK, Finucane MM et al. National, regional, and global trends in adult overweight and obesity prevalences. Popul Health Metr. 2012;10:16. https://doi.org/10.1186/1478-7954-10-22
Arnold M, Pandeya N, Byrnes G, Renehan AG, Stevens GA, Ezzati M, et al. Global burden of cancer attributable to high body-mass index in 2012: a population-based study. Lancet Oncol. 2015;16:36–46.
Bhaskaran K, Douglas I, Forbes H, dos-Santos-Silva I, Leon DA, Smeeth L. Body-mass index and risk of 22 specific cancers: a population-based cohort study of 5.24 million UK adults. Lancet. 2014;384:755–65.
Samuel VT, Shulman GI. Mechanisms for insulin resistance: common threads and missing links. Cell. 2012;148:852–71.
Speliotes EK, Willer CJ, Berndt SI, Monda KL, Thorleifsson G, Jackson AU, et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat Genet. 2010;42:937–48.
Heid IM, Jackson AU, Randall JC, Winkler TW, Qi L, Steinthorsdottir V, et al. Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution. Nat Genet. 2010;42:949–60.
Yang J, Loos RJF, Powell JE, Medland SE, Speliotes EK, Chasman DI, et al. FTO genotype is associated with phenotypic variability of body mass index. Nature. 2012;490:267–72.
Scott RA, Lagou V, Welch RP, Wheeler E, Montasser ME, Luan Ja, et al. Large-scale association analyses identify new loci influencing glycemic traits and provide insight into the underlying biological pathways. Nat Genet. 2012;44:991–1005.
Dick KJ, Nelson CP, Tsaprouni L, Sandling JK, Aissi D, Wahl S, et al. DNA methylation and body-mass index: a genome-wide analysis. Lancet. 2014;383:1990–8.
Wahl S, Drong A, Lehne B, Loh M, Scott WR, Kunze S, et al. Epigenome-wide association study of body mass index, and the adverse outcomes of adiposity. Nature. 2017;541:81–86.
Chambers JC, Loh M, Lehne B, Drong A, Kriebel J, Motta V, et al. Epigenome-wide association of DNA methylation markers in peripheral blood from Indian Asians and Europeans with incident type 2 diabetes: a nested case-control study. Lancet Diabetes Endocrinol. 2015;3:526–34.
Palli D, Berrino F, Vineis P, Tumino R, Panico S, Masala G, et al. A molecular epidemiology project on diet and cancer: the EPIC-Italy Prospective Study. Design and baseline characteristics of participants. Tumori. 2003;89:586–93.
Beulens JWJ, Monninkhof EM, Verschuren WMM, van der Schouw YT, Smit J, Ocke MC, et al. Cohort profile: the EPIC-NL study. Int J Epidemiol. 2010;39:1170–8.
Lund E, Kumle M, Braaten T, Hjartaker A, Bakken K, Eggen E, et al. External validity in a population-based national prospective study—the Norwegian Women and Cancer Study (NOWAC). Cancer Causes Control. 2003;14:1001–8.
Hallmans G, Agren A, Johansson G, Johansson A, Stegmayr B, Jansson J-H, et al. Cardiovascular disease and diabetes in the Northern Sweden Health and Disease Study Cohort—evaluation of risk factors and their interactions. Scand J Public Health. 2003;61:18–24.
Chadeau-Hyam M, Vermeulen RCH, DGAJ Hebels, Castagne R, Campanella G, Portengen L, et al. Prediagnostic transcriptomic markers of chronic lymphocytic leukemia reveal perturbations 10 years before diagnosis. Ann Oncol. 2014;25:1065–72.
Dumeaux V, Borresen-Dale A-L, Frantzen J-O, Kumle M, Kristensen VN, Lund E. Gene expression analyses in breast cancer epidemiology: the Norwegian Women and Cancer postgenome cohort study. Breast Cancer Res. 2008;10:R13 https://doi.org/10.1186/bcr1859
DGAJ Hebels, Georgiadis P, Keun HC, Athersuch TJ, Vineis P, Vermeulen R, et al. Performance in omics analyses of blood samples in long-term storage: opportunities for the exploitation of existing biobanks in environmental health research. Environ Health Perspect. 2013;121:480–7.
Ferrari SLP, Cribari-Neto F. Beta regression for modelling rates and proportions. J Appl Stat. 2004;31:799–815.
Higgins JPT, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002;21:1539–58.
Chen YA, Lemire M, Choufani S, Butcher DT, Grafodatskaya D, Zanke BW, et al. Discovery of cross-reactive probes and polymorphic CpGs in the Illumina Infinium HumanMethylation450 microarray. Epigenetics. 2013;8:203–9.
Dumeaux V, Olsen KS, Nuel G, Paulssen RH, Borresen-Dale AL, Lund E Deciphering normal blood gene expression variation—the NOWAC Postgenome Study. PLoS Genet. 2010;6:e1000873
Kannel WB, Vasan RS, Keyes MJ, Sullivan LM, Robins SJ. Usefulness of the triglyceride-high-density lipoprotein versus the cholesterol-high-density lipoprotein ratio for predicting insulin resistance and cardiometabolic risk (from the Framingham offspring cohort). Am J Cardiol. 2008;101:497–501.
Matthews DR, Hosker JP, Rudenski AS, Naylor BA, Treacher DF, Turner RC. Homeostasis model assessment: insulin resistance and beta-cell function from fasting plasma glucose and insulin concentrations in man. Diabetologia. 1985;28:412–9.
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinform. 2012;13:86.
Reinius LE, Acevedo N, Joerink M, Pershagen G, Dahlén S-E, Greco D, et al. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PLoS ONE. 2012;7:e41361.
Michels KB, Binder AM, Dedeurwaerder S, Epstein CB, Greally JM, Gut I, et al. Recommendations for the design and analysis of epigenome-wide association studies. Nat Methods. 2013;10:949–55.
Altshuler DM, Durbin RM, Abecasis GR, Bentley DR, Chakravarti A, Clark AG, et al. An integrated map of genetic variation from 1,092 human genomes. Nature. 2012;491:56–65.
Pfeifferm L, Wahl S, Pilling LC, Reischl E, Sandling JK, Kunze S et al. DNA methylation of lipid-related genes affects blood lipid levels. Circ Cardiovasc Genet. 2015;8:334–342
Li G, Gu H-M, Zhang D-W. ATP-binding cassette transporters and cholesterol translocation. IUBMB Life. 2013;65:505–12.
Vaughan AM, Oram JF. ABCA1 and ABCG1 or ABCG4 act sequentially to remove cellular cholesterol and generate cholesterol-rich HDL. J Lipid Res. 2006;47:2433–43.
Klucken J, Buchler C, Orso E, Kaminski WE, Porsch-Ozcurumez M, Liebisch C, et al. ABCG1 (ABC8), the human homolog of the Drosophila white gene, is a regulator of macrophage cholesterol and phospholipid transport. Proc Natl Acad Sci USA. 2000;97:817–22.
Kennedy MA, Barrera GC, Nakamura K, Baldan A, Tarr P, Fishbein MC, et al. ABCG1 has a critical role in mediating cholesterol efflux to HDL and preventing cellular lipid accumulation. Cell Metab. 2005;1:121–31.
Olivier M, Tanck MW, Out R, Villard EF, Lammers B, Bouchareychas L, et al. Human ATP-binding cassette G1 controls macrophage lipoprotein lipase bioavailability and promotes foam cell formation. Arterioscler Thromb Vasc Biol. 2012;32:2223–31.
Mahaney MC, Blangero J, Comuzzie AG, Vandeberg JL, Stern MP, Maccluer JW. Plasma HDL, cholesterol, triglycerides, and adiposity—a quantitative genetic test of the conjoint trait hypothesis in the San Antonio Family Heart study. Circulation. 1995;92:3240–8.
Frazier-Wood AC, Aslibekyan S, Absher DM, Hopkins PN, Sha J, Tsai MY, et al. Methylation at CPT1A locus is associated with lipoprotein subfraction profiles. J Lipid Res. 2014;55:1324–30.
Gagnon F, Aissi D, Carrie A, Morange P-E, Tregouet D-A. Robust validation of methylation levels association at CPT1A locus with lipid plasma levels. J Lipid Res. 2014;55:1189–91.
Tobi EW, Goeman JJ, Monajemi R, Gu H, Putter H, Zhang Y, et al. DNA methylation signatures link prenatal famine exposure to growth and metabolism. Nat Commun. 2014;5:5592.
Stefanovic-Racic M, Perdomo G, Mantell BS, Sipula IJ, Brown NF, O’Doherty RM. A moderate increase in carnitine palmitoyltransferase 1a activity is sufficient to substantially reduce hepatic triglyceride levels. Am J Physiol—Endocrinol Metab. 2008;294:E969–77.
Drzewinska J, Walczak-Drzewiecka A, Ratajewski M. Identification and analysis of the promoter region of the human DHCR24 gene: involvement of DNA methylation and histone acetylation. Mol Biol Rep. 2011;38:1091–101.
McGrath KCY, Li XH, Puranik R, Liong EC, Tan JTM, Dy VM, et al. Role of 3β-hydroxysteroid-Δ24 reductase in mediating antiinflammatory effects of high-density lipoproteins in endothelial cells. Arterioscler Thromb Vasc Biol. 2009;29:877–82.
Wu BJ, Chen K, Shrestha S, Ong KL, Barter PJ, Rye K-A. High-density lipoproteins inhibit vascular endothelial inflammation by increasing 3β-hydroxysteroid-Δ24 reductase expression and inducing heme oxygenase-1. Circ Res. 2013;112:278–88.
Wu C, Miloslavskaya I, Demontis S, Maestro R, Galaktionov K. Regulation of cellular response to oncogenic and oxidative stress by seladin-1. Nature. 2004;432:640–5.
Chang YT, Huang CS, Yao CT, Su SL, Terng HJ, Chou HL, et al. Gene expression profile of peripheral blood in colorectal cancer. World J Gastroenterol. 2014;20:14463–71.
Marshall KW, Mohr S, Khettabi FE, Nossova N, Chao S, Bao W, et al. A blood-based biomarker panel for stratifying current risk for colorectal cancer. Int J Cancer. 2010;126:1177–86.
Nishimura S, Manabe I, Nagasaki M, Eto K, Yamashita H, Ohsugi M, et al. CD8+ effector T cells contribute to macrophage recruitment and adipose tissue inflammation in obesity. Nat Med. 2009;15:914–20.
Shimokawa T, Furukawa Y, Sakai M, Li M, Miwa N, Lin Y-M, et al. Involvement of the FGF18 gene in colorectal carcinogenesis, as a novel downstream target of the β-catenin/T-cell factor complex. Cancer Res. 2003;63:6116–20.
Bienz M, Clevers H. Linking colorectal cancer to Wnt signaling. Cell. 2000;103:311–20.
Grant SF, Thorleifsson G, Reynisdottir I, Benediktsson R, Manolescu A, Sainz J, et al. Variant of transcription factor 7-like 2 (TCF7L2) gene confers risk of type 2 diabetes. Nat Genet. 2006;38:320–3.
Welters HJ, Kulkarni RN. Wnt signaling: relevance to beta-cell biology and diabetes. Trends Endocrinol Metab. 2008;19:349–55.
Das M, Sha J, Hidalgo B, Aslibekyan S, Do AN, Zhi D, et al. Association of DNA methylation at CPT1A locus with metabolic syndrome in the genetics of lipid lowering drugs and Diet Network (GOLDN) Study. PLoS ONE. 2016;11:e0145789.
Demerath EW, Guan W, Grove ML, Aslibekyan S, Mendelson M, Zhou YH, et al. Epigenome-wide association study (EWAS) of BMI, BMI change and waist circumference in African American adults identifies multiple replicated loci. Hum Mol Genet. 2015;24:4464–79.
Author contributions
Samples from EPIC-Italy were provided by Vittorio Krogh, Domenico Palli, Salvatore Panico, Carlotta Sacerdote, and Rosario Tumino. Samples from EPIC-Netherlands were provided by H. Bas Bueno-de-Mesquita, Anne M. May, N. Charlotte Onland-Moret, Elio Riboli, and W. M. Monique Verschuren. Samples and data from NOWAC were provided by Eiliv Lund, Nicolle Mode, and Torkjel M. Sandanger. Samples and data from NSHDS were provided by Ingvar A. Bergdahl, Beatrice Melin, and Per Lenner. Giuseppe Matullo provided DNA methylation profiles, and Giovanni Fiorito performed management and data quality assurance for the EPICOR study. Soterios A. Kyrtopoulos provided DNA methylation profiles from the EnviroGenoMarkers project. Laboratory analyses were performed by Silvia Polidoro (EPIC-Italy and NOWAC), Simonetta Guarrera (EPIC-Italy), Panagiotis Georgiadis (DNA methylation in EnviroGenoMarkers), Theo M. C. M. de Kok and Jos C. S. Kleinjans (gene expression in EnviroGenoMarkers), and Karen A. Lillycrop and Robert Murray (EPIC-Netherlands). Measurements of blood lipids, glucose, and insulin for EPIC-Italy samples were provided by Licia Iacoviello. Bisulphite pyrosequencing of the Italian replication samples was conducted by Valentina Fiano and Morena Trevisan. Marc Chadeau-Hyam supervised the statistical analyses; Gianluca Campanella compiled the data, reviewed its quality, devised and carried out all statistical analyses. Gianluca Campanella, Marc Gunter and Silvia Polidoro drafted the manuscript. Philippe Froguel provided critical comments and contributed to the manuscript preparation. Paul Elliott, Paolo Vineis, and Marc Chadeau-Hyam coordinated the work and contributed to writing the manuscript. All authors approved the final version of this article.
Funding
EPIC-Italy was financially supported by the Italian Association for Cancer Research (AIRC). Genome-wide DNA methylation profiling and bisulphite pyrosequencing of EPIC-Italy samples was financially supported by the Human Genetics Foundation (HuGeF) and Compagnia di San Paolo. The EnviroGenoMarkers project was financially supported by the European Union (grant agreement 226756 to Soterios A. Kyrtopoulos). EPIC-Netherlands was financially supported by the Dutch Ministry of Public Health, Welfare, and Sports (VWS), by the Netherlands Cancer Registry, by LK Research Funds, by Dutch Prevention Funds, by the Netherlands Organisation for Health Research and Development (ZON), and by the World Cancer Research Fund (WCRF). Genome-wide DNA methylation profiling of EPIC-Netherlands samples was financially supported by internal Imperial College funds. Genome-wide DNA methylation and gene expression profiling of NOWAC samples was financially supported by the European Research Council for frontier research, Advanced Grant TICE—Transcriptomics in Cancer Epidemiology (number 232997, period 2009–2014). Gianluca Campanella received a Doctoral Prize studentship awarded by the Engineering and Physical Sciences Research Council (EPSRC). Paul Elliott is a National Institute for Health Research (NIHR) senior investigator and acknowledges support from the NIHR Biomedical Research Centre at Imperial College Healthcare NHS Trust and Imperial College London, and the NIHR Health Protection Unit on Health Impact of Environmental Hazards. He is supported by the Medical Research Council and Public Health England as part of joint funding for the MRC-PHE Centre for Environment and Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Campanella, G., Gunter, M.J., Polidoro, S. et al. Epigenome-wide association study of adiposity and future risk of obesity-related diseases. Int J Obes 42, 2022–2035 (2018). https://doi.org/10.1038/s41366-018-0064-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-018-0064-7
This article is cited by
-
A prospective study of smoking-related white blood cell DNA methylation markers and risk of bladder cancer
European Journal of Epidemiology (2024)
-
DNA methylation and cardiovascular disease in humans: a systematic review and database of known CpG methylation sites
Clinical Epigenetics (2023)
-
Genetics and Epigenetics in Obesity: What Do We Know so Far?
Current Obesity Reports (2023)
-
ABCA1 and ABCG1 DNA methylation in epicardial adipose tissue of patients with coronary artery disease
BMC Cardiovascular Disorders (2021)
-
DNA methylation signatures of incident coronary heart disease: findings from epigenome-wide association studies
Clinical Epigenetics (2021)