Abstract
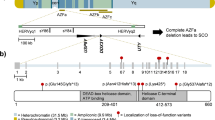
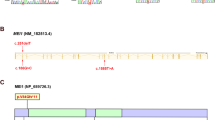
Male infertility pertains to male’s inability to cause pregnancy in a fertile female. It accounts for 40–50% of infertility in human. In the study, presented here, a large consanguineous family of Pakistani origin segregating male infertility in autosomal recessive manner was investigated. Exome sequencing revealed a homozygous frameshift variant [NM_001040108: c.3632delA, p.(Asn1211Metfs*49)] in DNA mismatch repair gene MLH3 (MutL Homolog) that segregated with male infertility within the family. This is the first loss-of-function homozygous variant in the MLH3 gene causing severe oligozoospermia leading to male infertility. Previous studies have demonstrated association of infertility with gene knockout in the mice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Oud MS, Volozonoka L, Smits RM, Vissers LELM, Ramos L, Veltman JA. A systematic review and standardized clinical validity assessment of male infertility genes. Hum Reprod. 2019;34:932–41.
Matzuk MM, Lamb DJ. The biology of infertility: research advances and clinical challenges. Nat Med. 2008;14:1197–213.
Lin YN, Matzuk MM. Genetics of male fertility. Methods Mol Biol. 2014;1154:25–37.
Krausz C, Riera-Escamilla A. Genetics of male infertility. Nat Rev Urol. 2018;15:369–84.
Krausz C, Escamilla AR, Chianese C. Genetics of male infertility: from research to clinic. Reprod. 2015;150:159–74.
World Health Organization. WHO laboratory manual for the examination and processing of human semen. 5th ed. Geneva: WHO Press; 2010.
Howe B, Umrigar A, Tsien F. Chromosome preparation from cultured cells. J Vis Exp. 2014;83:e50203.
Li H, Durbin R. Fast, and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics. 2010;26:589–95.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164.
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;2016:285–91.
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet. 2014;46:310–5.
Lipkin SM, Moens PB, Wang V, Lenzi M, Shanmugarajah D, Gilgeous A, et al. Meiotic arrest, and aneuploidy in MLH3-deficient mice. Nat Genet. 2002;31:385–90.
Firestein R, Nagy PL, Daly M, Huie P, Conti M, Cleary ML. Male infertility, impaired spermatogenesis, and azoospermia in mice deficient for the pseudophosphatase Sbf1. J Clin Invest. 2002;109:1165–72.
Fiedler SE, Sisson JH, Wyatt TA, Pavlik JA, Gambling TM, Carson JL, et al. Loss of ASP but not ROPN1 reduces mammalian ciliary motility. Cytoskeleton (Hoboken). 2012;69:22–32.
Wang TF, Kleckner N, Hunter N. Functional specificity of MutL homologs in yeast: evidence for three Mlh1-based heterocomplexes with distinct roles during meiosis in recombination and mismatch correction. Proc Natl Acad Sci USA. 1999;96:13914–9.
Paquis‐Flucklinger V, Santucci‐Darmanin S, Paul R, Saunieres A, Turc‐Carel C, Desnuelle C. Cloning, and expression analysis of a meiosis specific MutS homolog: the human MSH4 gene. Genomics. 1997;44:188–94.
Santucci‐Darmanin S, Neyton S, Lespinasse F, Saunieres A, Gaudray P, Paquis‐Flucklinger V. The DNA mismatch‐repair MLH3 protein interacts with MSH4 in meiotic cells, supporting a role for this MutL homolog in mammalian meiotic recombination. Hum Mol Genet. 2002;11:1697–706.
Surtees JA, Argueso JL, Alani E. Mismatch repair proteins: key regulators of genetic recombination. Cytogenet Genome Res. 2004;107:146–59.
Her C, Zhao N, Wu X, Tompkins JD. MutS homologues hMSH4 and hMSH5: diverse functional implications in humans. Front Biosci. 2007;12:905–11.
Guarné A, Ramon-Maiques S, Wolff EM, Ghirlando R, Hu X, Miller JH, et al. Structure of the MutL C-terminal domain: a model of intact MutL and its roles in mismatch repair. EMBO J. 2004;23:4134–45.
Mahfouz R, Sharma R, Thiyagarajan A, Kale V, Gupta S, Sabanegh E, et al. Semen characteristics and sperm DNA fragmentation in infertile men with low and high levels of seminal reactive oxygen species. Fertil Steril. 2010;94:2141–6.
Agarwal A, Said TM. Sperm chromatin assessment. In: Gardner DK, Weissman A, Howles CM, Shoham Z, editors. Textbook of assisted reproductive techniques, vol. 1, 4th ed. Boca Raton, Florida: Taylor & Francis Group; 2012. p. 75.
Terribas E, Bonache S, García-Arévalo M, Sánchez J, Franco E, Bassas L, et al. Changes in the expression profile of the meiosis-involved mismatch repair genes in impaired human spermatogenesis. J Androl. 2010;31:346–57.
Gunes S, Al-Sadaan M, Agarwal A. Spermatogenesis, DNA damage and DNA repair mechanisms in male infertility. Reprod Biomed Online. 2015;31:309–19.
Liu HX, Zhou XL, Liu T, Werelius B, Lindmark G, Dahl N, et al. The role of hMLH3 in familial colorectal cancer. Cancer Res. 2003;63:1894–9.
Markandona O, Dafopoulos K, Anifandis G, Messini CI, Dimitraki M, Tsezou A, et al. Single-nucleotide polymorphism rs 175080 in the MLH3 gene and its relation to male infertility. J Assist Reprod Genet. 2015;32:1795–9.
Zhao X, Mu C, Ma J, Dai X, Jiao H. The association of four SNPs in DNA mismatch repair genes with idiopathic male infertility in northwest China. Int J Immunogenet. 2019;46:451–8.
Xu K, Lu T, Zhou H, Bai L, Xiang Y. The role of MSH5 C85T and MLH3 C2531T polymorphisms in the risk of male infertility with azoospermia or severe oligozoospermia. Clin Chim Acta. 2010;411:49–52.
Chen S, Wang G, Zheng X, Ge S, Dai Y, Ping P, et al. Whole-exome sequencing of a large Chinese azoospermia and severe oligospermia cohort identifies novel infertility causative variants and genes. Hum Mol Genet. 2020;29:2451–9.
Olkinuora A, Nieminen TT, Martensson E, Rohlin A, Ristimaki A, Koskenvuo L, et al. Biallelic germline nonsense variant of MLH3 underlies polyposis predisposition. Genet Med. 2019;21:1868–73.
Acknowledgements
We thank the family members for participating in this study.
Funding
This work was supported by the Pakistan Academy of Science, Pakistan (https://www.paspk.org) (PAS-171) to WA.
Author information
Authors and Affiliations
Contributions
SN designed the study, drafted the manuscript, and performed segregation analysis. SH performed exome data analysis. IU and MN performed the clinical tests. BSH and JN collected the samples and analyzed the clinical data. WA designed the study, provided funds, and edited, and finalized the manuscript. All the authors have approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The study was approved by the Institutional Review Board (IRB) of Quaid-i-Azam University, Islamabad, Pakistan (IRB# QAU-171). The family, displaying male infertility, was ascertained from Khyber Pakhtunkhwa (KP) province of Pakistan. Informed written consent to carry out the study was obtained from family members.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Nawaz, S., Ullah, M.I., Hamid, B.S. et al. A loss-of-function variant in DNA mismatch repair gene MLH3 underlies severe oligozoospermia. J Hum Genet 66, 725–730 (2021). https://doi.org/10.1038/s10038-021-00907-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-021-00907-z