Abstract
Some long noncoding RNAs (lncRNAs), which harbor microRNAs in their gene sequence and are also known as microRNA host gene derived lncRNAs (lnc-MIRHGs), play a dominant role alongside miRNAs, or both perform biological functions synergistically or independently. However, only a small number of lnc-MIRHGs have been identified. Here, multiple liver injury datasets were analyzed to screen and identify the target lncRNA Mir122hg. Mir122hg was mainly enriched in liver tissues with human-mouse homology. In both CCl4-induced acute liver injury and Dgal/LPS-induced fulminant liver failure in mice, Mir122hg was sharply downregulated at the early stage, while a subsequent significant increase was only found in the CCl4 group with liver recovery. Overexpression and silencing assays confirmed that Mir122hg played a protective role in acute injury by promoting hepatocyte proliferation in vivo and in vitro. Consistent with the results of gene enrichment analysis, Mir122hg binding to C/EBPα affected its transcriptional repression, promoted gene transcription of downstream chemokines, Cxcl2, Cxcl3, and Cxcl5, and exerted pro-proliferative effects on hepatocytes through activation of the AKT/GSK-3β/p27 signaling pathway by CXC/CXCR2 complexes. This study identifies a novel lncRNA with protective effects in acute liver injury and demonstrates that the binding of Mir122hg-C/EBPα promotes hepatocyte proliferation via upregulation of CXC chemokine and activation of AKT signaling.
Similar content being viewed by others
Introduction
Liver injury is a common pathological basis for the development and progression of many liver diseases and can be induced by various pathogens, including viruses, bacteria, alcohol, drugs, metabolism, and trauma, manifested as inflammatory cell infiltration, cytokine release, and cell death1. Severe acute attack resulting in massive hepatocyte (HC) death and shortages of regenerated HCs can lead to acute liver failure, multiple organ failure, and even death2. The prognosis of acute liver injury and liver failure is closely related to the degree of cell death and regeneration of residual HCs3.
Long noncoding RNAs (lncRNAs) are a class of RNAs >200 nt with no protein-coding potential. Compared to messenger RNAs (mRNAs), lncRNAs are less conserved in species but have cell line-, tissue- or development-specific expression properties4. Growing evidence shows that many lncRNAs are aberrantly altered in liver disease and can regulate cellular metabolism, inflammatory response, autophagy, proliferation and apoptosis by direct or indirect regulation of key protein-coding genes5. For example, lncRNA-H19, which has been reported to have significant regulatory roles in embryonic growth, tumorigenesis, and cell differentiation, can also promote hepatic steatosis through the expression of MLXIPL, a transcription factor that induces lipogenesis and is involved in the development of hepatic lipid metabolism disorders6. In studies of acute liver injury, deletion of TUG1 protected against lipopolysaccharide (LPS)-induced hepatocyte inflammation by regulating miR-140/TNF7. In a fulminant liver failure mouse model induced by D-galactosamine (Dgal)/LPS, overexpressed lncRNA-NEAT1 resulted in elevated apoptosis and impaired hepatocyte proliferation8.
Some lncRNAs are generated from genomic loci that contain microRNAs (miRNAs), and such lncRNAs with miRNA sequences in their genes are also called miRNA-host-gene-derived lncRNAs (lnc-MIRHGs). Approximately 17.5% of miRNAs are produced from lnc-MIRHGs, which are usually thought to be precursor genes of miRNAs or nonfunctional byproducts of miRNA processing9. However, recent studies have identified many lnc-MIRHGs that play independent roles in a variety of key biological processes10. For example, MIR100HG produces spliced and stable lncRNAs that are significantly upregulated in the G1 phase of the cell cycle; it serves as a binding platform for the interaction between HuR and target mRNAs, regulating cell proliferation regardless of the level of miR1009.
In this study, we identified the mouse miR122 host-gene-derived lncRNA (Mir122hg), which was further verified to be homologous to human MIR122HG. In CCl4-treated mice, overexpression of Mir122hg attenuated liver injury by promoting hepatocyte proliferation; consistently, exacerbated liver injury was found when Mir122hg was silenced. miR122 levels were unaffected by overexpression or silencing of Mir122hg. The possible regulatory mechanism of Mir122hg was its binding and inhibition of the transcription factor CCAAT enhancer-binding protein alpha (C/EBPα), promoting the transcription of downstream chemokines and the activation of protein kinase B (AKT) signaling, which might provide new molecular mechanisms for biomedical and therapeutic applications in liver injury.
Materials and methods
RNA-seq data and gene enrichment analysis
Previously published RNA-seq datasets related to liver injury, namely, the paracetamol (APAP)-induced liver injury dataset (GSE111828), partial hepatectomy dataset (GSE125007), and bacterial infection-induced liver injury dataset (GSE122741) (NCBI Gene Expression Omnibus database, GEO), were analyzed using molecular biology methods. Sequence Read Archive files were uncompressed to complete fastqc quality testing, and the Spliced Transcripts Alignment to a Reference software11 was used for accurate alignment to obtain the read counts of each gene. R software with the edgeR package was used to analyze the raw gene count data. The differentially expressed genes (DEGs) were screened under the condition of |log2FoldChange | >2, adj. p value <0.05, and the DEGs in all three datasets displayed an overlap region in the Venn diagram. Genes closely associated with the target gene were screened under the conditions of |correlation coefficient R | > 0.5 and p value <0.05, and Gene Ontology analysis (GO) and Kyoto Encyclopedia of Genes and Genomes pathway analysis (KEGG) were performed by R software with the clusterprofiler package.
Animals
All 8-week-old male Balb/c mice were purchased from the Institute of Experimental Animal Science, Chinese Academy of Sciences (Beijing, China). All animals were housed in a pathogen-free animal house at Tianjin Medical University on a 12 h light/12 h dark cycle. This study was conducted in strict accordance with approved guidelines and was approved by the Animal Care and Use Committee of Tianjin Medical University.
Acute liver injury induced by carbon tetrachloride (CCl4) and D-gal/LPS
Mice were injected intraperitoneally with CCl4 (1 mg/kg) for the acute liver injury model, and different groups of mice were sacrificed at 6 h, 12 h, 24 h, 48 h, 96 h and 8 d after injection. Mice were injected intraperitoneally with Dgal (700 mg/kg)/LPS (100 µg/kg) for the fulminant liver failure model, and different groups of mice were sacrificed at 2 h, 4 h and 6 h after injection. Liver tissues and venous blood were taken for analysis.
Rapid amplification of cDNA ends (RACE)
RACE experiments were performed with the Smart RACE cDNA Amplification Kit (Clontech, Palo Alto, CA, USA) according to the manufacturer’s instructions, as described previously12. The 5′ and 3′ specific primers for Mir122hg are shown in Supplementary Table 1. PCR amplification, linearization of the vector pRACE plasmid, cloning and sequencing were performed.
Nuclear-cytoplasmic fractionation
Cytoplasmic and nuclear RNA extractions from primary HCs and AML12 (mouse immortalized hepatocyte cell) were performed with the PARIS™ Kit Am1921 (Invitrogen, Grand Island, NY, USA) according to the manufacturer’s instructions as described previously12.
Treatment with adeno-associated virus 8 (AAV8) in vivo
Mice were injected with control AAV8-TBG (Addgene, Beijing Zhongyuan, Ltd. Beijing, China) or overexpression AAV8-Mir122hg via the tail vein (1 × 1011 viral particles/mouse) for 14 d. Control AAV8-sh-Control (Addgene, Beijing Zhongyuan, Ltd. Beijing, China) or gene silencing AAV8-sh-Mir122hg were injected via the tail vein (1 × 1011 viral particles/mouse) for 14 d. Primers for adeno-associated viral plasmid construction are shown in Supplementary Table 2.
Culture and processing of primary HCs and hepatocyte cell lines
Primary HCs were isolated from mice as previously described12 and were cultured in Dulbecco’s modified Eagle’s medium (DMEM, Invitrogen, Camarillo, CA) supplemented with 10% fetal bovine serum (FBS, Gibco, Gaithersburg, MD, USA), 100 units/ml penicillin and 100 μg/ml streptomycin. AML12 and L02 (human immortalized hepatocytes) and HepG2 and Huh7 (human hepatoma cells) were purchased from the Shanghai Cell Bank of the Chinese Academy of Sciences, and the base media were DMEM-F12 (AML12) and Roswell Park Memorial Institute-1640 (Invitrogen, Camarillo, CA) (L02) and DMEM (HepG2 and Huh7) with the same FBS and antibiotics as above. In addition, AML12 cells were cultured with 1× insulin-transferrin-sodium selenite media supplement (Sigma‒Aldrich, St. Louis, MO, USA). Primary HCs and AML12 cells were treated in vitro with tumor necrosis factor alpha (TNFα, 10 ng/ml) (Sigma‒Aldrich, St. Louis, MO, USA) for 12–24 h to induce cell injury. The AKT signaling pathway-specific blocker MK2206 (10 ng/ml) (Med Chem Express, Danvers, USA) was used in primary HCs for 24 h, or the CXC receptor 2 (CXCR2) antagonist SB225002 (10 ng/ml) (Med Chem Express, Danvers, USA) was used in primary HCs for 48 h in the signaling-blockade experiments.
Treatment with lentivirus (LV) or small interfering RNA (si) in vitro
For gene overexpression, cultured cells were infected with LV-Mir122hg or LV-Control for 48 h (1 × 109 viral particles/well). Primers for LV plasmid construction are shown in Supplementary Table 2. For gene silencing, cultured cells were treated with si-Mir122hg, si-Cebpa, si-MIR122HG, or si-Negative Control (si-NC) (GenePharma, Shanghai, China) transfected by Lipofectamine Max (Invitrogen, Carlsbad, CA) for 36 h. The siRNA sequences are shown in Supplementary Table 3.
Serum enzyme assay
Serum levels of alanine aminotransferase (ALT), aspartate aminotransferase (AST), and lactate dehydrogenase (LDH) were measured using a commercial test kit (Nanjing Jiancheng Corp., Nanjing, China) according to the manufacturer’s protocol.
Standard polymerase chain reaction (PCR) and quantitative real-time PCR (qPCR)
Total RNA was extracted using TRIzol reagent (Takara, Dalian, China). RNA was treated with DNase I (Takara, Dalian, China) to remove potential chromosomal DNA contamination prior to cDNA synthesis. First-strand cDNA was synthesized using random primers and AMV Reverse Transcriptase (Thermo Fisher Scientific, Basingstoke, UK), and miRNA first-strand cDNA was synthesized using Tailing Reactions (Sangon Biotech, Shanghai, China). Standard PCRs were performed using Taqase (CoWin Biosciences, Beijing, China). The target PCR products were compared by agarose gel electrophoresis. For qPCR, all reactions were performed according to the manufacturer’s instructions with SYBR Green master mix (Takara, Dalian, China). The primer sequences used for PCR are shown in Supplementary Table 5.
Western blot (WB) analysis
WB analysis was performed as described previously12. The antibodies against PCNA(Cell Signaling, 13110), HMGB1(Cell Signaling, 6893), LC3B(Cell Signaling, 12741), p62(Cell Signaling, 5114), Becline(Cell Signaling, 3495), BAX(abcam, ab32503), BCL2(Proteintech, 12789-1-AP), Cas8(abcam, ab25901), PARP1(Proteintech, 13371-1-AP), AKT(Cell Signaling, 9272), p-AKT(ser473) (Cell Signaling, 4060), p-AKT(thr308) (Cell Signaling, 13038), p-PTEN(ser380)(Cell Signaling, 9551), p-GSK-3β(ser9) (Cell Signaling, 5558), p-c-Raf(ser259) (Cell Signaling, 9421), mTOR(Cell Signaling, 2983), p-mTOR(ser2448) (Cell Signaling, 5536), p27(abcam, ab32034), p-p27(s10)(abcam, ab62364), CDK2(abcam, ab32147), CDK4(Santa Cruz, sc-23896), CyclinB1(abcam, ab181593), CyclinD1(abcam, ab134175), Cyclin E(Santa Cruz, sc-377100), NEDD4 L(Santa Cruz, sc-514954), C/EBPα(Cell Signaling, 8178) and GAPDH(Cell Signaling, 5174) were diluted in TBS containing 5% milk and 0.1% Tween 20. Signals were detected using a chemiluminescence system (Millipore, Bedford, MA, USA).
Histology and immunohistochemistry (IHC)
Hematoxylin-eosin staining (HE) and IHC analysis were performed as described previously12. The antibodies against PCNA (Cell Signaling, 13110), Ki67 (Abcam, ab16667), and HMGB1 (Cell Signaling, 6893) were diluted in phosphate buffered saline (PBS). The reaction products were visualized using diaminobenzidine and monitored by microscopy.
Confocal microscopy
Primary HCs and AML12 cells were replated on polylysine-precoated glass coverslips. The cells were treated with 4% paraformaldehyde overnight at 4 °C, permeabilized in 1% Triton X-100 for 0–60 min, and blocked with 10% goat serum for 30 min at room temperature. p-AKT (Ser473) (Cell Signaling, 4060) and p27 (Abcam, ab32034) were incubated overnight at 4 °C, and unrelated isotype rabbit IgG (Millipore, PP64B) was used as a negative control. Goat anti-rabbit IgG (Invitrogen, Alexa Fluor 594) was incubated for 1 h at room temperature in the dark, and the nuclei were stained with DAPI (5 µg/ml). The cells were finally washed with PBS and mounted with antifade mounting medium (P0126, Beyotime, Shanghai, China). All immunofluorescence was then visualized by a confocal microscope (LSM 700).
RNA‒protein pulldown and mass spectrometry
The target linear DNA template was obtained by PCR amplification, agarose gel electrophoresis, and gel recovery with Mir122hg overexpression plasmid and primers containing the T7 promoter sequence (Supplementary Table 4) according to the manufacturer’s instructions, followed by application of the TranscriptAid T7 High Yield Transcription Kit (Thermo Fisher Scientific, Basingstoke, UK) for transcription in vitro to obtain the sense and anti-sense RNA of Mir122hg. The PierceTM RNA 3′ END Desthiobiotinylation Kit (Thermo Fisher Scientific, Basingstoke, UK) was used to perform desthiobiotinylation ligation to the 3′ ends of the sense and antisense RNA. The PierceTM Magnetic RNA‒Protein Pull-Down Kit (Thermo Fisher Scientific, Basingstoke, UK) was then used to perform the RNA‒Protein pulldown experiments. Briefly, 50 pmol biotin-labeled RNA was incubated with 50 µl magnetic beads at room temperature with mixing for 30 min, and the supernatant was removed on a magnetic separator. Nuclear protein lysate from primary HCs, for which nuclear-cytoplasmic protein fractionation was performed with a PARIS™ Kit Am1921, was added and incubated at 4 °C with mixing for 60 min. The target RNA-pulldown proteins were obtained after washing and elution, followed by electrophoresis with polyacrylamide gels and silver staining according to the manufacturer’s instructions (Silver-stain Kit, Beyotime Biotechnology, Shanghai, China). Gel bands of interest were subjected to in-gel digestion as described previously13. Then, dehydration in acetonitrile and drying in a SpeedVac (Thermo Fisher Scientific, Basingstoke, UK), trypsin digestion with 50 mM ammonium bicarbonate (Promega Biotech Co., Ltd. China) and desalting with a μ-C18 Ziptip were performed.
Mass spectrum: The digested samples were injected into a Nano-LC system (EASY-nLC 1000, Thermo Fisher Scientific, Basingstoke, UK). Each sample was separated by a C18 column at a flow rate of 300 nL/min. The HPLC eluate was electrosprayed directly onto an Orbitrap Q-Exactive mass spectrometer (Thermo Fisher Scientific, Basingstoke, UK). For the MS1 survey scan, the automatic gain control target was 1e6 with a resolution of 70,000. MS2 spectra were acquired at a resolution of 17,500. MS/MS data were retrieved using Proteome Discoverer software, and the overall false discovery rate of peptides was less than 1%. Proteins with a score <2 were removed.
RNA binding protein immunoprecipitation assay (RIP)
RIP was performed using the Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore, Bedford, MA, USA) according to the manufacturer’s instructions. Hepatocyte suspensions were isolated from mouse livers and washed twice with precooled PBS. Complete RIP lysis buffer with the same volume of cell pellets was added, mixed gently, incubated on ice for 5 min, transferred to –80 °C, and centrifuged at 4 °C after one freeze‒thaw cycle. Ten microliters of supernatant was transferred to a new EP tube as input, and another 100 µl of supernatant was transferred to the antibody (C/EBPα, Proteintech, 18311-1-AP, 5 µg, and negative control IgG, 5 µg) complex with 900 µl of RIP buffer and incubated overnight at 4 °C with mixing. Afterward, the supernatant was removed on a magnetic separator and washed 6 times repeatedly, followed by elution to purify the RNA for detection.
Chromatin immunoprecipitation (ChIP)
ChIP was performed using the ChIP Assay Kit (Beyotime Biotechnology, Shanghai, China) according to the manufacturer’s instructions. Briefly, primary HCs were treated with 1% formaldehyde for chromatin cross-linking and terminated with glycine. The cells were washed with PBS, followed by resuspension in 200 µl of SDS lysis buffer and phenylmethylsulfonyl fluoride. Samples were treated on ice with fifteen 10-s pulses for ultrasonic shearing at a power setting of 30% and centrifuged at 14000 × g for 10 min. The supernatant was transferred to a new EP tube. Then, 20 µl of supernatant was transferred as input, and the remaining supernatant was added to 1800 µl ChIP Dilution Buffer and incubated with primary antibody (C/EBPα, Proteintech, 18311-1-AP, 5 µg, and negative control IgG, 5 µg) overnight at 4 °C. Then, 60 µl of Protein A/G Agarose was added and incubated with mixing at 4 °C for 1 h. After multiple washes, DNA was eluted and recovered with phenol‒chloroform methods. Primers for ChIP experiments can be found in Supplementary Table 6.
Human liver tissues
Liver tissues from patients with liver failure (n = 6) and healthy liver tissues from patients with hepatic hemangioma (n = 6) were obtained from the Liver Transplant Department of Tianjin First Central Hospital. Detailed clinical information of the 12 patients can be found in Supplementary Table 7. This study was approved by the Ethics Committee of Tianjin First Central Hospital, and individual permission was obtained through a standard informed consent procedure. This investigation was in accordance with the Declaration of Helsinki regarding the use of human tissues.
Statistical analysis
Graphs were plotted using GraphPad Prism software (La Jolla, CA, USA), and WB bands were quantified by ImageJ software (National Institutes of Health, Bethesda, MD, USA). Gene enrichment analysis was performed by R software (version 3.6.3)14 (Foundation for Statistical Computing, Vienna, Austria, https://www.R-project.org) with the clusterprofiler package. The data are shown as the mean ± SD of at least triplicate experiments. For statistical analysis of RNA levels, data were calculated using the 2 − ΔΔCt method. Differences between the two groups were analyzed by a two-tailed Student’s t test, and p < 0.05 was considered statistically significant.
Results
Identification and expression profiles of Mir122hg in acute liver injury
To investigate the role of lncRNAs in acute liver injury, we analyzed the APAP-induced, partial hepatectomy, and bacterial infection-induced liver injury RNA-seq datasets from the GEO database. The expression heatmap of DEGs in these datasets is shown in Fig. 1a. After intersection of the DEGs, we obtained 176 differentially expressed genes, among which 16 were lncRNAs (Fig. 1b). The 16 lncRNAs were then detected in CCl4- and Dgal/LPS-induced mouse models, and Gm29966 showed a consistent decreasing trend in both models (Supplementary Fig. 1a-d) and the largest enrichment in liver tissues (Supplementary Fig. 1e). Therefore, lncRNA-Gm29966 was selected as the target gene for subsequent study. Sequence search and genome-wide localization from the Ensembl Genome Browser database (www.ensembl.org/) revealed that the putative Gm29966 is an intergenic lncRNA with two transcripts, 001 and 002, and transcript 002 covers the full length of miR122 (Supplementary Fig. 2a). Therefore, we hypothesized that Gm29966 was the miR122 host-gene-derived lncRNA and named it Mir122hg.
a Heatmap of DEGs from the APAP-induced, partial hepatectomy, and bacterial infection-induced liver injury RNA-seq datasets from the GEO database, with red representing high expression and green representing low expression. b DEGs in three datasets displayed an overlapping region in the Venn diagram, and 176 genes, including 16 lncRNAs, were differentially expressed in all three datasets. c–e Mice in each group (n = 8 per group) were sacrificed at 6 h, 12 h, 24 h, 48 h, 96 h and 8 d after CCl4 treatment, and the livers were taken for photographing and HE staining c. Venous blood was taken for the detection of serum enzymes d. RNA levels of Mir122hg-001 and 002 in the liver were detected by qPCR e. f–h Mice in each group (n = 8 per group) were sacrificed at 2 h, 4 h and 6 h after Dgal/LPS treatment, and the livers were taken for photographing and HE staining f; venous blood was taken for the detection of serum enzymes g; RNA levels of Mir122hg-001 and 002 in the liver were detected by qPCR h. The data are shown as the mean ± SD of at least triplicate experiments. *p < 0.05 vs. Control.
In the CCl4-induced mouse model, slight liver injury with partial hepatocyte swelling was already visible in the liver tissue morphology staining at 6 h posttreatment, together with a slight enhancement of serum ALT and AST. Liver injury continued to worsen at 12 h and peaked at 24 h, with high levels of serum ALT, AST, and LDH, and HE staining showed large areas of hepatocyte necrosis with fusion and massive inflammatory cell infiltration. At 48 h, liver damage began to decrease slightly, and at 96 h-8 d, significantly smaller areas of hepatocyte necrosis and a basic return to normal enzyme levels were found (Fig. 1c, d, Supplementary Fig. 2b). In this mouse model from the onset to recovery of acute liver injury, both Mir122hg-001 and 002 exhibited consistent expression changes in the liver, showing a progressive decrease at 6 h, 12 h and 24 h after CCl4 treatment followed by a rapid rebound to above-normal levels at 48 h and a subsequent recovery to normal levels at 96 h and 8 d (Fig. 1e).
In the Dgal/LPS-induced mouse model, no significant changes in morphology or serum enzymes were observed at 2 h and 4 h posttreatment. However, liver damage was sharply aggravated at 6 h, with a dark macroscopic appearance, extensive hepatocyte necrosis and erythrocyte sludge, and significantly elevated serum enzymes (Fig. 1f, g, Supplementary Fig. 2c). In this high-mortality mouse model of fulminant liver failure, Mir122hg- 001 and 002 exhibited consistent changes in the liver tissues, demonstrating a significant decrease at 2 h after Dgal/LPS treatment and a drop to trough values at 4 h and 6 h (Fig. 1h).
Properties and distribution of Mir122hg
The Mir122hg sequences clarified by RACE were slightly different from the data in Ensembl. The full length of Mir122hg-001 was 1095 nt, with only one exon and no introns (Supplementary Fig. 2d). In addition, we identified three new transcripts, Mir122hg-003, 004, and 005 (Fig. 2a).
a Sequences of five transcripts of Mir122hg were detected by RACE. b Expression of Mir122hg, miR122 and the neighboring genes in 10 organs of mice (n = 8) was detected by PCR and agarose gel electrophoresis. c Comparison of five transcripts of Mir122hg in liver tissues by qPCR. d Comparison of Mir122hg-001 and 002 in primary HCs, HSCs, KCs, and LSECs. e, f Primary HCs e and AML12 f were treated with TNFα for 12–24 h in vitro, and the expression levels of Mir122hg-001 and 002 were measured by qPCR. g, h Nuclear-cytoplasmic RNA and protein fractionation were performed in primary HCs g and AML12 h, and the RNA levels of Gapdh, Neat1, and Mir122hg-001 and the protein levels of GAPDH and PCNA were measured. The data are shown as the mean ± SD of at least triplicate experiments. *p < 0.05 vs. Control or Mir122hg-001 or Primary HC.
Gene expression in ten organs of normal mice was determined (Fig. 2b). Mir122hg-001, 002 and miR122-5p were all significantly upregulated in the liver and downregulated in other organs, and transcripts 003, 004 and 005 were only expressed in the liver. The neighboring gene Nedd4l was mainly expressed in the liver, lung, kidney, stomach and small intestine, Alpk2 in the heart and muscle, Malt1 in the spleen and lung, and other genes showed no significant differences in different organs (Fig. 2b, Supplementary Fig. 2e, f). Figure 2c shows that Mir122hg-001 was the most abundant transcript in the liver, and transcripts 003, 004, and 005 were all expressed in trace amounts.
Primary HCs, hepatic stellate cells (HSCs), Kupffer cells (KCs), and liver sinusoidal endothelial cells (LSECs) were isolated from mouse livers. Both Mir122hg-001 and 002 were predominantly expressed in primary HCs (Fig. 2d). TNFα is an injury-induced inflammatory cytokine and is commonly used in studies of hepatocyte injury and death in vitro15,16. On the other hand, it has also been reported to promote the expansion of hepatocytes in 3D culture and enable serial passages and long-term culture for more than 6 months17. Therefore, primary HCs and AML12 cells were treated with TNFα for the cell injury models in vitro.
Consistent with the results in vivo, Mir122hg-001 and 002 in primary HCs were significantly decreased at 12 h and 24 h after TNFα treatment (Fig. 2e). Both Mir122hg-001 and 002 showed low baseline expression in AML12 and were significantly decreased at 24 h, rather than 12 h, after TNFα treatment (Fig. 2f). In addition, Mir122hg- 001 and 002 were predominantly localized in the nucleus in both primary HCs and AML12, and nuclear localization was not changed in CCl4-induced liver injury (Fig. 2g, h and Supplementary Fig. 3a).
As a newly identified gene, we predicted the protein-coding ability of Mir122hg-001 through the online tools CPC2 and CPAT (http://cpc2.cbi.pku.edu.cn/), and the prediction p values were 0.02 and 0.03, respectively. Supported by the characteristic of nuclear localization, it was confirmed as a noncoding gene. Taken together, Mir122hg-001 and 002, with significantly higher expression levels for the former, were both mainly located in the hepatocyte nucleus and consistently changed in models of hepatocyte injury in vivo and in vitro. Therefore, our subsequent study mainly focused on the Mir122hg-001 transcript.
Mir122hg significantly correlates with Ki67 and PCNA in the CCl4-induced model
The first finding that drew our attention to the exact role of Mir122hg in acute liver injury was the levels of proliferation-related Ki67 and PCNA (Fig. 3a, b). In the CCl4-induced model, Ki67 levels in the liver showed a slight decrease followed by a sharp increase (Fig. 3b), which was in strong agreement with Mir122hg-001 (correlation R = 0.60, p < 0.05) (Fig. 3c). In addition, the protein levels, rather than the mRNA levels, of PCNA were consistent with the trend of Ki67 (Fig. 3a and Supplementary Fig. 3b, c), and the correlation R value with Mir122hg-001 was 0.51 (p < 0.05) (Fig. 3d). Consistently, IHC analysis of PCNA further showed that hepatocyte proliferation in the CCl4-induced model was mild at 24 h, obvious at 48 h and 96 h, and largely normal at 8 d (Fig. 3e). In addition, we also observed a significant change in autophagy-related LC3B-II/I, apoptosis-related BAX, and BCL2, suggesting upregulated hepatocyte autophagy and apoptosis in the CCl4-induced model (Fig. 3a).
a–e Mice in each group (n = 8 per group) were sacrificed at 6 h, 12 h, 24 h, 48 h, 96 h and 8 d after CCl4 treatment; protein levels of PCNA, LC3B, BAX and BCL-2 in the liver were detected by WB a; mRNA levels of Ki67 in the liver were measured by qPCR b; and PCNA expression was also detected by IHC e; mRNA levels of Ki67 and protein levels of PCNA in each group were analyzed with Mir122hg-001 by Pearson correlation analysis c, d. The data are shown as the mean ± SD of at least triplicate experiments. *p < 0.05 vs. Control.
In the Dgal/LPS-induced model, no obvious change was found in PCNA and Ki67, and the levels of LC3B II/I and BAX were somewhat reduced at 6 h (Supplementary Fig. 3f–i), suggesting a lack of cell proliferation and inhibition of autophagy and apoptosis in this model. In addition, the changes in miR122 expression in acute liver injury were quite different from those in Mir122hg. In the CCl4-treated group, the expression of miR122-3p/5p was significantly decreased at 48 h and returned to normal at 8 d (Supplementary Fig. 3d, e). In the Dgal/LPS-treated group, the expression levels of miR122-3p/5p were both significantly increased at 4 h and 6 h (Supplementary Fig. 3j, k). In conclusion, the CCl4-induced model comprehensively reflected the process of liver injury and recovery, and changes in Mir122hg were associated with hepatocyte proliferation.
Attenuation of CCl4-induced liver injury in cells overexpressing Mir122hg and aggravation of silencing via the direct regulation of hepatocyte proliferation in vivo and in vitro
To explore the function of Mir122hg, AAV8 was used for overexpression or silencing experiments in the CCl4-induced model. Overexpression of Mir122hg-001 was confirmed by qPCR (Fig. 4a). The upregulation of serum ALT, AST, LDH, protein levels of HMGB1 and liver necrosis at 24 h after CCl4 treatment was significantly attenuated in the overexpression group, indicating that overexpressed Mir122hg attenuated CCl4-induced liver injury (Fig. 4a, b, Supplementary Fig. 4a, c). Both Mir122hg-001 and 002 were downregulated significantly in the silencing experiment in vivo (Fig. 4c, Supplementary Fig. 4d), and the upregulation of serum enzymes, HMGB1 and liver necrosis at 48 h after CCl4 treatment were significantly aggravated, suggesting that silencing Mir122hg aggravated CCl4-induced liver injury (Fig. 4c–e, Supplementary Fig. 4d).
a–e For the overexpression experiment in vivo, mice were injected with AAV8-Mir122hg or control AAV8-TBG for 14 d, followed by CCl4 or oil treatment for 24 h (n = 8 per group); for the silencing experiment in vivo, mice were injected by tail vein with AAV8-sh-Mir122hg or AAV8-sh-control for 14 d, followed by CCl4 or oil treatment for 48 h (n = 8 per group); RNA levels of Mir122hg-001, Hmgb1, Pcna, Ki67 in liver and serum ALT, AST of venous blood were detected a, c; protein levels of PCNA, HMGB1, LC3B, p62 in liver were detected by WB b, d; liver macroscopic pictures, HE staining, and IHC detection of HMGB1, PCNA, Ki67 were performed, respectively e. f–i For the overexpression experiment in vitro, primary HCs were treated with lentivirus LV-Mir122hg or LV-Control for 48 h, followed by TNFα treatment for 24 h; for the silencing experiment in vitro, primary HCs were treated with si-Mir122hg or control si-NC for 36 h, followed by TNFα treatment for 24 h; the RNA levels of Mir122hg-001, Pcna, and Ki67 were measured by qPCR f, g, and the protein levels of PCNA, LC3B, and p62 were detected by WB h, i. The data are shown as the mean ± SD of at least triplicate experiments. *p < 0.05 vs. Control, #p < 0.05 vs. AAV8-TBG + CCl4_24 h or AAV8-sh-Control+CCl4_48 h or LV-Control+TNFα or si-NC + TNFα.
The mRNA levels of the inflammatory factors Il-6, Il-10, and Mcp1 were elevated, except for Tnfα and Il-1β, at 24 h after CCl4 treatment, and inconsistent changes were found in the overexpression group (Supplementary Fig. 4a). Only Tnfα and Mcp1 were significantly elevated at 48 h after CCl4 treatment, and no significant change was found in the silencing group (Supplementary Fig. 4d). Therefore, the role of Mir122hg in the transcription of these inflammatory factors could not be determined.
As previously described, Mir122hg is closely associated with hepatocyte proliferation. The upregulated protein levels of PCNA, autophagy-related LC3BII/I and autophagic substrate SQSTM1 (p62) at 24 h and 48 h after CCl4 treatment, including elevated Ki67 at 48 h, which represented increased hepatocyte proliferation and autophagy, were further promoted by overexpressed Mir122hg and inhibited by silenced Mir122hg in vivo (Fig. 4a–e, Supplementary Fig. 4c).
For further validation in vitro, lentivirus-mediated overexpression and siRNA were used in primary HCs, and the RNA levels of Mir122hg-001 were confirmed by qPCR (Fig. 4f, g, Supplementary Fig. 4f, h). Overexpression of Mir122hg upregulated the mRNA levels of Ki67 and the protein levels of PCNA, LC3BII/I, and p62, while silencing Mir122hg downregulated the above indicators (Fig. 4f–i). Moreover, changes in Ki67, PCNA, LC3B, and p62 regulated by Mir122hg were stable in TNFα-induced hepatocyte injury, further suggesting its direct regulation of hepatocyte proliferation and autophagy (Fig. 4f–i). In addition, Mir122hg did not affect the levels of apoptosis-related proteins, such as Cas8, BAX, and BCL2, both in vivo and in vitro, suggesting that it does not regulate apoptosis (Supplementary Fig. 4b, e, g, i).
Mir122hg regulates the cell cycle via the AKT/GSK-3β/p27 signaling pathway in vivo and in vitro
AKT signaling is a central survival pathway in liver injury due to its important regulatory role in liver regeneration; hepatocytes and nonparenchymal cells activate AKT signaling through cytokine interactions, promoting hepatocyte proliferation and inhibiting cell death in response to drug or toxic stimuli18. Therefore, the changes in AKT and downstream signaling molecules in the CCl4-induced model were further investigated in our study.
Examination of liver tissues after CCl4 treatment showed that the protein levels of phosphorylated p-AKT (ser473), p-AKT (thr308), and p-GSK-3β were increased, the cell cycle inhibitors CDKN1B (p27) and p-p27 were decreased, and the cell cycle-associated proteins CDK2 and Cyclin E were increased, with no significant change in the levels of other AKT-related signaling molecules, such as AKT (pan), mTOR, p-mTOR, p-PTEN, p-c-Raf and cell cycle-related CDK4. AKT signaling and the downstream cell cycle were significantly activated at 24 h and 48 h after CCl4 treatment, which were further promoted by overexpressing Mir122hg and inhibited by silencing Mir122hg in vivo (Fig. 5a, b, Supplementary Fig. 5a, b).
a, b For the overexpression experiment in vivo, mice were injected with AAV8-Mir122hg or control AAV8-TBG for 14 d, followed by CCl4 or oil treatment for 24 h (n = 8 per group). For the silencing experiment in vivo, mice were injected with AAV8-sh-Mir122hg or AAV8-sh-control for 14 d, followed by CCl4 or oil treatment for 48 h (n = 8 per group). The levels of AKT signaling-related p-AKT (Ser473), p-AKT (Thr308), p-GSK-3β, cell cycle-inhibitory p27, p-p27, and the cell cycle-related proteins cyclin E and CDK2 in each group were detected by WB a, b. c–f For the overexpression in vitro, primary HCs were treated with lentivirus LV-Mir122hg or LV-Control for 48 h, followed by TNFα treatment for 24 h; for the silencing experiment in vitro, primary HCs were treated with si-Mir122hg or control si-NC for 36 h, followed by TNFα treatment for 24 h; cellular p-AKT (ser473), p-AKT (Thr308), p-GSK-3β, p27, p-p27, cyclin E, CDK2 levels were detected by WB c, e; p-AKT (ser473) and p27 levels were also detected by confocal microscopy d, f. g–i Primary HCs were treated with lentivirus LV-Mir122hg or LV-Control for 48 h, followed by treatment with TNFα and AKT specific blocker MK2206 or control DMSO for 24 h; Cellular RNA levels of Mir122hg-001, Pcna, Ki67 were measured by qPCR g; protein levels of p-AKT(ser473), p-AKT(thr308), p-GSK-3β, p-c-Raf, mTOR, p-mTOR h, and downstream proteins LC3B, p27, p-p27, Cyclin E, CDK2, and PCNA were detected by WB i. The data are shown as the mean ± SD of at least triplicate experiments. *p < 0.05 vs. Control, #p < 0.05 vs. LV-Mir122hg+TNFα + DMSO.
In experiments of primary HCs in vitro, overexpressed Mir122hg upregulated the protein levels of p-AKT (Ser473), p-AKT (Thr308), p-GSK-3β, Cyclin E, and CDK2 and downregulated the protein levels of p27 and p-p27, while the exact opposite regulation of the above indicators was found when Mir122hg was silenced (Fig. 5c–f). Moreover, changes in p-AKT, p-GSK-3β, Cyclin E, CDK2, p27, and p-p27 regulated by Mir122hg were stable in TNFα-induced hepatocyte injury, further suggesting the direct regulation of Mir122hg in AKT signaling and the cell cycle (Fig. 5c–f). In addition, no obvious changes in the protein levels of other AKT-related signaling molecules, such as mTOR, p-mTOR, p-PTEN, and p-c-Raf, or the mRNA levels of Cdk2, Cdk4, CyclinB1, CyclinD1, and Cyclin E were found in Mir122hg-overexpressing or -silenced cells in vitro (Supplementary Fig. 5c–f).
For further validation, the AKT signaling pathway-specific blocker MK2206 was used to confirm whether Mir122hg regulated the cell cycle via AKT. As shown in Fig. 5h, MK2206 treatment resulted in a significant decrease in the protein levels of p-AKT (Ser473), p-AKT (Thr308), p-GSK-3β, mTOR and p-mTOR and a significant increase in LC3BII/I, suggesting a good blocking effect of AKT signaling (Fig. 5h). Downregulation of p27 and p-p27 and upregulation of Ki67, PCNA, Cyclin E, and CDK2 resulting from overexpression of Mir122hg were significantly suppressed after treatment with MK2206, suggesting that the blockade of AKT signaling eliminated the pro-proliferative effect of Mir122hg in primary HCs (Fig. 5g–i).
In addition, only overexpression experiments were performed in AML12 cells. Overexpression of Mir122hg in AML12 cells did not affect the levels of cell proliferation, autophagy, or the activation of AKT signaling, which was consistent with the results determined by qPCR, WB and confocal microscopy (Supplementary Fig. 6a–d). We speculated that these results were related to the extremely low baseline level of Mir122hg in AML12.
Mir122hg regulates CXC chemokine transcription and activates the AKT signaling pathway through CXCR2
Since Mir122hg is a nuclear-localized and miRNA host-gene-derived lncRNA, how is it involved in regulating AKT signaling? We first investigated whether it acts as a cis-regulator to regulate the transcription of neighboring genes or as a miRNA precursor. However, the results were negative. In CCl4-induced liver injury, correlation analysis showed no significant association between Mir122hg and miR122-3p/5p (Supplementary Fig. 7a–d). Overexpression or silencing of Mir122hg also did not affect the expression levels of miR122-3p/5p and its target gene NEDD4L in vivo and in vitro (Supplementary Fig. 7e, f, 8a–d). In addition, no significant change was found in the expression of the neighboring genes in primary HCs treated with overexpressed or silenced Mir122hg (Supplementary Fig. 8a, b).
To explore the mechanism of Mir122hg, we analyzed the GSE111828, GSE125007, and GSE122741 data again, and 723 genes with significant correlation with Mir122hg were identified for further GO and KEGG analysis (Supplementary Table 8). The resultant cytokine-related signaling pathway caught our attention immediately (Fig. 6a).
a Under the condition of |correlation coefficient R | > 0.5 and p < 0.05, genes that were closely associated with Mir122hg were screened in the three RNA-seq datasets of APAP-, hepatectomy-, and bacterial infection-induced liver injury, and further GO and KEGG analyses were performed by R software with the clusterProfiler package. b, c For the overexpression in vitro, primary HCs were treated with lentivirus LV-Mir122 hg or LV-Control for 48 h, followed by TNFα treatment for 24 h; for the silencing experiment in vitro, primary HCs were treated with si-Mir122 hg or control si-NC for 36 h, followed by TNFα treatment for 24 h; mRNA levels of Cxcr1, Cxcr2, Cxcl3, Cxcl5, Cxcl7, and Cxcl8 were measured by qPCR. d, f Primary HCs were treated with lentivirus LV-Mir122hg or LV-Control for 12 h, followed by treatment with CXCR2 specific antagonist SB225002 or control DMSO for 48 h; cellular RNA levels of Mir122hg-001, Pcna, Ki67 were measured by qPCR d, protein levels of p-AKT(ser473), p-AKT(thr308), p-GSK-3β, p-c-Raf, mTOR, p-mTOR e, and p27, p-p27(s10), Cyclin E, CDK2 and PCNA were detected by WB f. The data are shown as the mean ± SD of at least triplicate experiments. *p < 0.05 vs. Control, #p < 0.05 vs. LV-Control+TNFα or si-NC + TNFα or LV-Mir122hg+ DMSO.
Chemokines are a class of small molecular proteins belonging to cytokines, including CXC, CC, CX3C and C. The CXC chemokines CXCL1-3 and CXCL5-8 act as ligands for CXCR2, and their binding can affect hepatocyte injury, repair and regeneration, which in turn affects the recovery of the entire liver19,20. As shown in Fig. 6b, c, overexpressed Mir122hg in primary HCs significantly upregulated the mRNA levels of Cxcl1, Cxcl2, Cxcl3, Cxcl5, and Cxcl7, and the silencing of Mir122hg significantly downregulated the levels of Cxcl2, Cxcl3, Cxcl5, and Cxcl7. Moreover, changes in the mRNA levels of these CXC chemokines regulated by overexpressed or silenced Mir122hg were stable in TNFα-induced hepatocyte injury, further suggesting the transcriptional regulation of Mir122hg.
Further validation was performed in primary HCs with the specific CXCR2 receptor antagonist SB225002, which mildly upregulated the RNA level of Mir122hg-001, with no significant changes in AKT signaling-related molecules or cell cycle-related proteins. Therefore, treatment with SB225002 alone did not affect the activation of AKT and the cell cycle but might lead to the compensatory elevation in Mir122hg (Fig. 6d–f). On the other hand, upregulation of P-AKT (Ser473), p-AKT (Thr308), p-GSK-3β, Ki67, Cyclin E, PCNA, and CDK2 and downregulation of p27 resulting from overexpression of Mir122hg were significantly inhibited by SB225002 (Fig. 6d–f). These results indicated that CXCR2 blockade inhibited the activating effect of Mir122hg on AKT signaling and the cell cycle in vitro.
Mir122hg binds C/EBPα and affects its transcriptional repression
To confirm the regulatory mechanism, we labeled the full-length transcript of Mir122hg-001 with biotin and incubated it for an RNA pull-down assay with nuclear protein lysate from primary HCs. In addition, mass spectrometry was performed to identify the proteins specifically bound to Mir122hg-001. Among the 11 possible binding proteins, C/EBPα was selected for further validation because of its cellular localization and functional relevance (Fig. 7a). The specific binding of C/EBPα to Mir122hg-001 was further confirmed by reverse RIP experiments (Fig. 7b, c).
a Sense and antisense strand RNA of Mir122hg-001 was obtained by in vitro transcription, and RNA‒protein pulldown experiments were performed with biotin magnetic beads; the protein samples obtained were subjected to silver staining, and mass spectrometry revealed 11 possible binding proteins. b, c qPCR and PCR (including agarose gel electrophoresis) detection of Mir122hg-001 retrieved by anti-C/EBPα-specific antibody compared with Control IgG in the RIP assay with primary HCs. d Primary HCs were infected with LV-Control or LV-Mir122hg, ChIP analyses were performed on the indicated gene promoter regions using an anti-C/EBPα-specific antibody, and enrichment is shown relative to the input. e, f Primary HCs were treated with si-Mir122hg or si-Cebpa or control si-NC for 36 h. Cellular RNA levels of Cebpa, Pcna, Ki67, Cxcr1, Cxcr2, Cxcl1, Cxcl2, Cxcl3, Cxcl5, Cxcl7 and Cxcl8 were measured by qPCR e. Protein levels of C/EBPα, p-AKT (ser473), p-AKT (thr308), p-GSK-3β, p-c-Raf, p27, p-p27 (s10), Cyclin E, CDK2 and PCNA were detected by WB f. The data are shown as the mean ± SD of at least triplicate experiments. *p < 0.05 vs. Control or si-NC.
According to the qPCR and WB results, overexpression or silencing of Mir122hg did not affect the mRNA and protein levels of C/EBPα (Supplementary Fig. 8a–d). Previous studies reported the transcriptional repression of chemokines by the transcription factor C/EBPα21, and further analysis of the JASPAR database (https://jaspar.genereg.net) identified possible binding sites for C/EBPα to the promoter regions of the downstream CXC chemokine genes Cxcl1, Cxcl2, Cxcl3, Cxcl5, and Cxcl7 (Supplementary Fig. 8e). Overexpression of Mir122hg in primary HCs was found to significantly inhibit the binding of C/EBPα to the promoter regions of Cxcl2, Cxcl3, and Cxcl5 in the ChIP assay (Fig. 7d). This suggested that Mir122hg might bind and affect the transcriptional repression of C/EBPα, which in turn promoted downstream transcription of CXC chemokines.
Further validation was performed in primary HCs with si-Cebpa to clarify whether silencing C/EBPα in vitro disrupted the Mir122hg-C/EBPα complex and affected the function of Mir122hg. As shown in Fig. 7e, f, cellular mRNA levels of Ki67, Cxcl3, and Cxcl5 were significantly upregulated after C/EBPα silencing in hepatocytes; protein levels of p-AKT (Ser473), p-AKT (Thr308) and downstream p-GSK-3β were upregulated, p27 levels were decreased, Cyclin E and CDK2 levels were significantly upregulated, suggesting that silenced C/EBPα activated the AKT signaling pathway and cell cycle. Moreover, the inhibition of the AKT signaling pathway, cell cycle, and cell proliferation caused by si-Mir122hg in hepatocytes was abrogated by silencing C/EBPα. The above results strongly suggested that Mir122hg exerts its regulatory effects through C/EBPα.
Identification of the human homologous gene MIR122HG
Human MIR122HG was reported in only one study22. The neighboring genes of MIR122HG are the same as those of mouse Mir122hg in the Ensembl database; between them, the gene sequence homology is as high as 21%, and the homology region in the genome is where the miR122 gene is located. Gene expression for each organ in healthy people was obtained from the GTEx database (https://gtexportal.org/), and MIR122HG was also found to be significantly enriched in the liver (Supplementary Fig. 9a). MIR122HG levels in liver tissues of six patients with liver failure were significantly lower than those in healthy liver tissues (Supplementary Fig. 9b). The baseline levels of MIR122HG in L02 and HepG2 cells were much lower than those in Huh7 cells (Supplementary Fig. 9c), which were further selected for silencing experiments in vitro. Silencing MIR122HG inhibited cell proliferation, as shown by downregulated mRNA and protein levels of CyclinD1 and protein levels of PCNA (Supplementary Fig. 9d, e). Taken together, these results were sufficient to demonstrate the human-mouse homology of MIR122HG.
Discussion
In this study, we first identified the homologous Mir122hg in mice. Consistent with the hepatic specificity of miR122, Mir122hg showed characteristics of hepatocyte enrichment and nuclear localization and was expressed at low levels in other organs. MiRNAs are 21–22 nt-long ribonucleic acids that often play an important role in regulating biological processes, including the cell cycle, differentiation, development, and immune response, by inhibiting mRNA translation or regulating mRNA stability23. As the major miRNA in the liver, miR122 has been extensively studied and shown to be involved in the regulation of various hepatic processes, including cholesterol and lipid metabolism, mitochondrial function, control of circadian rhythms, and polyploidy regulation24. Ashish et al.22 first reported human MIR122HG, elaborating that most lnc-MIRHG do not use the canonical cleavage and polyadenylation pathway but instead use microprocessor cleavage to terminate transcription; however, the biological function of MIR122HG was not investigated in the article.
The lnc-MIRHG loci can produce both miRNAs and lncRNAs, with miRNAs playing a dominant functional role, or both acting synergistically or independently10. Lnc-MIRHGs can function as precursors of miRNAs; for example, cancer cells epigenetically regulate the expression of MIRHGs to alter the levels of miRNAs25. On the other hand, many lnc-MIRHGs perform functions independently. Similar to other types of lncRNAs, lnc-MIRHGs also function via “competing endogenous RNAs”, “DNA interactors” and “protein interactors” mechanisms10. For example, MIR100HG plays an oncogenic role in breast cancer by directly interacting with the promoter of p27 to form RNA‒DNA triplex structures, which attenuate the transcription of p2726. MIR31HG uses its 5′ terminal region to interact with the PAS-B domain of HIF-1α, enhancing the chromatin recruitment of HIF-1α and p300 cofactors to their target gene promoters and promoting the HIF-1 transcriptional network27. To explore the possible biological functions of hepatocyte-enriched Mir122hg, CCl4-induced acute liver injury and Dgal/LPS-induced fulminant liver failure in mice were performed. Mir122hg was first sharply decreased at the initial stage of liver injury, but a subsequent increase was only found in the recovered CCl4 group, not the severe Dgal/LPS group. Mir122hg was significantly correlated with the proliferation-related indices Ki67 and PCNA and was further confirmed to play a protective role in acute liver injury by promoting hepatocyte proliferation in subsequent experiments in vivo and in vitro.
Liver regeneration is an important and necessary process for recovery after liver injury, in which phosphatidylinositol 3-kinase (PI3K)/Akt signaling is the core survival pathway. For example, treatment with the nano-CO donor SMA/CORM2 significantly increased CO levels in the liver and circulatory system and promoted liver regeneration and recovery through the PI3K/AKT/mTOR signaling pathway in APAP-induced liver injury28. It was reported that PRDM4 induced cell cycle arrest and downregulated cellular cyclin E, cyclin D1, and CDK4 levels by upregulating p27 and CDKN1A (p21) via inhibition of AKT signaling29. Therefore, in this study, we focused on the changes in AKT and downstream signaling molecules. Mir122hg promoted hepatocyte proliferation both in vivo and in vitro by activating AKT/GSK-3β signaling, inhibiting p27 expression, and significantly upregulating CDK2, Cyclin E, PCNA, and Ki67 levels during acute liver injury. In addition, our results showed that Mir122hg also promoted hepatocyte autophagy. Paradoxically, activated AKT/mTOR signaling is an important inhibitor of autophagy30, so other potential autophagy-related mechanisms are speculated to be involved, which needs to be further explored in the future.
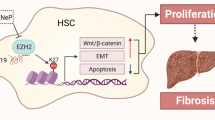
In general, nuclear lncRNAs play either cis-acting or trans-acting roles to modulate chromatin, regulate gene transcription or processing, or organize nuclear structures, and cytoplasmic lncRNAs are known to regulate RNA stability, protein translation, and signal transduction31. In this study, Mir122hg was mainly localized in the nucleus of hepatocytes but did not act through cis-regulatory effects or as a precursor of miR122. Combined with the results of gene enrichment analysis, Mir122hg promoted gene transcription of downstream CXC chemokines in trans and exerted pro-proliferative effects on hepatocytes through activation of the AKT/GSK-3β/p27 signaling pathway by CXC/CXCR2 complexes (Fig. 8).
The CXC chemokine family plays an important role in liver injury and liver regeneration. It comprises 15 ligands, seven of which (CXCL1-3, CXCL5-8) contain a glutamic acid–leucine–arginine motif, enabling their binding to CXCR220. CXC chemokine levels are upregulated in models of partial hepatectomy; blocking CXC or CXCR2, but not CXCR1, reduces liver regenerative capacity; stimulation of cultured hepatocytes with CXC chemokine results in increased proliferation similar to hepatocyte growth factor induction19. It has also been reported that CXCL5 direct treatment of HepG2 cells activates PI3K/AKT and extracellular regulated protein kinase (ERK) signaling with upregulated p-AKT and p-ERK1/2 and promotes cell proliferation, migration and invasion32. Liver parenchyma infusion of CXC chemokines successfully promotes stable regeneration of transplanted hepatocytes in the damaged liver33. These findings fully illustrate the pro-proliferative effect of CXC chemokines on hepatocytes.
The transcription factor C/EBPα belongs to the C/EBP family and has the highest expression levels in liver and adipose tissue. C/EBPα is an important negative regulator of cell proliferation, the cell cycle and gene expression34. In a CCl4-induced rat model of liver fibrosis, C/EBPα was significantly decreased in activated HSCs; overexpressed C/EBPα in vitro inhibited HSC activation, extracellular matrix synthesis, αSMA expression, and lipid droplet formation35. In another study, aging was found to increase susceptibility to alcoholic liver injury in mice and humans by downregulating the neutrophil Sirtuin1-C/EBPα-Mir223 axis and increasing downstream IL-1β, IL-6, TNFα, and CXCL1 secretion21. Consistent with our results, we found transcriptional repression of downstream CXC chemokines by C/EBPα in hepatocytes. Combined Mir122hg-C/EBPα inhibits its binding to the promoter regions of the target genes Cxcl2, Cxcl3, and Cxcl5, which in turn promotes transcription. On the other hand, the mechanism regulating the expression of Mir122hg in acute liver injury is still not clear and needs to be further studied in follow-up experiments.
In conclusion, our work defines a human-mouse homologous gene and reveals a novel mechanism by which Mir122hg-C/EBPα binding promotes CXC chemokine transcription, activates the AKT/GSK-3β/p27 signaling pathway via CXC/CXCR2 complexes, and promotes hepatocyte proliferation. The protective effect of Mir122hg in acute liver injury may become a potential therapeutic target in the future.
References
Gao, Y. et al. CCL22 signaling contributes to sorafenib resistance in hepatitis B virus-associated hepatocellular carcinoma. Pharmacol. Res. 157, 104800 (2020).
Michalopoulos, G. K. & Bhushan, B. Liver regeneration: biological and pathological mechanisms and implications. Nat. Rev. Gastro. Hepat. 18, 40–55 (2021).
Dong, V., Nanchal, R. & Karvellas, C. J. Pathophysiology of acute liver failure. Nutr. Clin. Pract. 35, 24–29 (2020).
Ulitsky, I. Evolution to the rescue: using comparative genomics to understand long non-coding RNAs. Nat. Rev. Genet. 17, 601–614 (2016).
De Vincentis, A. et al. Long noncoding RNAs in nonalcoholic fatty liver disease and liver fibrosis: state-of-the-art and perspectives in diagnosis and treatment. Drug Discov. Today 25, 1277–1286 (2020).
Wang, H. et al. Long non‐coding RNA (lncRNA) H19 induces hepatic steatosis through activating MLXIPL and mTORC1 networks in hepatocytes. J. Cell. Mol. Med. 24, 1399–1412 (2020).
Liu, Q. M. et al. Silencing lncRNA TUG1 Alleviates LPS-Induced Mouse Hepatocyte Inflammation by Targeting miR-140/TNF. Front. Cell. Dev. Biol. 8, 616416 (2020).
Wang, Q. et al. Long noncoding RNA NEAT1 suppresses hepatocyte proliferation in fulminant hepatic failure through increased recruitment of EZH2 to the LATS2 promoter region and promotion of H3K27me3 methylation. Exp. Mol. Med. 52, 461–472 (2020).
Wu, T. et al. Development of diagnostic criteria and a prognostic score for hepatitis B virus-related acute-on-chronic liver failure. Gut 67, 2181–2191 (2018).
Sun, Q., Song, Y. J. & Prasanth, K. V. One locus with two roles: microRNA‐independent functions of microRNA‐host‐gene locus‐encoded long noncoding RNAs. Wires RNA 12, e1625 (2021).
Dobin, A. & Gingeras, T. R. Mapping RNA-seq Reads with STAR. Curr. Protoc. Bioinfor. 51, 11.14.11–11.14.19 (2015).
Zhang, K. et al. The liver-enriched lnc-LFAR1 promotes liver fibrosis by activating TGFβ and Notch pathways. Nat. Commun. 8, 1–16 (2017).
Bai, X. et al. Development of a DNA‐templated peptide probe for photoaffinity labeling and enrichment of the histone modification reader. Proteins Angew. Chem. Int. Ed. Engl. 128, 8125–8129 (2016).
R Development Core Team. R: A language and environment for statistical computing. (R Foundation for Statistical Computing, 2020).
Ezquerro, S. et al. Ghrelin reduces TNF-α–induced human hepatocyte apoptosis, autophagy, and pyroptosis: role in obesity-associated NAFLD. J. Clin. Endocrinol. Metab. 104, 21–37 (2019).
Jing, Z.-T. et al. AKT activator SC79 protects hepatocytes from TNF-α-mediated apoptosis and alleviates d-Gal/LPS-induced liver injury. Am. J. Physiol. Gastrointest. Liver. Physiol. 316, G387–G396 (2019).
Peng, W. C. et al. Inflammatory cytokine TNFα promotes the long-term expansion of primary hepatocytes in 3D culture. Cell 175, 1607–1619 (2018). e1615.
Ozaki, M. Cellular and molecular mechanisms of liver regeneration: Proliferation, growth, death and protection of hepatocytes. Semin. Cell. Dev. Biol. 100, 62–73 (2020).
Van Sweringen, H. L. et al. CXC chemokine signaling in the liver: impact on repair and regeneration. Hepatology 54, 1445–1453 (2011).
Veenstra, M. & Ransohoff, R. M. Chemokine receptor CXCR2: physiology regulator and neuroinflammation controller? J. Neuroimmunol. 246, 1–9 (2012).
Ren, R. et al. Aging exaggerates acute-on-chronic alcohol-induced liver injury in mice and humans by inhibiting neutrophilic sirtuin 1-C/EBPα-miRNA-223 axis. Hepatology 75, 646–660 (2021).
Dhir, A., Dhir, S., Proudfoot, N. J. & Jopling, C. L. Microprocessor mediates transcriptional termination of long noncoding RNA transcripts hosting microRNAs. Nat. Struct. Mol. Biol. 22, 319–327 (2015).
Gebert, L. F. & MacRae, I. J. Regulation of microRNA function in animals. Nat. Rev. Mol. Cell. Biol. 20, 21–37 (2019).
Luna, J. M. et al. Argonaute CLIP defines a deregulated miR-122-bound transcriptome that correlates with patient survival in human liver cancer. Mol. Cell. 67, 400–410 (2017). e407.
Daniunaite, K. et al. Clinical significance of miRNA host gene promoter methylation in prostate cancer. Hum. Mol. Genet. 26, 2451–2461 (2017).
Wang, S. et al. LncRNA MIR100HG promotes cell proliferation in triple-negative breast cancer through triplex formation with p27 loci. Cell. Death. Dis. 9, 1–11 (2018).
Shih, J. W. et al. Long noncoding RNA LncHIFCAR/MIR31HG is a HIF-1α co-activator driving oral cancer progression. Nat. Commun. 8, 1–16 (2017).
Song, B. et al. Nano-designed carbon monoxide donor SMA/CORM2 exhibits protective effect against acetaminophen induced liver injury through macrophage reprograming and promoting liver regeneration. J. Control. Release 331, 350–363 (2021).
Yang, W. T., Chen, M., Xu, R. & Zheng, P. S. PRDM4 inhibits cell proliferation and tumorigenesis by inactivating the PI3K/AKT signaling pathway through targeting of PTEN in cervical carcinoma. Oncogene 40, 3318–3330 (2021).
Lin, P. H. et al. Melatonin activates cell death programs for the suppression of uterine leiomyoma cell proliferation. J. Pineal Res. 68, e12620 (2020).
Sun, Q., Hao, Q. & Prasanth, K. V. Nuclear long noncoding RNAs: key regulators of gene expression. Trends Genet. 34, 142–157 (2018).
Zhou, S. L. et al. Overexpression of CXCL5 mediates neutrophil infiltration and indicates poor prognosis for hepatocellular carcinoma. Hepatology 56, 2242–2254 (2012).
Joshi, M. et al. Chemokine‐mediated robust augmentation of liver engraftment: a novel approach. Stem. Cell. Transl. Med 4, 21–30 (2015).
Cassel, T. N. & Nord, M. C/EBP transcription factors in the lung epithelium. Am. J. Physiol. Lung. Cell. Mol. Physiol. 285, L773–L781 (2003).
Sharma, A. et al. Lipopolysaccharide reverses hepatic stellate cell activation through modulation of cMyb, small mothers against decapentaplegic, and CCAAT/enhancer‐binding protein C/EBP transcription factors. Hepatology 72, 1800–1818 (2020).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Nos. 81870429; 82170630; 32171125).
Author information
Authors and Affiliations
Contributions
Z.Y. drafted the manuscript; Z.Y., Y.L., S.S., B.G., and M.Z. performed the experiments. Z.Y., K.Z. and W.H. analyzed the data; L.Z., C.C., and L.Z. helped to collect the samples. T.H., W.H., and W.J. conceived the study and revised the final manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yu, Z., Li, Y., Shao, S. et al. Protective effect of hepatocyte-enriched lncRNA-Mir122hg by promoting hepatocyte proliferation in acute liver injury. Exp Mol Med 54, 2022–2035 (2022). https://doi.org/10.1038/s12276-022-00881-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-022-00881-2