Abstract
The Y-chromosome haplogroup C2b1a3a2-F8951 is the paternal lineage of the Aisin Gioro clan, the most important brother branch of the famous Mongolic-speaking population characteristic haplogroup C2*-Star Cluster (C2b1a3a1-F3796). However, investigations on its internal phylogeny are still limited. In this study, we used whole Y-chromosome sequencing to update its phylogenetic tree. In the revised tree, C2b1a3a2-F8951 and C2*-Star Cluster differentiated 3852 years ago (95% CI = 3295–4497). Approximately 3558 years ago (95% CI = 3013–4144), C2b1a3a2-F8951 was divided into two main subclades, C2b1a3a2a-F14753 and C2b1a3a2b-F5483. Currently, samples of C2b1a3a2-F8951 were mainly from the House of Aisin Gioro clan, the Ao family from Daur and some individuals mainly from northeast China. Although other haplogroups are also found in the Ao family, including C2b1a2-M48, C2b1a3a1-F3796, C2a1b-F845, and N1c-M178, the haplogroup C2b1a3a2-F8951 is still the most distinct genetic component. For haplogroup C2b1a3a2-F8951, the time of the most recent common ancestor of the House of Aisin Gioro clan and the Ao family were both very late, just a few hundred years ago. Some family-specific Y-SNPs of the House of Aisin Gioro and the Ao family were also discovered. This revision evidently improved the resolving power of Y-chromosome phylogeny in northeast Asia, deepening our understanding of the origin of these two families, even the Mongolic-speaking population.
Similar content being viewed by others
Introduction
In recent years, the male-specific region of the Y-chromosome (MSY) has shown to be a powerful tool in paternity testing, genetic genealogy, biogeographical ancestry, and analysis of population demographic structure [1,2,3]. Two genetic markers in MSY, Y-STRs (short tandem repeats), and Y-SNPs (single-nucleotide polymorphisms), are often used to identify the genetic relationships between samples, but both of them have substantial limitations. Distinct patrilineal lineages can easily be identified by high-resolution Y-STR haplotyping [4, 5], but there are two widely used but very different mutation rates for Y-STR, which are often controversial in paternal lineage dating [6, 7]. The Y-SNP markers have a lower mutation rate than Y-STRs, so they are stable in paternity identification. However, except for the inevitably incomplete resolution of the SNP-defined phylogenic tree, exponentially available Y-SNP data bring disorder to geneticists [8].
Compared with the common Y-STR and Y-SNP markers, whole Y-chromosome sequencing has unparalleled advantages, even though it is more expensive and contains more precise workflows. First, the new Y-SNP sites, even family-specific Y-SNPs, can be discovered quickly and efficiently [9, 10]. Second, after ancient DNA data correction, the molecular clock based on whole-sequence is gradually becoming a more reasonable choice in Y-chromosome dating than Y-STR methods [11, 12].
The people in northeastern China have profoundly influenced the demographic history of the eastern part of Eurasia. While some general population genetics studies have been carried out [13,14,15,16], little research has been conducted to explain the history of important genealogies, in addition to that of the Aisin Gioro clan [17]. The House of Aisin Gioro, as the core of Manchu, conquered other Jurchen tribes and established the last empire in Chinese history—the Qing Dynasty. The paternal lineage of this clan belongs to haplogroup C2b1a3a2-F8951, the most important brother branch of the famous Mongolic-speaking population characteristic haplogroup C2*-Star Cluster [17]. On the basis of the scattered Y-STR data in published literature, previous researchers speculated that the Aisin Gioro clan may share a patrilineal ancestor with some of the current Daur population [17].
The origin of the Daur population, in other words, the genetic relationship between the Daur, the ancient Khitan, and the Mongolian peoples, has been a hot topic in population genetics and ethnohistory for a long time [17,18,19]. Initially, the Daur population settled on the north beach of the Heilongjiang River (Amur River) [20]. After the invasion of the Russian Cossacks in the 17th century, the Daur people gradually migrated southward [21]. In the Qing Dynasty, the Daur people acquired a high social status and were often dispatched to suppress rebel forces [22]. Now, they mainly live in Hulunbuir (Inner Mongolia Autonomous Region), Qiqihar (a city near Hulunbuir, in Heilongjiang Province), Tacheng (Xinjiang Autonomous Region), and the far east of Russia. In the Qing Dynasty, the paternal social units of the Daur population were composed of Hala and its subordinate Mokun [23]. Currently, Hala has the same meaning as the surname. There are ~20 surnames in the Daur population, of which, the ancient clan of Ao has the largest population [23]. Based on a large amount of Y-STR investigation in East Asia (unpublished data in our laboratory), we found that a similar haplotype was shared by the House of Aisin Gioro and some individuals from the Ao clan.
In this study, we analyzed the paternal lineage of the Ao family, including Y-STR and Y-SNP data. Moreover, we revised phylogenetic tree of haplogroup C2b1a3a2-F8951 according to sequences from the House of Aisin Gioro, the clan of Ao and other scattered but genetically related samples mainly from northeast China.
Materials and methods
Samples and ethical requirements
Saliva or blood samples of 633 individuals from more than 40 populations were collected (see Table S1) from healthy males from populations in eastern Eurasia between 2003 and 2018. Except for two sequences from the Aisin Gioro clan [17], we selected 13 samples to conduct whole Y-chromosome sequencing, containing six samples from the Ao clan and seven samples from northeast China, including Daur, Mongolian, and Ewenk. The ethics committee for biological research at the School of Life Sciences at Fudan University approved the study.
Workflows for next-generation sequencing and statistical analysis
Genomic DNA was extracted using the DP-318 Kit (Tiangen Biotechnology, Beijing, China) according to the manufacturer’s protocol. The DNA was sent for sequencing on the Illumina HiSeq2000 platform (Illumina, San Diego, CA, USA). A series of bait libraries were designed to capture the sequences of an ~7.33M region on the Y-chromosome. A mature procedure, including DNA shearing, adding an adaptor, and gel electrophoresis, which we described previously, were performed prior to next-generation sequencing [24]. Typical pipelines were employed to perform whole-sequencing analyses and determine genotypes and haplogroups for each sample [25, 26]. The raw sequence data reported in this study have been deposited in the Genome Sequence Archive (GSA) [27] in the BIG Data Center (Members BIGDC 2017), Beijing Institute of Genomics (BIG), Chinese Academy of Sciences, under an accession number HRA000024, which is publicly accessible at http://bigd.big.ac.cn/gsa-human.
Standard procedures (bwa+samtools) were used to analyze the sequencing results [25, 28]. Some details of the statistical analysis were consistent with the relevant chapters of our previous papers [10, 24]. The revised phylogenetic tree for F8951 was constructed under regulations of the Y Chromosome Consortium (YCC). Beast v.2.4.3 was employed to estimate the coalescent times for haplogroup F8951 [29]. To gain a better substitution model, a Bayesian skyline coalescent tree prior was chosen with the bModelTest package in Beast 2.0 software [29]. The calculation was conducted with 10 million iterations and sampling every 1000 steps. Our results were visualized in Tracer v.1.6 and FigTree v1.4.2 with a burn-in of 20%, and all effective sample sizes were above 200.
The corresponding Y-STR data for these sequencing samples and a series of available Y-STR haplotypes from previous studies, which are close to the profiles of the Ao and Aisin Gioro clans, were collected (Table S1).
Result
The paternal lineage of the Ao family
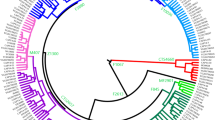
A total of five haplogroups were discovered in the Ao family, including haplogroups C2b1a3a1-F3796 (N = 5), C2b1a2-M48 (N = 12), C2a1b-F845 (N = 4), N1c-M178 (N = 1), and C2b1a3a2-F8951 (N = 6). For haplogroups C2b1a2-M48, C2b1a3a1-F3796, and C2b1a3a2-F8951, a large amount of data has been accumulated from published literature (Table S1), so additional Y-STR profiles were collected and used to construct the reduced-median network (Figs. 1 and 2). As shown in Fig. 1a, the five samples from the Ao family were scattered in the C2b1a3a1-F3796 network. Except for one sample that was located in a branch containing slightly more Tungusic samples, the other four samples were clustered with populations of Mongolian origin. Since the C2b1a3a1-F3796 haplogroup is high in almost all Mongolic populations [30, 31] and the Daur people are an ancient part of the Mongolic populations, this distribution is very reasonable. The sample close to Tungusic groups can be explained as extended genetic communication with Ewenki and Oroqen since the Qing Dynasty. In Fig. 1b, in addition to one sample that may be differentiated from the Kazakh population, most of the samples clustered to the root Tungusic population rather than the later differentiated Kalmyk population. C2b1a2-M48, the absolute main paternal lineage of the Tungusic population [32] and an important paternal lineage of the Mongolic population [33], has been widely distributed in the Mongolian plateau before the expansion of Mongols. It is very likely that the Ao family obtained this genetic component from the nearby Ewenki and Oroqen or other ancient populations. According to unpublished data from our laboratory, C2a1b-F845 is a new downstream haplogroup, with high frequency in the southern Chinese population, such as Tujia, Miao and Yao. This suggests that the genetic components of the Ao family are not only from northeast Asia but also include the unknown gene flow from distant southern China. Haplogroup N1c-M178 is widely distributed in northeast Asia with many unknown branches [34], so it is normal that this haplogroup appeared once in the Ao family.
Y-STR network of C2b1a3a1-F3796 and C2b1a2-M48. a Y-STR network of C2b1a3a1-F3796 based on 15 Y-STRs (DYS19, DYS389I/II, DYS390, DYS391, DYS392, DYS393, DYS437, DYS438, DYS439, DYS448, DYS456, DYS458, DYS635, and Y-GATA H4). b Y-STR network of C2b1a2-M48 based on 9 Y-STRs (DYS389I/II, DYS389I/II, DYS390 DYS391, DYS392, DYS393, DYS437, DYS438, and DYS439). To determine the root of C2b1a2-M48, we added the typical Y-STR value of C2-M217 as a reference, located to the right of the red line in the lower right corner of the figure. Note: For the raw Y-STR data used to construct the Network, please see the “used for network” section in Tale S1. Different colors represent different populations. The size of the pie chart is positively related to the size of the population. The distance between the circles symbolizes the genetic distance between corresponding populations
Y-STR network of C2b1a3a2-F8951 based on 15 Y-STRs (DYS19, DYS389I/II, DYS390, DYS391, DYS392, DYS393, DYS437, DYS438, DYS439, DYS448, DYS456, DYS458, DYS635, and Y-GATA H4). Note: For the raw Y-STR data used to construct the Network, please see the “used for network” section in Tale S1. Different colors represent different populations. The size of the pie chart is positively related to the size of the population. The distance between the circles symbolizes the genetic distance between corresponding populations
For C2b1a3a2-F8951, the samples from the Aisin Gioro family are characterized by DYS389I = 14 and DYS439 = 10, while the samples from the Ao family are characterized by DYS389I = 13 and DYS439 = 11 (Table S1). As shown in Fig. 2, samples from the Aisin Gioro family and Xinbin-Manchu group form a clade in the lower right part of the Y-STR network, whereas other samples from the Ao clan and the Daur, Mongolian, Xibe, Oroqen, and Buryat groups cluster together on the other left branch. Moreover, all the samples of the Ao family cluster with other Daur, Mongolian, Buryat, and Evenk groups, forming the largest circle in the figure. It shows that these samples of the Ao family came from a male ancestor in a relatively recent time, and his descendants have successfully spread in the Daur group and even other Mongolian groups around them. In other words, the haplogroup C2b1a3a2-F8951 is likely to be the core paternal lineage of the Ao family.
Above all, although various haplogroups have been found in the Ao family, C2b1a3a2-F8951 is still the most distinct genetic component of all. In addition, it is the paternal lineage of the Aisin Gioro clan, the most important brother branch of the haplogroup C2b1a3a1-F3796. It is necessary to carry out an investigation on its internal phylogeny.
Revised phylogeny and time estimates for haplogroup C2b1a3a2-F8951
Based on whole Y-chromosome sequencing data, we show a revised phylogenetic tree of haplogroup C2b1a3a2-F8951 in Fig. 3. The new tree contained several subclades and a series of new polymorphisms. Thirty Y-SNP markers were unique for the Aisin Gioro clan, while 91 Y-SNP markers were distinct for the Ao clan. The new phylogenetic tree for haplogroup C2b1a3a2-F8951 is divided into two distinct secondary branches, C2b1a3a2a-F14753 and C2b1a3a2b-F5483. Two samples from the Aisin Gioro clan belong to one subbranch, C2b1a3a2a-F14750, while six samples from the Ao clan belong to the other subbranch, C2b1a3a2b-F5483. A sample from the Hulunbuir-Ewenk group has a closer genetic relationship with the Aisin Gioro clan, but their ancestors separated 2350 years ago (95% CI = 1840–2885). The sample from the Xinbin-Manchu group is the closest known sample to the Aixinjue family (Fig. 2), but unfortunately, we are unable to obtain its sequencing data in this study. In addition, we must note that six scattered samples, which also carry the genetic marker F5483, were all from minorities (Daur, Mongolian, and Evenk) in Hulunbuir. Considering that Hulunbuir has been managed by the Daur population for hundreds of years and is now the largest settlement of the Daur population, we speculate that these samples may be a result of recent spread from the Ao family.
Revised phylogenetic tree and time estimates for haplogroup C2b1a3a2-F8951 (Note: The age of the Aisin Gioro clan cited the previous result [17]. The blue numbers indicate the number of mutations for each segment of the tree.)
As shown in Fig. 1, the divergence time between haplogroup C2b1a3a2-F8951 and its most closely related lineage (represented by sample ELT00023) is ~3852 years old. Although the haplogroup C2b1a3a2-F8951 appeared very early, the time of the most recent common ancestor (TMRCA) of the Aisin Gioro and the Ao clan are both relatively young. The Aisin Gioro family originated ~470 years ago, while the Ao clan has a genetic history of ~787 years. This may be due to the population bottleneck caused by the long-term battles between many nationalities and shortages of living resources in ancient northeast China.
Discussion
It is easy to speculate that sometimes families who share the same surname tend to share the same Y-chromosome. Previous scholars have researched this kind of case using Y-STR and Y-SNP data [5, 35,36,37,38]. However, in this study, the Aisin Gioro family is the leader of Manchus, speaking a language that belongs to a branch of Tungusic, while the Ao family has the largest population among all Daur populations, speaking a language that belongs to a branch of Mongolic (http://www.ethnologue.com). Although they have different surnames, languages, ethnic identities, and no common patrilineal ancestor in their memories, they did share a paternal ancestor of 3558 (95% CI = 3013–4144) years ago by Y-chromosome whole-sequence analysis. It is difficult to find specific historical events to answer why the two families developed independently because we know very little about the history of their differentiated time. Interestingly, TMRCA of the Aisin Gioro and Ao clans are both a few hundred years old. This may be the result of a bottleneck caused by the long-term battles and migrations in ancient northeast China, as mentioned above. In particular, we have sufficient evidence to prove that the Daur people experienced a terrible population loss in the invasion of the Russian Cossacks in the 17th century and subsequent migrations [21, 22]. To understand the early genetic traits of these two families, it is necessary to conduct more work on DNA testing covering more populations, including ancient DNA analysis.
After discovering the paternal kinship and related time between the two families, the next two interesting questions are the ancestors and birthplaces of haplogroup C2b1a3a2-F8951. First, haplogroup C2b1a3a2-F8951 is the most important brother branch of haplogroup C2b1a3a-F3796 (Fig. 3). There is no doubt that haplogroup C2b1a3a- F3796 plays a vital role in the formation of the Mongolian population, even though it may not be the paternal type of Genghis Khan himself [30, 31]. We speculate that the earliest ancient population, carrying haplogroup C2b1a3a2-F8951, also has a direct relationship with the origin of the Mongolians. This hypothesis is more convincing when we consider that the Daur language also belongs to a branch of Mongolic and conserved some ancient Mongolian vocabulary, which was lost in modern Mongolian [39]. Second, in northeast China, it was not until ~3000 years ago that Dong-Hu, the first ancient ethic group, appeared in history [40]. By coincidence, the time when Dong-Hu appeared was just a little later than that of C2b1a3a2-F8951 (~3558). Furthermore, it is widely accepted that Mongolians have developed from Dong-Hu. Therefore, we conclude that haplogroup C2b1a3a2-F8951, carried by the Aisin Gioro and Ao families, may be traced back to the ancestors of Dong-Hu.
Although, except for the Aisin Gioro family, the known samples of C2b1a3a2-F8951 are almost entirely from Hulunbuir, it is clearly a result of the migration of the Daur. According to previous research on the Aisin Gioro family [17] and the memories of the Daur people [22], we can only speculate that the northern bank of Heilongjiang was an important settlement to the population of C2b1a3a2-F8951 before the 17th century. More genetic investigations are needed to trace the birthplace of this haplogroup.
As mentioned above, the origin of the Daur is mainly related to the Khitan and Mongolians [17,18,19]. On the one hand, the representative haplogroups of Mongolic-speaking populations, including C2b1a2-M48, C2b1a3a1-F3796, C2b1a1a1-F1756, and C2a1a1a1-M407, have been studied thoroughly [9, 10, 30,31,32]. This paper confirms that some Daur populations represented by the Ao family are indeed an ancient branch of the Mongolic-speaking population. On the other hand, the ancient Khitan population disappeared for almost a thousand years. To clarify whether they have a patrilineal relationship, it is necessary to compare the samples from the modern Daur with those from the ancient Khitan. We need more families and more samples to understand the origin of the Daur more comprehensively.
In summary, our research provides a somewhat atypical application of whole-sequence analysis to genealogy study. The revised phylogeny tree of C2b1a3a2-F8951 evidently improved the resolving power of Y-chromosome phylogeny in northeast Asia, deepening our understanding of the origin of the Aisin Gioro and Ao families, even the Mongolic-speaking population. However, in East Asia, the genetic history of many important families has not been ascertained, such as the Wanyan family (the founder of Jin Dynasty) and the Liu family (the founder of Han Dynasty). These may be important research directions for future anthropology scholars.
References
Kayser M. Forensic use of Y-chromosome DNA: a general overview. Hum Genet. 2017;136:621–5.
Shi CM, Li C, Ma L, Chi L, Zhao J, Yuan W, et al. Inferring Chinese surnames with Y-STR profiles. Forensic Sci Int Genet. 2018;33:66–71.
Toscanini U, Gaviria A, Pardo-Seco J, Gomez-Carballa A, Moscoso F, Vela M, et al. The geographic mosaic of Ecuadorian Y-chromosome ancestry. Forensic Sci Int Genet. 2018;33:59–65.
Claerhout S, Vandenbosch M, Nivelle K, Gruyters L, Peeters A, Larmuseau M, et al. Determining Y-STR mutation rates in deep-routing genealogies: identification of haplogroup differences. Forensic Sci Int Genet. 2018;34:1–10.
Kayser M, Vermeulen M, Knoblauch H, Schuster H, Krawczak M, Roewer L. Relating two deep-rooted pedigrees from Central Germany by high-resolution Y-STR haplotyping. Forensic Sci Int Genet. 2007;1:125–8.
Wei W, Ayub Q, Xue Y, Tyler-Smith C. A comparison of Y-chromosomal lineage dating using either resequencing or Y-SNP plus Y-STR genotyping. Forensic Sci Int Genet. 2013;7:568–72.
Zhivotovsky LA, Underhill PA, Cinnioglu C, Kayser M, Morar B, Kivisild T, et al. The effective mutation rate at Y chromosome short tandem repeats, with application to human population-divergence time. Am J Hum Genet. 2004;74:50–61.
Larmuseau MH, Van Geystelen A, Kayser M, van Oven M, Decorte R. Towards a consensus Y-chromosomal phylogeny and Y-SNP set in forensics in the next-generation sequencing era. Forensic Sci Int Genet. 2015;15:39–42.
Wei LH, Huang YZ, Yan S, Wen SQ, Wang LX, Du PX, et al. Phylogeny of Y-chromosome haplogroup C3b-F1756, an important paternal lineage in Altaic-speaking populations. J Hum Genet. 2017;62:915–8.
Huang YZ, Wei LH, Yan S, Wen SQ, Wang CC, Yang YJ, et al. Whole sequence analysis indicates a recent southern origin of Mongolian Y-chromosome C2c1a1a1-M407. Mol Genet Genom. 2018;293:657–63.
Helgason A, Einarsson AW, Guethmundsdottir VB, Sigurethsson A, Gunnarsdottir ED, Jagadeesan A, et al. The Y-chromosome point mutation rate in humans. Nat Genet. 2015;47:453–7.
Karmin M, Saag L, Vicente M, Wilson SM, Jarve M, Talas UG, et al. A recent bottleneck of Y chromosome diversity coincides with a global change in culture. Genome Res. 2015;25:459–66.
Han Y, Li L, Liu X, Chen W, Yang S, Wei L, et al. Genetic analysis of 17 Y-STR loci in Han and Korean populations from Jilin Province, Northeast China. Forensic Sci Int Genet. 2016;22:8–10.
Chen WQ, Feng Z, Jin W, Zhang YJ, Wang Q, Li P, et al. Genetic analysis of 17 Y-STR loci from 1026 individuals of Han populations in Jilin Province, Northeast China. Int J Leg Med. 2018;132:1309–11.
He J, Guo F. Population genetics of 17 Y-STR loci in Chinese Manchu population from Liaoning Province, Northeast China. Forensic Sci Int Genet. 2013;7:e84–5.
Guo F, Zhang L, Jiang X. Population genetics of 17 Y-STR loci in Xibe ethnic minority from Liaoning Province, Northeast China. Forensic Sci Int Genet. 2015;16:86–7.
Wei LH, Yan S, Yu G, Huang YZ, Yao DL, Li SL, et al. Genetic trail for the early migrations of Aisin Gioro, the imperial house of the Qing dynasty. J Hum Genet. 2017;62:407–11.
Kong QP, Yao YG, Liu M, Shen SP, Chen C, Zhu CL, et al. Mitochondrial DNA sequence polymorphisms of five ethnic populations from northern China. Hum Genet. 2003;113:391–405.
Chen S. A study on the origin of Daur. Ethno-Natl Stud. 1959;8:43–50.
Narangoa L, Cribb R. Historical Atlas of Northeast Asia, 1590–2010: Korea, Manchuria, Mongolia, Eastern Siberia. New York: Columbia University Press;2014.
Dmytryshyn, B, Crownhart-Vaughan, EAP, Vaughan, T. Russia’s conquest of Siberia, 1558–1700: a documentary record, vol. 1. Portland: The Press of the Oregon Historical Society;1985.
Aola AL. The western expedition and defense of Solon. Beijing, China: Minzu University of China Press;2017.
Duo WZ. Hala and Mokun of Daur in Qiqihar. Heilongjiang History. 2014;1:367–78.
Wang LX, Lu Y, Zhang C, Wei LH, Yan S, Huang YZ, et al. Reconstruction of Y-chromosome phylogeny reveals two neolithic expansions of Tibeto-Burman populations. Mol Genet Genom. 2018;293:1–8.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Li H, Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 2009;25:1754–60.
Wang Y, Song F, Zhu J, Zhang S, Yang Y, Chen T. GSA: genome sequence. Genom Proteom Bioinform. 2017;15:14–8.
Li H, Durbin R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics . 2010;26:589–95.
Bouckaert R, Heled J, Kuhnert D, Vaughan T, Wu CH, Xie D, et al. BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput Biol. 2014;10:e1003537.
Wei LH, Yan S, Lu Y, Wen SQ, Huang YZ, Wang LX, et al. Whole-sequence analysis indicates that the Y chromosome C2*-Star Cluster traces back to ordinary Mongols, rather than Genghis Khan. Eur J Hum Genet. 2018;26:230–7.
Zerjal T, Xue Y, Bertorelle G, Wells RS, Bao W, Zhu S, et al. The genetic legacy of the Mongols. Am J Hum Genet. 2003;72:717–21.
Zhong H, Shi H, Qi XB, Xiao CJ, Jin L, Ma RZ, et al. Global distribution of Y-chromosome haplogroup C reveals the prehistoric migration routes of African exodus and early settlement in East Asia. J Hum Genet. 2010;55:428–35.
Rootsi S, Zhivotovsky LA, Baldovic M, Kayser M, Kutuev IA, Khusainova R, et al. A counter-clockwise northern route of the Y-chromosome haplogroup N from Southeast Asia towards Europe. Eur J Hum Genet. 2007;15:204–11.
Wells RS, Yuldasheva N, Ruzibakiev R, Underhill PA, Evseeva I, Blue-Smith J, et al. The Eurasian heartland: a continental perspective on Y-chromosome diversity. Proc Natl Acad Sci USA. 2001;98:10244–9.
McEvoy B, Bradley DG. Y-chromosomes and the extent of patrilineal ancestry in Irish surnames. Hum Genet. 2006;119:212–9.
Martinez-Cadenas C, Blanco-Verea A, Hernando B, Busby GB, Brion M, Carracedo A, et al. The relationship between surname frequency and Y chromosome variation in Spain. Eur J Hum Genet. 2016;24:120–8.
Sole-Morata N, Bertranpetit J, Comas D, Calafell F. Y-chromosome diversity in Catalan surname samples: insights into surname origin and frequency. Eur J Hum Genet. 2015;23:1549–57.
King TE, Jobling MA. Founders, drift, and infidelity: the relationship between Y chromosome diversity and patrilineal surnames. Mol Biol Evol. 2009;26:1093–102.
Erdentai, Oyondalai, Asaraltu. Some Interpretation of the vocabulary of “Mongolian Secret History”. Hohhot: Inner Mongolia People Publishing House;1980.
Guo DC. Supplement, correction and study of “Wang Hui”. Changchun: Jilin Education Audiovisual Press;1991.
Acknowledgements
We especially thank all our sample donors and all of the people who helped with sample collection, including Fu-Rong Su, A-li, Bai-Ling Jin, Xin Jin, Yue-Ling, and Zhong-Mei Ao of the Hulunbuir Daur Ethnic Culture Research Association. This study was supported by the National Natural Science Foundation of China (NSFC Nos. 81671874 and 31671297) and the Ministry of Science and Technology of China (MOST) (2016YFC0900300).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, CZ., Wei, LH., Wang, LX. et al. Relating Clans Ao and Aisin Gioro from northeast China by whole Y-chromosome sequencing. J Hum Genet 64, 775–780 (2019). https://doi.org/10.1038/s10038-019-0622-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-019-0622-4
This article is cited by
-
Whole mitochondrial genome analysis of the Daur ethnic minority from Hulunbuir in the Inner Mongolia Autonomous Region of China
BMC Ecology and Evolution (2022)