Abstract
Background/objectives
CYP2A6 (CYP2A5 in mice) is mainly expressed in the liver. Hepatic CYP2A6 expression is increased in patients with non-alcoholic fatty liver disease (NAFLD). In mice, hepatic CYP2A5 is induced by high fat diet (HFD) feeding. Hepatic CYP2A5 is also increased in monosodium glutamate-induced obese mice. NAFLD is associated with obesity. In this study, we examined whether obesity is related to CYP2A6.
Subjects/methods
Obesity genetic association study: The SAGE is a comprehensive genome-wide association study (GWAS) with case subjects having a lifetime history of alcohol dependence and control subjects never addicted to alcohol. We used 1030 control individuals with self-reported height and weight. A total of 12 single nucleotide polymorphisms (SNP) within the CYP2A6 gene were available. Obesity was determined as a BMI ≥30: 30–34.9 (Class I obesity) and ≥35 (Class II and III obesity). Animal experiment study: CYP2A5 knockout (cyp2a5−/−) mice and wild type (cyp2a5+/+) mice were fed HFD for 14 weeks. Body weight was measured weekly. After an overnight fast, the mice were sacrificed. Liver and blood were collected for biochemical assays.
Results
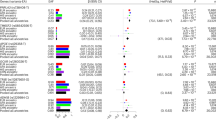
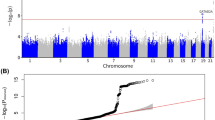
Single marker analysis showed that three SNPs (rs8192729, rs7256108, and rs7255443) were associated with class I obesity (p < 0.05). The most significant SNP for obesity was rs8192729 (odds ratio (OR) = 1.94, 95% confidence intervals = 1.21–3.10, p = 0.00582). After HFD feeding, body weight was increased in cyp2a5−/− mice to a greater extent than in cyp2a5+/+ mice, and fatty liver was more pronounced in cyp2a5−/− mice than in cyp2a5+/+ mice. PPARα deficiency in cyp2a5−/− mice developed more severe fatty liver, but body weight was not increased significantly.
Conclusion
CYP2A6 is associated with human obesity; CYP2A5 protects against obesity and NAFLD in mice. PPARα contributes to the CYP2A5 protective effects on fatty liver but it opposes to the protective effects on obesity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Leclercq IA, Farrell GC, Field J, Bell DR, Gonzalez FJ, Robertson GR. CYP2E1 and CYP4A as microsomal catalysts of lipid peroxides in murine nonalcoholic steatohepatitis. J Clin Invest. 2000;105:1067–75.
Lu Y, Zhuge J, Wang X, Bai J, Cederbaum AI. Cytochrome P450 2E1 contributes to ethanol-induced fatty liver in mice. Hepatology . 2008;47:1483–94.
Lu Y, Wu D, Wang X, Ward SC, Cederbaum AI. Chronic alcohol-induced liver injury and oxidant stress are decreased in cytochrome P4502E1 knockout mice and restored in humanized cytochrome P4502E1 knock-in mice. Free Radic Biol Med. 2010;49:1406–16.
Abdelmegeed MA, Banerjee A, Yoo SH, Jang S, Gonzalez FJ, Song BJ. Critical role of cytochrome P450 2E1 (CYP2E1) in the development of high fat-induced non-alcoholic steatohepatitis. J Hepatol. 2012;57:860–6.
Zong H, Armoni M, Harel C, Karnieli E, Pessin JE. Cytochrome P-450 CYP2E1 knockout mice are protected against high-fat diet-inducedobesity and insulin resistance. Am J Physiol Endocrinol Metab. 2012;302:E532–9.
Larsen MC, Bushkofsky JR, Gorman T, Adhami V, Mukhtar H, Wang S, et al. Cytochrome P450 1B1: an unexpected modulator of liver fatty acid homeostasis. Arch Biochem Biophys. 2015;571:21–39.
Su T, Ding X. Regulation of the cytochrome P450 2A genes. Toxicol Appl Pharmacol. 2004;199:285–94.
Niemelä O, Parkkila S, Juvonen RO, Viitala K, Gelboin HV, Pasanen M. Cytochromes P450 2A6, 2E1, and 3A and production of protein-aldehyde adducts in the liver of patients with alcoholic and non-alcoholic liver diseases. J Hepatol. 2000;33:893–901.
Lu Y, Zhuge J, Wu D, Cederbaum AI. Ethanol induction of CYP2A5: permissive role for CYP2E1. Drug Metab Dispos. 2011;39:330–6.
Lu Y, Zhang XH, Cederbaum AI. Ethanol induction of CYP2A5: role of CYP2E1-ROS-Nrf2 pathway. Toxicol Sci. 2012;128:427–38.
Hong F, Liu X, Ward SS, Xiong H, Cederbaum AI, Lu Y. Absence of cytochrome P450 2A5 enhances alcohol-induced liver injury in mice. Dig Liver Dis. 2015;47:470–7.
Chen X, Ward SC, Cederbaum AI, Xiong H, Lu Y. Alcoholic fatty liver is enhanced in CYP2A5 knockout mice: the role of the PPARα-FGF21 axis. Toxicology . 2017;379:12–21.
Tomankova V, Liskova B, Skalova L, Bartikova H, Bousova I, Jourova L, et al. Altered cytochrome P450 activities and expression levels in the liver and intestines of the monosodium glutamate-induced mouse model of human obesity. Life Sci. 2015;133:15–20.
Kersten S, Seydoux J, Peters JM, Gonzalez FJ, Desvergne B, Wahli W. Peroxisome proliferator-activated receptor alpha mediates the adaptive response to fasting. J Clin Invest. 1999;103:1489–98.
Leone TC, Weinheimer CJ, Kelly DP. A critical role for the peroxisome proliferator-activated receptor alpha (PPARalpha) in the cellular fasting response: the PPARalpha-null mouse as a model of fatty acid oxidation disorders. Proc Natl Acad Sci USA. 1999;96:7473–8.
Huang J, Jia Y, Fu T, Viswakarma N, Bai L, Rao MS, et al. Sustained activation of PPARα by endogenous ligands increases hepatic fatty acid oxidation and prevents obesity in ob/ob mice. FASEB J. 2012;26:628–38.
Gao Q, Jia Y, Yang G, Zhang X, Boddu PC, Petersen B, et al. PPARα-deficient ob/ob obese mice become more obese and manifest severe hepatic steatosis due to decreased fatty acid oxidation. Am J Pathol. 2015;185:1396–408.
Bierut LJ, Agrawal A, Bucholz KK, Doheny KF, Laurie C, Pugh E, et al. A genome-wide association study of alcohol dependence. Proc Natl Acad Sci USA. 2010;107:5082–87.
Wang KS, Liu X, Zhang QY, Pan Y, Aragam N, Zeng M. Meta-analysis of two genome-wide association studies identifies 3 new loci for alcohol dependence. J Psychiatr Res. 2011;45:1419–25.
Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21:263–5.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Zhou X, Zhuo X, Xie F, Kluetzman K, Humphreys WG, Ding X. Role of CYP2A5 in the clearance of nicotine and cotinine: insights from studies on a Cyp2a5-null mouse model. J Pharmacol Exp Ther. 2010;332:578–87.
Ardlie KG, Kruglyak L, Seielstad M. Patterns of linkage disequilibrium in the human genome. Nat Rev Genet. 2002;3:299–309.
Motomura W, Inoue M, Ohtake T, Takahashi N, Nagamine M, Tanno S, et al. Up-regulation of ADRP in fatty liver in human and liver steatosis in mice fed with high fat diet. Biochem Biophys Res Commun. 2006;340:1111–8.
Luo Y, McKeehan WL.Stressed liver and muscle call on adipocytes with FGF21. Front Endocrinol (Lausanne). 2013;4:194
Holland WL, Adams AC, Brozinick JT, Bui HH, Miyauchi Y, Kusminski CM, et al. An FGF21-adiponectin-ceramide axis controls energy expenditure and insulin action in mice. Cell Metab. 2013;17:790–7.
Lin Z, Tian H, Lam KS, Lin S, Hoo RC, Konishi M. Itoh, et al. Adiponectin mediates the metabolic effects of FGF21 on glucose homeostasis and insulin sensitivity in mice. Cell Metab. 2013;17:779–89.
Li Y, Wright GL, Peterson JM. C1q/TNF-related protein 3 (CTRP3) function and regulation. Comp Physiol. 2017;7:863–78.
Kirby GM, Nichols KD, Antenos M. CYP2A5 induction and hepatocellular stress: an adaptive response to perturbations of heme homeostasis. Curr Drug Metab. 2011;12:186–97.
Cui Y, Wang Q, Yi X, Zhang X. Effects of fatty acids on CYP2A5 and Nrf2 expression in mouse primary hepatocytes. Biochem Genet. 2016;54:29–40.
Wang XH, Cui XX, Sun XQ, Wang XH, Li XC, Qi Y, et al. High fat diet-induced hepatic 18-carbon fatty acids accumulation up-regulates CYP2A5/CYP2A6 via NF-E2-related factor 2. Front Pharmacol. 2017;8:233.
Montagner A, Polizzi A, Fouché E, Ducheix S, Lippi Y, Lasserre F, et al. Liver PPARα is crucial for whole-body fatty acid homeostasis and is protective against NAFLD. Gut. 2016;65:1202–14.
Peeters A, Baes M. Role of PPARa in hepatic carbohydrate metabolism. PPAR Res. 2010; 2010:572405.
Gurevich-Panigrahi T, Panigrahi S, Wiechec E, Los M. Obesity: pathophysiology and clinical management. Curr Med Chem. 2009;16:506–21.
Donnelly KL, Smith CI, Schwarzenberg SJ, Jessurun J, Boldt MD, Parks EJ. Sources of fatty acids stored in liver and secreted via lipoproteins in patients with nonalcoholic fatty liver disease. J Clin Invest. 2005;115:1343–51.
Kharitonenkov A, Shiyanova TL, Koester A, Ford AM, Micanovic R, Galbreath EJ, et al. FGF-21 as a novel metabolic regulator. J Clin Invest. 2005;115:1627–35.
Fisher FM, Estall JL, Adams AC, Antonellis PJ, Bina HA, Flier JS, Kharitonenkov A, et al. Integrated regulation of hepatic metabolism by fibroblast growth factor 21 (FGF21) in vivo. Endocrinology . 2011;52:2996–3004.
Badman MK, Pissios P, Kennedy AR, Koukos G, Flier JS, Maratos-Flier E. Hepatic fibroblast growth factor 21 is regulated by PPARalpha and is a key mediator of hepatic lipid metabolism in ketotic states. Cell Metab. 2007;5:426–37.
Potthoff MJ, Kliewer SA, Mangelsdorf DJ. Endocrine fibroblast growth factors 15/19 and 21: from feast to famine. Genes Dev. 2012;26:312–24.
Inagaki T, Dutchak P, Zhao G, Ding X, Gautron L, Parameswara V, et al. Endocrine regulation of the fasting response by PPARalpha-mediated induction of fibroblast growth factor 21. Cell Metab. 2007;5:415–25.
Lundåsen T, Hunt MC, Nilsson LM, Sanyal S, Angelin B, Alexson SE, et al. PPARalpha is a key regulator of hepatic FGF21. Biochem Biophys Res Commun. 2007;360:437–40.
Chakravarthy MV, Pan Z, Zhu Y, Tordjman K, Schneider JG, Coleman T, et al. “New” hepatic fat activates PPARalpha to maintain glucose, lipid, and cholesterol homeostasis. Cell Metab. 2005;1:309–22.
Asrih M, Veyrat-Durebex C, Poher AL, Lyautey J, Rohner-Jeanrenaud F, Jornayvaz FR. Leptin as a potential regulator of FGF21. Cell Physiol Biochem. 2016;38:1218–25.
Véniant MM, Hale C, Helmering J, Chen MM, Stanislaus S, Busby J, et al. FGF21 promotes metabolic homeostasis via white adipose and leptin in mice. PLoS One. 2012;7:e40164.
Peterson JM, Seldin MM, Wei Z, Aja S, Wong GW. CTRP3 attenuates diet-induced hepatic steatosis by regulating triglyceride metabolism. Am J Physiol Gastrointest Liver Physiol. 2013;305:G214–24.
Andrikopoulos S, Blair AR, Deluca N, Fam BC, Proietto J. Evaluating the glucose tolerance test in mice. Am J Physiol Endocrinol Metab. 2008;295:E1323–32.
Acknowledgements
Funding support for the SAGE was provided through the NIH Genes, Environment and Health Initiative (GEI) Grant U01 HG004422. SAGE is one of the GWAS funded as part of the GENEVA under GEI. Assistance with phenotype harmonization and genotype cleaning, as well as with general study coordination, was provided by the GENEVA Coordinating Center (Grant U01 HG004446). Assistance with data cleaning was provided by the National Center for Biotechnology Information. Support for collection of datasets and samples were provided by COGA (Grant U10 AA008401), COGEND (Grant P01 CA089392), and FSCD (Grant R01 DA013423). Funding support for genotyping, which was performed at the Johns Hopkins University Center for Inherited Disease Research, was provided by NIH GEI Grant U01HG004438, NIAAA, NIDA, and the NIH contract “High throughput genotyping for studying the genetic contributions to human disease” (HHSN268200782096C). The datasets used for the analyses described in this manuscript were obtained from dbGaP at http://www.ncbi.nlm.nih.gov/projects/gap/cgibin/study.cgi study_id = phs000092.v1.p1 through dbGaP accession number phs000092.v1.p.1. We thank Dr. Xinxin Ding for the cyp2a5−/− mice, Dr. Jerome Lasker for anti-CYP2E1 IgG, and Dr. Risto Juvonen for anti-CYP2A5 IgG.
Funding
This study is partially supported by USPHS grants AA-024723 from the National Institute on Alcohol Abuse and Alcoholism and National Institute of Health grant C06RR0306551.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Wang, K., Chen, X., Ward, S.C. et al. CYP2A6 is associated with obesity: studies in human samples and a high fat diet mouse model. Int J Obes 43, 475–486 (2019). https://doi.org/10.1038/s41366-018-0037-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-018-0037-x