Abstract
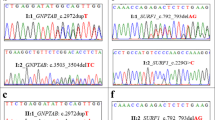
Mucolipidosis (ML) (OMIM 607840 & 607838) is a rare autosomal recessive inherited disorder that occurs due to the deficiency of golgi enzyme uridine diphosphate (UDP)– N-acetylglucosamine-1-phosphotransferase (GlcNAc-phosphotransferase) responsible for tagging mannose-6-phosphate for proper trafficking of lysosomal enzymes to lysosomes. Variants in GlcNAc-phosphotransferase (GNPTAB (α, β subunits) and GNPTG (γ subunits) are known to result in impaired targeting of lysosomal enzymes leading to Mucolipidosis (ML) Type II or Type III. We analyzed 69 Indian families of MLII/III for clinical features and molecular spectrum and performed in silico analysis for novel variants. We identified 38 pathogenic variants in GNPTAB and 5 pathogenic variants in GNPTG genes including missense, frame shift, deletion, duplication and splice site variations. A total of 26 novel variants were identified in GNPTAB and 4 in GNPTG gene. In silico studies using mutation prediction software like SIFT, Polyphen2 and protein structure analysis further confirmed the pathogenic nature of the novel sequence variants detected in our study. Except for a common variant c.3503_3504delTC in early onset MLII, we could not establish any other significant genotype and phenotype correlation. This is one of the largest studies reported till date on Mucolipidosis II/III in order to identify mutation spectrum and any recurrent mutations specific to the Indian ethnic population. The mutational spectrum information in Indian patients will be useful in better genetic counselling, carrier detection and prenatal diagnosis for patients with ML II/III.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Raas-Rothschild A, Pohl S, Braulke T. Multiple enzyme deficiencies: defects in transport: mucolipidosis II alpha/beta; mucolipidosis III alpha/beta and mucolipidosis III gamma. In Mehta AB, Winchester B, editors. Lysosomal storage diseases: a practical guide. Wiley-Blackwell;2012. pp. 121–6..
Kudo M, Bao M, D’Souza A, Ying F, Pan H, Roe BA, et al. The α-and β-subunits of the human UDP-N-acetylglucosamine: lysosomal enzyme phosphotransferase are encoded by a single cDNA. J Biol Chem 2005;280:36141–9.
Filocamo M, Morrone A. Lysosomal storage disorders: molecular basis and laboratory testing. Hum genomics 2011;5:156.
Leroy JG, Cathey SS, Friez MJ. GNPTAB-related disorders. In: GeneReviews®. Seattle: University of Washington; 2019.
Leroy JG, Cathey SS, Friez MJ. Mucolipidosis III alpha/beta. In: GeneReviews®. Seattle: University of Washington; 2012.
Qian Y, Lee I, Lee WS, Qian M, Kudo M, Canfield WM, et al. Functions of the α, β, and γ subunits of UDP-GlcNAc: lysosomal enzyme N-acetylglucosamine-1-phosphotransferase. J Biol Chem 2010;285:3360–70.
Plante M, Claveau S, Lepage P, Lavoie ÈM, Brunet S, Roquis D, et al. A single causal mutation in the N‐acetylglucosamine‐1‐phosphotransferase gene (GNPTAB) in a French Canadian founder population. Clin Genet 2008;73:236–44.
Wang Y, Ye J, Qiu WJ, Han LS, Gao XL, Liang LL, et al. Identification of predominant GNPTAB gene mutations in eastern Chinese patients with mucolipidosis II/III and a prenatal diagnosis of mucolipidosis II. Acta Pharmacologica Sin 2019;40:279–87.
Velho RV, Harms FL, Danyukova T, Ludwig NF, Friez MJ, Cathey SS, et al. The lysosomal storage disorders mucolipidosis type II, type III alpha/beta, and type III gamma: update on GNPTAB and GNPTG mutations. Hum Mutat 2019;40:842–64.
Otomo T, Muramatsu T, Yorifuji T, Okuyama T, Nakabayashi H, Fukao T, et al. Mucolipidosis II and III alpha/beta: mutation analysis of 40 Japanese patients showed genotype–phenotype correlation. J Hum Genet 2009;54:145–51.
Mucolipidosis II alpha/beta. Genetics home reference: your guide to understanding genetic conditions. National Library of Medicine, Bethesda, USA; 2020 https://ghr.nlm.nih.gov/condition/mucolipidosis-ii-alpha-beta#diagnosis
Verma J,C, Thomas D, Sharma S, Jhingan G, Saxena R, Kohli S, et al. Inherited metabolic disorders: prenatal diagnosis of lysosomal storage disorders. Prenat Diagnosis. 2015;35:1137–47.
Eswar N, Webb BM, Marti-Renom A, Madhusudhan MS, Eramian D, Shen M-Y, Pieper U, et al. Curr Protoc Bioinf 2006;15:1–5.
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y. The I-TASSER suite: protein structure and function prediction. Nat Methods. 2015;12:7.
Lovell SC, Davis IW, Arendall WB III, De Bakker PI, Word JM, Prisant MG, et al. Structure validation by Cα geometry: ϕ, ψ and Cβ deviation. Proteins: Struct, Funct, Bioinforma 2003;50:437–50.
Guex N, Peitsch MC. SWISS‐MODEL and the Swiss‐Pdb Viewer: an environment for comparative protein modeling. Electrophoresis. 1997;18:2714–23.
Delano WL. Pymol: An open-source molecular graphics tool. CCP4 Newsletter On Protein Crystallography. 40:82–92.
Qian Y, Van Meel E, Flanagan-Steet H, Yox A, Steet R, Kornfeld S. Analysis of mucolipidosis II/III GNPTAB missense mutations identifies domains of UDP-GlcNAc: lysosomal enzyme GlcNAc-1-phosphotransferase involved in catalytic function and lysosomal enzyme recognition. J Biol Chem 2015;290:3045–56.
Velho RV, De Pace R, Klünder S, Sperb-Ludwig F, Lourenço CM, Schwartz IV. et al. Analyses of disease-related GNPTAB mutations define a novel GlcNAc-1-phosphotransferase interaction domain and an alternative site-1 protease cleavage site. Hum Mol Genet. 2015;24:3497–505.
Aggarwal S, Coutinho MF, Dalal AB, Jain SJ, Prata MJ, Alves S. Prenatal skeletal dysplasia phenotype in severe MLII alpha/beta with novel GNPTAB mutation. Gene. 2014;542:266–8.
Cury GK, Matte U, Artigalás O, Alegra T, Velho RV, Sperb F, et al. Mucolipidosis II and III alpha/beta in Brazil: analysis of the GNPTAB gene. Gene. 2013;524:59–64.
De Pace R, Coutinho MF, Koch‐Nolte F, Haag F, Prata MJ, Alves S, et al. Related Mutations Inhibit the Exit from the endoplasmic reticulum and proteolytic cleavage of G lc NA c‐1‐phosphotransferase precursor protein (GNPTAB). Hum Mutat. 2014;35:368–76.
Tappino B, Chuzhanova NA, Regis S, Dardis A, Corsolini F, Stroppiano M, et al. Molecular characterization of 22 novel UDP‐N‐acetylglucosamine‐1‐phosphate transferase α‐and β‐subunit (GNPTAB) gene mutations causing mucolipidosis types IIα/β and IIIα/β in 46 patients. Hum Mutat. 2009;30:E956–73.
Acknowledgements
The authors are thankful to the patients and their families who volunteered in the study, for their kind cooperation and for allowing us to use their medical and clinical information for the benefit of others. The authors are glad to acknowledge the support provided by the research institution Centre for DNA Fingerprinting and Diagnostics, Hyderabad. This research project was supported by a research grant from the Indian Council of Medical Research (ICMR)-Department of Health Research, Government of India (GIA/31(Vii)/2014-DHR). Ms. Divya Pasumarthi is the recipient of Senior Research Fellowship (F.NO.45/25/2018-HUM/BMS) from the Indian Council of Medical Research (ICMR). We are grateful to the coordinators and all the members of Task force on Lysosomal storage Disorders of the Indian Council of Medical Research (ICMR)-Department of Health Research, Government of India: Dr. Roli Mathur and Dr. Babbanjee.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pasumarthi, D., Gupta, N., Sheth, J. et al. Identification and characterization of 30 novel pathogenic variations in 69 unrelated Indian patients with Mucolipidosis Type II and Type III. J Hum Genet 65, 971–984 (2020). https://doi.org/10.1038/s10038-020-0797-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-020-0797-8
This article is cited by
-
Quaternary diagnostics scheme for mucolipidosis II and detection of novel mutation in GNPTAB gene
Journal of Genetic Engineering and Biotechnology (2021)