Abstract
The phenotype of coeliac disease varies considerably for incompletely understood reasons. We investigated whether established coeliac disease susceptibility variants (SNPs) are individually or cumulatively associated with distinct phenotypes. We also tested whether a polygenic risk score (PRS) based on genome-wide associated (GWA) data could explain the phenotypic variation. The phenotypic association of 39 non-HLA coeliac disease SNPs was tested in 625 thoroughly phenotyped coeliac disease patients and 1817 controls. To assess their cumulative effects a weighted genetic risk score (wGRS39) was built, and stratified by tertiles. In our PRS model in cases, we took the summary statistics from the largest GWA study in coeliac disease and tested their association at eight P value thresholds (PT) with phenotypes. Altogether ten SNPs were associated with distinct phenotypes after correction for multiple testing (PEMP2 ≤ 0.05). The TLR7/TLR8 locus was associated with disease onset before and the SH2B3/ATXN2, ITGA4/UBE2E3 and IL2/IL21 loci after 7 years of age. The latter three loci were associated with a more severe small bowel mucosal damage and SH2B3/ATXN2 with type 1 diabetes. Patients at the highest wGRS39 tertiles had OR > 1.62 for having coeliac disease-related symptoms during childhood, a more severe small bowel mucosal damage, malabsorption and anaemia. PRS was associated only with dermatitis herpetiformis (PT = 0.2, PEMP2 = 0.02). Independent coeliac disease-susceptibility loci are associated with distinct phenotypes, suggesting that genetic factors play a role in determining the disease presentation. Moreover, the increased number of coeliac disease susceptibility SNPs might predispose to a more severe disease course.
Similar content being viewed by others
Introduction
Food antigens do not generally cause a systemic immune response in healthy individuals, but rather lead to the induction of oral tolerance. However, in approximately 1–2% of individuals, the ingestion of dietary gluten results in the development of coeliac disease, an immune-mediated enteropathy [1]. In some patients, coeliac disease develops already in early childhood, while in others, the tolerance to gluten is lost at a considerably older age. A hallmark of the disease is gluten-dependent small bowel mucosal damage that ranges from minor inflammatory changes to total villous atrophy with crypt hyperplasia [2]. The enteropathy is often coupled with malabsorption and gastrointestinal symptoms, such as diarrhoea, but the disease also presents with diverse extraintestinal manifestations affecting various organs including skin and musculoskeletal system [2, 3]. This multifaceted clinical picture is further diversified by several associated conditions, such as type 1 diabetes and autoimmune thyroidal disease [2].
The predisposition for coeliac disease runs in families, and relatives of coeliac patients are at an increased risk [4]. The disease susceptibility is largely conferred by human leucocyte antigen (HLA) haplotypes encoding for DQ2 or DQ8 heterodimers, which are necessary but not sufficient for disease development [2]. Genome-wide association (GWA) and follow-up studies have identified 94 SNPs in 43 non-HLA risk loci that by themselves modify the disease risk modestly [5,6,7,8]. However, combining their independent cumulative effects into a genetic risk score (GRS) improves the prediction of the disease risk [9]. Moreover, by combining the loci of small effect, including those that do not achieve genome wide significance into a polygenic risk score (PRS), it has been possible to examine the influence of several thousands of risk alleles to coeliac disease susceptibility [7, 10].
Currently, there’s only limited information about the connection of the non-HLA risk SNPs with different coeliac disease phenotypes. Therefore, the aim of this study was to investigate whether previously identified coeliac disease-susceptibility SNPs are associated with distinct disease phenotypes and to gain insight into possible biological pathways and processes underlying the identified genotype–phenotype associations. Moreover, the objective was to study whether GRS or PRS give clues to the factors contributing to the clinical heterogeneity of the disease.
Materials and methods
Study population
Altogether 1048 biopsy-proven coeliac disease patients were recruited by a nationwide newspaper advertisements and the assistance of the Finnish Coeliac Society in Tampere University and Tampere University Hospital 2005–2010. The study protocol was approved by the Regional Ethics Committee of Tampere University Hospital and all study subjects or young children’s legal guardians gave written informed consent. All participants were interviewed either by a physician or by a study nurse with expertise in coeliac disease. The structured interviews included questions on coeliac disease diagnosis, symptoms at the time of diagnosis and in childhood, and associated medical conditions. All relevant medical information was confirmed from the patient records. Whole blood samples were drawn for genetic analysis. As the presence of relatives can lead to genetic bias (inflation of type 1 error), the current study considered only one coeliac case per family resulting in 625 cases grouped into relevant phenotypes (Table 1). The median age of the patients was 41 ranging from 0.5 to 79 years. The individual was selected randomly among the family members with full genotype data available. The healthy controls (n = 1817) with information on gender and HLA-type were obtained from the FINRISK and Health 2000 population cohorts [11]. Ethical committee’s approvals were available from the National Public Health Institute’s ethical committee and the Ethical committee in epidemiology and public health at the hospital district of Helsinki and Uusimaa.
Genotyping and quality control
HLA-DQ typing was performed with the TaqMan chemistry, SSP DQB1 and DRB1 low-resolution kit (Olerup SSP AB, Saltsjöbaden, Sweden), or DELFIA® Coeliac Disease Hybridization Assay Kit (PerkinElmer Life and Analytical Sciences, Wallac Oy, Turku, Finland) [12, 13]. For patients and controls missing HLA-typing results the missing GWAS SNPs were further imputed in full HLA haplotypes with HIBAG R package [14].
All participants had been genotyped for a SNP set on an Illumina 610-Quad BeadChip array (Illumina Inc., San Diego, CA, USA) [5]. The SNPs with established risk for coeliac disease were selected. We included the 39 non-HLA coeliac disease SNPs identified in the earlier GWAS [5] and directly genotyped in our samples. As most of the SNPs identified in Trynka et al. study [6] were either not available at our array or had low genotyping frequency, they were imputed (further details under the “imputation” section).
Genotypes were stored in BC│Genome v.4.0 (BC│Platforms, Espoo, Finland) and quality checks performed [15]. The genotyped and imputed SNPs passed the quality control (QC) filters for missing genotype rate < 5%, missing genotype rates differences between the cases and controls (<3%), and minor allele frequency (MAF > 1%) [16]. All markers were (P > 1 × 10−6) in HWE in the controls [6, 16]. Allelic associations with phenotypes were tested in a case–control analysis and within cases, further detailed under the ‘statistical analysis’ section.
Imputation
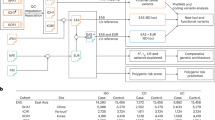
The genotypes of 94 coeliac disease risk variants [5, 6] were selected to be phased and imputed by using a Finnish population-specific panel of 3775 high-coverage (25–30×) whole-genome sequences (SISu v3) as here described: dx.doi.org/10.17504/protocols.io.nmndc5e. SISu v3 panel was generated at the Broad Institute of MIT and Harvard and at the McDonnell Genome Institute at Washington University; and jointly processed at the Broad Institute.
Genotyping data produced with our chip platform were lifted over to genome build version 38 (GRCh38/hg38) following the protocol described here: https://doi.org/10.17504/protocols.io.nqtddwn. In sample-wise quality control (QC), samples with sex discrepancies, high genotype missingness (>5%), excess heterozygosity (±4SD) and non-Finnish ancestry were removed. In variant-wise QC, variants with high missingness (>2%), deviation from Hardy–Weinberg equilibrium (HWE) (P < 1e-6) and minor allele count <3 were removed. Pre-phasing (default parameters, except the number of conditioning haplotypes was set to 20,000) and phasing of genotyped data were performed with Eagle 2.3.5 (https://data.broadinstitute.org/alkesgroup/Eagle/)
Imputation was carried out with Beagle 4.1 (version 08Jun17.d8b, https://faculty.washington.edu/browning/beagle/b4_1.html) as described in the following protocol: dx.doi.org/10.17504/protocols.io.nmndc5e. Variant callset was produced by following GATK best-practices. Genotype-, sample- and variant-wise QC was applied in an iterative manner by using the Hail framework v0.1 [https://github.com/hail-is/hail].
In the post-imputation QC, the SNPs with good imputation quality metrics (INFO score ≥0.8) and MAF > 0.01 were included. The VCF files with the genotypes probabilities dosages were handled by using bcftools (https://samtools.github.io/bcftools/)
Functional identification of enriched pathways related to the phenotype-associated SNPs
Given that coeliac disease associated SNPs might not be causal variants but situated in their close proximity e.g in high linkage disequilibrium (LD) with them, FUMA (Functional Mapping and Annotation of GWAS) platform [17] was used for the functional annotation of the 39 genotyped SNPs. Publicly available GWAS summary statistic results in which our cohort has been included [5] and a pre-defined list of associated SNPs with phenotypes in this study were provided. All proxies in LD (r2 ≥ 0.8) with the phenotype-associated SNPs were identified using HaploReg v.4.1 [18]. The RegulomeDB 2.0 [19] was used to assign all the variants a score ranging from 1a to 7. Scores from 1a to 3a are likely to affect the expression of a gene. The lesser the score the higher likehood, thus 1a indicating a maximum and 3a a minimum likelihood to affect gene expression [19]. Variants scoring ≥3a but predicted as likely deleterious [Combined Annotation Dependent Depletion (CADD) score closest or >12.37)] were also included [20]. For those phenotype-associated SNPs scoring >3a, we selected their proxy with the lowest RegulomeDB score [19]. Using publicly available databases to study tissue-specific gene expression, we examined significant eQTLs associations (FDR < 0.05) with the functional variants, located nearby (cis-eQTL) or distal (trans-eQTL) to genes. They were assessed in the whole and peripheral blood using the Blood eQTL [21], BIOS QTL [22] and eQTLGen [23] browsers. Moreover, as the phenotype of coeliac disease varies considerable and the symptoms may affect the function of several cell types tissues we also used the Genotype-Tissue Expression data (brain, nerve, colon, oesophagus, EBV-transformed lymphocytes, muscle-skeletal, pancreas, skin, small intestine, stomach, cultured fibroblasts, thyroid, whole blood) [24]. Their participation in pathways and biological processes were identified by using KEGG (Kyoto Encyclopedia of Genes and Genomes) and GO (Gene Ontology) databases [25].
Genetic risk score
In order to assess the cumulative effects of the genotyped 39 non-HLA SNPs on phenotypes, we created a weighted GRS model (wGRS39), calculated by multiplying each allele by the natural logarithms of the previously reported OR values (β-coefficients) [5], followed by dividing their sum by the total number of alleles [9]. We analysed the wGRS39 model in tertiles, adjusted for sex, calculated according to the distribution of the average weighted risk alleles in the controls as published [26]. Based on these tertiles, all participants were categorised into the low-, medium- and high-risk groups.
Polygenic risk score
We derived a PRS using summary statistics from the most recent and largest GWAS in coeliac disease with 24,269 participants [6] as the discovery dataset and tested its association with phenotypes. After extracting only cases in our cohort, we performed LiftOver (https://genome.ucsc.edu/cgi-bin/hgLiftOver) to update our genotype data to a recent human genome build (hg19) and were left with 624 cases and 526,118 SNPs. We applied quality filters as described above and constructed the PRS accordingly [27] using the ‘fastscore’ option in PRSice software package [28]. PRSs were calculated as the sum of all risk variants carried by that individual weighted by the β-coefficients reported in the discovery GWAS [6]. To account for independent association signals, clumping retained the SNP with smallest P value in each 250 kilobases window and removed all those in LD (r2 > 0.2) with this SNP [7]. SNPs in the MHC region with long range LD (29.7–33.3 megabases) were excluded [10]. PRS was calculated on a priori set of significant thresholds (P < 0.001, P < 0.01, P < 0.05, P < 0.1, P < 0.2, P < 0.3, P < 0.4, P < 0.5) to identify the best fit PRS that was most predictive of an association.
Statistical analysis
Allelic associations using the genotyped 39 SNPs were tested by using the chi-squared method with one degree of freedom, measured by OR values and 95% confidence intervals. Logistic regression adjusted for SNP-gender interaction. We performed 104 permutation tests, generating an empirical PEMP2 for the evaluation of subgroups with a small sample size and adjusting for multiple tests [16]. Associations were statistically significant at PEMP2 ≤ 0.05. The Breslow–Day (BD) test examined the homogeneity of the association’s OR [PBD values < 0.02 (0.05/2)] in each male and gender strata [29].
Phenotype-association testing by using post-imputation genotype probabilities was performed under an additive genetic effect model using the frequentist likelihood score method implemented in SNPTEST v2.5.2 [30]. All the associations were adjusted for gender with the SNPTEST method Newml. For ChrX region we additionally used the ChrX-specific SNPTEST method with stratify-on gender option. Associations that reached out our permutation threshold (P value ≤ 0.001) were considered as statistically significant.
Using the Genetic Power Calculator [31], we computed the statistical power reached by the sample size set for each associated phenotype to detect allele association at permutation (P ≤ 0.001) and at nominal (P ≤ 0.01 and P ≤ 0.05) thresholds (detailed results in Supplementary Table S1). SNPs were assumed to be independent causal variants with no other causal variants in linkage disequilibrium (D′ = 1).
The eQTL associations by the selected variants located nearby (cis-eQTL) or distal (trans-eQTL) to genes were significant at a false discovery rate (FDR) < 0.05. An overall representation analysis adjusted by Benjamini & Hochberg procedure was performed by using WEB-based GEne SeT AnaLysis online toolkit [25] to identify significant (FDR < 0.05) eQTL gene-sets enriched in KEGG pathways and GO biological processes.
Logistic regression was used to test the associations of the wGRS39 tertiles with the coeliac phenotypes after comparing each group (medium [2nd tertile] and high [3rd tertile] wGRS39 risk categories) to a reference (low wGRS39 risk category, 1st tertile). The associations were quantified by ORs with 95% CIs, significant at P value < 0.05. In our case only model, we tested the associations between PRS and phenotypes using linear logistic regression, adjusting for age and sex. Multiple comparisons were addressed by applying 104 permutations to identify the best fit PRS (PEMP2 ≤ 0.05). Results are presented as change in variance (R2), with both unadjusted P values and PEMP2 values.
All data analysis, conversion of required input formats in the imputation analysis and graphs were performed using PLINK 1.90 (https://www.cog-genomics.org/plink/1.9), PLINK 2.0 (https://www.cog-genomics.org/plink/2.0/), PRSice v2 and RStudio (version 1.1.463-© 2009–2019 RStudio Inc., Boston, USA).
Results
Genetic associations with coeliac disease phenotypes
Of the previously identified 94 coeliac disease-associated variants, 39 were typed to sufficient extent and thus were selected for the study. When comparing coeliac disease patients to controls, ten SNPs were associated with specific phenotypes (Table 2, and Supplementary Table S2). Rs5979785 in the TLR7–TLR8 region was associated with coeliac disease diagnosis ≤7 years of age and stratifying for gender revealed that this association only occurs in girls (Breslow–Day test, P < 0.001). This SNP was associated with the age of coeliac disease diagnosis in girls (OR = 0.11; 95% CI = 0.03–0.46) also in the case only analysis comparing coeliac disease patients with the given phenotype to those without (detailed results in Supplementary Table S3). As regards coeliac disease diagnosis above 7 years of age in the case–control analysis, rs653178 (at SH2B3/ATXN2), rs13010713 (at ITGA4/UBE2E3), rs13151961 (at IL2/IL21), rs11712165 (at CD80) and rs10936599 (at MYNN) showed an association. Except for rs10936599, the same variants were also associated with an intermediate HLA risk. Moreover, rs653178, rs13010713 and rs13151961 showed an association with the presence of gastrointestinal symptoms and with the presence of more severe mucosal damage (TVA/SVA) at diagnosis. In addition, rs653178 was also associated with extraintestinal manifestation as well as coeliac disease associated conditions in general and more specifically with AITD and T1D.
In order to address the association of all the 94 coeliac disease associated SNPs, we carried out imputation. Out of the 2442 subjects in our study, 299 individuals (285 cases and 14 controls) were excluded due to heterozygosity and missingness. Of the 94 coeliac disease SNPs selected, five variants were missing (rs859637, rs2327832, rs12928822, rs58911644, rs4819388) and one (rs12998748) was multiallelic in the Finnish reference panel, and thus dropped out of the analysis. All the remaining 88 coeliac disease variants were successfully imputed and tested with the phenotypes (detailed results in Supplementary Table S4).
According to the results exploiting the imputed SNPs, five of the genotyped SNPs (rs5979785, rs653178, rs13010713, rs13151961, rs17810546) remained associated with the same phenotypes as in the original analysis. Seven imputed SNPs (rs3184504 at SH2B3/ATXN2, rs76830965 at IL12A, rs7616215 at CCR1, rs17264332 at OLIG3/TNFAIP3, rs243323 at SOCS1, rs13132308 at IL2/IL21 and rs990171 at IL18R1/IL18RAP), revealed as associated (P ≤ 0.001, adjusted for sex) with distinct phenotypes (Table 3, and Supplementary Table S4). Of these, rs76830965 was associated with coeliac disease diagnosis ≤7 years of age, the presence of gastrointestinal symptoms and malabsorption, intermediate HLA risk and coeliac disease autoantibodies. Moreover, rs3184504 was associated with coeliac disease diagnosis >7 years of age, extraintestinal manifestations, coeliac disease associated condition in general and with T1D specifically and also with intermediate HLA risk. The rest five imputed SNPs were associated with only one phenotype. Since 47.8% of cases were dropped out from the association analysis with the imputed SNPs, we carried out the remaining analysis with the 39 genotyped SNPs.
Functional annotation and pathway enrichment analysis
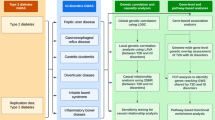
Of the 10 phenotype-associated genotyped SNPs, four variants (rs11712165, rs10936599, rs2327832 and rs13098911) had strong regulatory function on transcription with a RegulomeDB score ≤ 3a, indicating that the variant likely lies within a potential functional region. Of these rs13098911 was enriched with proxies with the lowest scores (1a and 1f). Out of the 148 extracted proxies to the other phenotype-associated SNPs (rs653178, rs13010713, rs17810546 and rs2298428), 42 had a putative functional role in gene regulation (RegulomeDB score ≤ 3a) (Supplementary Table S5). Thereafter we searched for eQTL effects for both the queried SNPs or the proxies that were likely to regulate gene expression or to be deleterious. We identified several significant highly tissue-specific eQTL effects (FDR < 0.05) (Supplementary Table S6). The eQTL genes connected to each of the phenotypes by the associated SNPs or their proxies were then subjected to KEGG pathways and GO terms analysis.
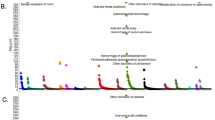
The analysis of the cis-eQTL genes of individual phenotypes, revealed significant enrichment only in the case of coeliac disease symptoms in childhood, neurological disorders, PVA and negative coeliac disease antibodies (Supplementary Table S7). The four cis-eQTL genes (CCR5, CCR3, CCR2, CXCR6) were enriched (FDR ≤ 0.002) in chemokine signalling and cytokine-cytokine receptor interaction KEGG pathways. These genes are likely to be involved in modulation of chemokine and cellular defence responses, cell chemotaxis second messenger-mediated signalling and divalent inorganic cation homoeostasis based on the GO associated terms (FDR ≤ 0.0001). Coeliac disease diagnosis above 7 years of age, gastrointestinal symptoms, coeliac disease-associated conditions, EI manifestations and intermediate HLA risk were the phenotypes connected to a set of trans-eQTL genes. Among KEGG pathways, they were most significantly enriched (FDR ≤ 0.01) in the B cell receptor signalling, Pertussis, NOD-like receptor signalling and in MicroRNAs in cancer. They were enriched (FDR ≤ 2.2E-09) for biological processes GO terms mostly involved in adaptive immune response, T cell activation, response to interferon-gamma, leucocyte cell-cell adhesion, cytokine secretion, regulation of leucocyte activation and response to molecule of bacterial origin (Supplementary Table S7).
Genetic risk score and polygenic risk score associations with coeliac phenotypes
In order to study the combined effect of the 39 SNPS on different phenotypes, we applied the wGRS39 tertiles (Supplementary Table S8). Patients at the highest wGRS39 tertiles had significantly higher risk for having coeliac disease-related symptoms during childhood (OR = 1.76, 95% CI = 1.12–2.77), a more severe small bowel mucosal damage (OR = 1.76, 95% CI = 1.08–2.90), malabsorption (OR = 1.62, 95% CI = 1.04–2.54) and anaemia (OR = 1.68, 95% CI = 1.03–2.78) (Table 4, and Supplementary Table S8).
As regards the PRS analysis we found a significant effect of PRS on DH at P values threshold (PT of 0.2, R2 = 0.06, P = 0.007, PEMP2 = 0.02) (Fig. 1a). Best fit PRS at PT of 0.001 predicted the presence of fractures but did not persist permutation correction (R2 = 0.025, P = 0.043, PEMP2 = 0.12) (Fig. 1b, and Supplementary Table S9).
Discussion
In this study, by using genotyped variants in our cohort, we identified a genotype-phenotype association for ten SNPs previously associated with coeliac disease susceptibility. In addition, our results demonstrate that combining 39 coeliac disease-associated genotyped SNPs into wGRS39 was more informative than a PRS to assess the genetic risk for distinct phenotypes of coeliac disease.
According to our results, rs5979785 located in the proximity of the TLR7 and TLR8 genes was associated with a decreased risk of diagnosis before 7 years of age in girls and notably this association was detected also in the case only comparison. Our RegulomeDB analysis did not retrieve rs5979785 as having regulatory function of transcription although it has previously been reported to decrease the TLR8 expression in the blood [5]. TLR7 and TLR8 are both members of the toll-like receptor family and they detect distinct forms of viral nucleic acids and initiate antiviral responses [32]. As distinct viruses, including enterovirus has been implicated as a risk factor for coeliac disease [33, 34], additional studies on this loci in relation with viral infections and early coeliac disease onset are called for.
Our results also revealed an association of rs653178 at SH2B3/ATXN2 with an increased risk to several other phenotypes, including concomitant T1D. Our findings are thus in accordance with previous studies where the 12q24/SH2B3 locus has been associated with both coeliac disease and T1D [8, 35, 36]. In these studies, rs3184504, a functional proxy in LD with rs653178 [37], has been indicated as the true SNP behind the association. Although rs3184504 was not included in our initial set of genotyped SNPs, we found it to be associated with concomitant T1D in the analysis carried out with the imputed genotypes.
In addition to having a likely regulatory function on transcription and an eQTL effect of the expression of SH2B3 along with many other genes in our analysis, rs3184504 SNP has been also associated with a higher expression of SH2B3 in the intestinal mucosal of patients with active coeliac disease [38], islet autoimmunity [39] and also implicated with bacterial infections [40], making the 12q24/SH2B3 locus associations in our cohort interesting for further research.
In pursue of clarifying the molecular mechanisms exerted by the susceptibility SNPs on the phenotypes, we identified that only three of our phenotype-associated SNPs had significant eQTL effects on regulating gene expression. This would suggest that the SNPs included in our analysis are mostly proxies and either situated in the proximity of the causative variant or further away. In the KEGG pathway and GO biological process analysis the phenotypes-associated rs13098911 SNP was identified as the only variant having cis-eQTL effects on a set of genes enriched in chemokine pathway. This finding most likely reflects the location of this variant in the chemokine gene cluster.
The enrichment results of the trans-eQTL genes identified largely identical pathways for most of our phenotypes. All these phenotypes were associated with rs653178 and its proxy rs3184504 revealed considerable effects on the expression of several distal genes. This pleiotropic effect of rs3184504 thus likely explains our finding of the pathway enrichment analysis of the trans-eQTL genes. Thus, further studies to infer the true phenotype-causal variants and the mechanisms that might mediate their effects on phenotypic variation are needed.
Regarding our weighted 39 SNP-based risk score model, the high-risk tertile in wGSR39 was associated with a higher risk of having coeliac disease symptoms in childhood, more severe small bowel mucosal damage, malabsorption and the occurrence of anemia. These phenotypes can be considered as signs of a more severe disease course [2, 41], thus raising the possibility that increased number of coeliac disease susceptibility SNPs might predispose to a more severe disease. In contrast, combining information from thousands of genomic variants into a PRS contributed to explain very little of the variance in phenotypes apart from DH. One major reason for this might be related to the fact that genetic loci not directly associated with disease status in large case–control GWAS may moderate the relationship between the coeliac polygenic burden and phenotypes [7, 10]. Thus, the main genetic contribution to phenotypic variations seems to derive from loci associated with the disease susceptibility.
A major strength of the current study is the clinically well-characterised large cohort of coeliac patients. In addition, a strength is the careful phenotyping of the patients, allowing us to investigate the association of different genotypes with various phenotypes. Moreover, the imputation analysis allowed us to explore the phenotype association of further 49 previously coeliac disease SNPs not typed in our cohort. The use of the most comprehensive available imputation panel in Finnish population increased our opportunities for identifying new coeliac disease SNPs associated with the phenotypes. Our highly conservative procedures in the imputation analysis diminished the risk of false positive associations. It must be noted though that 47.8% of coeliac disease patients dropped out from the analysis. Moreover, for distinct phenotypes, we had a fairly low number of patients which affected the statistical power in some cases. Moreover, we included in our GRS study only the 39 genotyped SNPs from independent coeliac disease risk loci, and not all the 94 previously associated with coeliac disease, and it might be the case that other coeliac susceptibility SNPs have a modulatory role on the phenotypes
We conclude that independent coeliac disease-susceptibility loci are associated with distinct coeliac disease phenotypes, suggesting that distinct SNPs might play a role in modulating the disease presentation in a yet to determined mechanism. Moreover, while PRS seems not to explain the variance in phenotypes, more severe coeliac disease phenotypes could possibly be contributed by higher amount of coeliac disease risk SNPs. Our GRS approach might thus be useful to identify patients at risk of developing a severe disease course unless identified and treated early. Further studies with a larger number of SNPs are called for in independent well-characterised patient cohorts to better understand how genetic variants contribute to the different coeliac disease phenotypes.
References
Singh P, Arora A, Strand TA, Leffler DA, Catassi C, Green PH, et al. Global prevalence of coeliac disease: systematic review and meta-analysis. Clin Gastroenterol Hepatol. 2018;16:826–36.
Lindfors K, Ciacci C, Kurppa K, Lundin KEA, Makharia GK, Mearin ML, et al. Coeliac disease. Nat Rev Dis Prim. 2019;5:3.
Leffler DA, Green PH, Fasano A. Extraintestinal manifestations of coeliac disease. Nat Rev Gastroenterol Hepatol. 2015;12:561–71.
Singh P, Arora S, Lal S, Strand TA, Makharia GK. Risk of coeliac disease in the first- and second-degree relatives of patients with coeliac disease: a systematic review and meta-analysis. Am J Gastroenterol. 2015;110:1539–48.
Dubois PC, Trynka G, Franke L, Hunt KA, Romanos J, Curtotti A, et al. Multiple common variants for coeliac disease influencing immune gene expression. Nat Genet. 2010;42:295–302.
Trynka G, Hunt KA, Bockett NA, Romanos J, Mistry V, Szperl A, et al. Dense genotyping identifies and localizes multiple common and rare variant association signals in coeliac disease. Nat Genet. 2011;43:1193–201.
Coleman C, Quinn EM, Ryan AW, Conroy J, Trimble V, Mahmud N, et al. Common polygenic variation in coeliac disease and confirmation of ZNF335 and NIFA as disease susceptibility loci. Eur J Hum Genet. 2016;24:291–7.
Smyth DJ, Plagnol V, Walker NM, Cooper JD, Downes K, Yang JH, et al. Shared and distinct genetic variants in type 1 diabetes and coeliac disease. N Engl J Med. 2008;359:2767–77.
Romanos J, Rosén A, Kumar V, Trynka G, Franke L, Szperl A, et al. Improving coeliac disease risk prediction by testing non-HLA variants additional to HLA variants. Gut. 2014;63:415–22.
Abraham G, Tye-Din JA, Bhalala OG, Kowalczyk A, Zobel J, Inouye M. Accurate and robust genomic prediction of coeliac disease using statistical learning. PLoS Genet. 2014;10:e1004374.
Borodulin K, Tolonen H, Jousilahti P, Jula A, Juolevi A, Koskinen S, et al. Cohort profile: the national FINRISK study. Int J Epidemiol. 2018;47:696–96i.
Monsuur AJ, de Bakker PI, Zhernakova A, Pinto D, Verduijn W, Romanos J, et al. Effective detection of human leukocyte antigen risk alleles in coeliac disease using tag single nucleotide polymorphisms. PLoS One. 2008;28:3–e2270.
Koskinen L, Romanos J, Kaukinen K, Mustalahti K, Korponay-Szabo I, Barisani D, et al. Cost-effective HLA typing with tagging SNPs predicts coeliac disease risk haplotypes in the Finnish, Hungarian, and Italian populations. Immunogenetics. 2009;61:247–56.
Zheng X, Shen J, Cox C, Wakefield JC, Ehm MG, Nelson MR, et al. HIBAG—HLA genotype imputation with attribute bagging. Pharmacogenomics J. 2014;14:192–200.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Clarke GM, Anderson CA, Pettersson FH, Cardon LR, Morris AP, Zondervan KT. Basic statistical analysis in genetic case-control studies. Nat Protoc. 2011;6:121–33.
Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Ward LD, Kellis M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 2012;40:D930–34.
Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012;22:1790–7.
Kircher M, Witten DM, Jain P, O’Roak BJ, Cooper GM, Shendure J. A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet. 2014;46:15.
Westra HJ, Peters MJ, Esko T, Yaghootkar H, Schurmann C, Kettunen J, et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat Genet. 2013;45:1238–43.
Zhernakova DV, Deelen P, Vermaat M, van Iterson M, van Galen M, Arindrarto W, et al. Identification of context-dependent expression quantitative trait loci in whole blood. Nat Genet. 2017;49:139–45.
Võsa U, Claringbould A, Westra H, Bonder MJ, Deelen P, Zeng B, et al. Unraveling the polygenic architecture of complex traits using blood eQTL meta analysis. bioRxiv 447367; https://doi.org/10.1101/447367.
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science. 2020;369:1318–30.
Liao Y, Wang J, Jaehnig EJ, Shi Z, Zhang B. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 2019;47:W199–205.
Weigl K, Thomsen H, Balavarca Y, Hellwege JN, Shrubsole MJ, Brenner H. Genetic risk score is associated with prevalence of advanced neoplasms in a colorectal cancer screening population. Gastroenterology. 2018;155:88–98.e10.
International Schizophrenia Consortium, Purcell SM, Wray NR, Stone JL, Visscher PM, O’Donovan MC, et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature. 2009;460:748–52.
Euesden J, Lewis CM, O’Reilly PF. PRSice: Polygenic Risk Score software. Bioinformatics. 2015;31:1466–8.
Liu K, Lai TY, Chiang SW, Chan VC, Young AL, Tam PO, et al. Gender specific association of a complement component 3 polymorphism with polypoidal choroidal vasculopathy. Sci Rep. 2014;4:7018.
Marchini J, Howie B, Myers S, McVean G, Donnelly P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet. 2007;39:906–13.
Purcell S, Cherny SS, Sham PC. Genetic power calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics. 2003;19:149–50.
Lester SN, Li K. Toll-like receptors in antiviral innate immunity. J Mol Biol. 2014;426:1246–64.
Cooper JD, Walker NM, Smyth DJ, Downes K, Healy BC, Todd JA. Follow-up of 1715 SNPs from the Wellcome Trust Case Control Consortium genome-wide association study in type I diabetes families. Genes Immun. 2009;10(Suppl 1):S85–S94.
Kahrs CR, Chuda K, Tapia G, Stene LC, Mårild K, Rasmussen T, et al. Enterovirus as trigger of coeliac disease: nested case-control study within prospective birth cohort. BMJ. 2019;364:l231.
Hagopian W, Lee HS, Liu E, Rewers M, She JX, Ziegler AG, et al. Co-occurrence of type 1 diabetes and coeliac disease autoimmunity. Pediatrics. 2017;140:e20171305.
Todd JA, Walker NM, Cooper JD, Smyth DJ, Downes K, Plagnol V, et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat Genet. 2007;39:857–64.
Li Y, He X, Schembri-King J, Jakes S, Hayashi J. Cloning and characterization of human Lnk, an adaptor protein with pleckstrin homology and Src homology 2 domains that can inhibit T cell activation. J Immunol. 2000;164:5199–206.
Plaza-Izurieta L, Castellanos-Rubio A, Irastorza I, Fernández-Jimenez N, Gutierrez G, CEGEC, et al. Revisiting genome wide association studies (GWAS) in coeliac disease: replication study in Spanish population and expression analysis of candidate genes. J Med Genet. 2011;48:493–6.
Sharma A, Liu X, Hadley D, Hagopian W, Chen WM, Onengut-Gumuscu S, et al. Identification of non-HLA genes associated with development of islet autoimmunity and type 1 diabetes in the prospective TEDDY cohort. J Autoimmun. 2018;89:90–100.
Zhernakova A, Elbers CC, Ferwerda B, Romanos J, Trynka G, Dubois PC, et al. Evolutionary and functional analysis of coeliac risk loci reveals SH2B3 as a protective factor against bacterial infection. Am J Hum Genet. 2010;86:970–7.
Zhernakova A, Elbers CC, Ferwerda B, Romanos J, Trynka G, Dubois PC, et al. Coeliac disease patients presenting with anemia have more severe disease than those presenting with diarrhea. Clin Gastroenterol Hepatol. 2013;11:1472–7.
Acknowledgements
This study was supported by grants from the Academy of Finland, the Sigrid Juselius Foundation, the Foundation for Pediatric Research and the Competitive State Research Financing of the Expert Area of Tampere University Hospital. The funding sources had no role in the design of the study, the collection, analysis, or interpretation of the data; the preparation or review of the manuscript; or the decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
JXMC: Study concept and design, acquisition of the in silico data, pre-processing chip genotyped data to imputation with SISu v3 panel, analysis and interpretation of the genotyped and imputed data, drafting of the manuscript, and statistical analysis. PS: Study concept and design, imputation of the genotyped data, and study supervision. KK: Acquisition of the data, analysis and interpretation of the data, and acquisition of funding. PL: Analysis and interpretation of the data. HH: Statistical analysis. MN: Analysis and interpretation of the functional annotation data. LLEK: Statistical analysis. DAY: Statistical analysis. EK, AS and AP: Chip data pre-imputation QC and genotype imputation with SISu v3 imputation reference panel. AP: Acquisition of the SISu v3 imputation reference panel data. KKa: Study concept and design, acquisition of the data, study supervision, and acquisition of funding. KL: Study concept and design, analysis and interpretation of the data, study supervision and acquisition of funding. All authors approved the final manuscript and contributed critical revisions to its intellectual content.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cerqueira, J.X.M., Saavalainen, P., Kurppa, K. et al. Independent and cumulative coeliac disease-susceptibility loci are associated with distinct disease phenotypes. J Hum Genet 66, 613–623 (2021). https://doi.org/10.1038/s10038-020-00888-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-020-00888-5
This article is cited by
-
Family-based association of 4q27chromosomal region covering IL2-IL21 genes with type 1 diabetes (T1D)—a study of genetic risk factors
International Journal of Diabetes in Developing Countries (2023)