Abstract
Severe congenital eye malformations, particularly microphthalmia and anophthalmia, are one of the main causes of visual handicap worldwide. They can arise from multifactorial, chromosomal, or monogenic factors and can be associated with extensive clinical variability. Genetic analysis of individuals with these defects has allowed the recognition of dozens of genes whose mutations lead to disruption of normal ocular embryonic development. Recent application of next generation sequencing (NGS) techniques for genetic screening of patients with congenital eye defects has greatly improved the recognition of monogenic cases. In this study, we applied clinical exome NGS to a group of 14 Mexican patients (including 7 familial and 7 sporadic cases) with microphthalmia and/or anophthalmia. Causal or likely causal pathogenic variants were demonstrated in ~60% (8 out of 14 patients) individuals. Seven out of 8 different identified mutations occurred in well-known microphthalmia/anophthalmia genes (OTX2, VSX2, MFRP, VSX1) or in genes associated with syndromes that include ocular defects (CHD7, COL4A1) (including two instances of CHD7 pathogenic variants). A single pathogenic variant was identified in PIEZO2, a gene that was not previously associated with isolated ocular defects. NGS efficiently identified the genetic etiology of microphthalmia/anophthalmia in ~60% of cases included in this cohort, the first from Mexican origin analyzed to date. The molecular defects identified through clinical exome sequencing in this study expands the phenotypic spectra of CHD7-associated disorders and implicate PIEZO2 as a candidate gene for major eye developmental defects.
Similar content being viewed by others
Introduction
Human eye development is a coordinated process that requires the ordered activation of numerous genes in a spatially and temporally controlled manner during embryonic life [1, 2]. Environmental and/or genetic factors disturbing this process can result in a range of congenital eye anomalies that lead to impaired visual function in affected subjects [3]. Developmental anomalies can affect many parts of the eye and are termed panocular defects, whereas other abnormalities may be restricted to specific structures of the anterior segment, posterior segment, or can alter the differentiation of particular cell populations, such as photoreceptors [4]. Collectively, congenital ocular defects are one of the most common causes of irreversible visual deficiency in humans, accounting for ~25% of severely visually impaired children [5].
Among genetic causes of congenital eye defects, monogenic etiology is of particular importance as it carries a high risk of recurrence in families and because affected subjects frequently suffer from associated extraocular anomalies [6, 7]. Monogenic ocular defects are also of great interest since they have allowed a better understanding of the molecular processes that occur during normal eye formation, by recognizing key genetic factors that cause aberrant eye development when mutated [8,9,10].
During the past years, it has been established that up to 25% of cases with severe congenital ocular defects such as microphthalmia (small eye) or anophthalmia (absent eye) are due to pathogenic variants in genes related to eye development, including SOX2 (MIM 184429), RAX (MIM 601881), OTX2 (MIM 600037), VSX2 (MIM 142993; formerly CHX10), FOXE3 (MIM 601094), and ALDH1A3 (MIM 600463), among others [11, 12]. When various molecular tools are applied for genetic screening, including Sanger sequencing, copy number variation analysis, and array CGH, a genetic cause can be demonstrated in up to 80% of cases of bilateral anophthalmia or microphthalmia, with SOX2 and OTX2 being the most commonly mutated genes [13]. However, in typical published cohorts, in which patients with a combination of phenotype severities and laterality are included, molecular diagnosis declines to ~20% [12, 14,15,16].
Typically, molecular diagnosis in subjects with eye anomalies has relied on Sanger sequencing, in a gene-by-gene basis. As the current number of genes associated to eye defects is high and is expected to continue to grow, Sanger sequencing has limited utility for an exhaustive molecular screening in those cases with suspected monogenic etiology. Recently, next generation sequencing (NGS) has emerged as an efficient method for systematic and simultaneous analysis of dozens of genes involved in congenital eye defects [17]. Thus, several cohorts of microphthalmic/anophthalmic patients have been analyzed by NGS, resulting in rates of molecularly solved cases between 11–36% [18,19,20,21]. It is anticipated that the extended application of NGS for molecular diagnosis will allow the establishment of more realistic mutational rates for genes involved in these disorders, both at the global and at the ethnic-specific levels. Concurrently, expansion of NGS-based diagnosis in clinical practice must be encouraged, as it will allow an improvement in current diagnostic rates and more rational follow-up of affected patients.
In order to contribute to a better molecular characterization of subjects with developmental eye defects, we applied NGS to a group of 14 Mexican patients with microphthalmia and/or anophthalmia. Causal mutations were demonstrated in 8 cases, for a molecular diagnostic rate of ~60%. Importantly, our results expand the mutational spectrum associated with eye malformations and demonstrates novel gene-phenotype correlations.
Materials and methods
Ethics statement and participants
The protocol was approved by the Institutional Review Board of the Institute of Ophthalmology “Conde de Valenciana”, at Mexico City. All procedures followed the tenets of the Helsinki Declaration and patients or their parents gave written permission for inclusion in the study. For each subject, exhaustive genealogical information was recorded and the occurrence of extraocular somatic defects and/or developmental abnormalities was investigated by a clinical geneticist.
A total of 14 unrelated Mexican probands affected by microphthalmia and/or anophthalmia were included in the study. Bilateral defects were observed in 12 patients. Four subjects had anophthalmia (bilateral in 1) and 13 presented with microphthalmia (bilateral in 6) (Table 1). Seven cases exhibited concomitant craniofacial, gastrointestinal, and/or neurologic anomalies and were classified as probably syndromic (Table 1). Familial transmission of the eye defect was observed in 7 cases (4 with presumed autosomal recessive inheritance (patients #3, 4, 9, and 11) and 3 with probable autosomal dominant transmission (patients #1, 8, and 10), by genealogy) while the remaining 7 cases were sporadic. Phenotypes of analyzed patients are summarized in Table 1. In order to exclude chromosomal anomalies, karyotype analysis was performed in subjects with concomitant severe malformations and intellectual disability (patients 6, 10, and 14 in Table 1) before NGS.
Next generation sequencing
DNA was extracted from peripheral blood leukocytes of all participants using QIAaMP DNA Blood kits (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. NGS library construction was performed using two commercial clinical exome kits according to manufacturer’s recommendations. As indicated in the Supplementary Table, a total of 83 genes involved in congenital eye defects were analyzed using either kit.
For patients 1–4, 7–9, and 13–14, library preparation and enrichment were performed using the Illumina TruSight One inherited disease panel (Illumina, San Diego, CA, USA). For these samples, DNA was enzymatically fragmented and purified. Index adapters were ligated to the 5′ and 3′ ends for subsequent amplification. Amplified fragments were hybridized to the Illumina TruSight One inherited disease panel that enables the enrichment of 4811 genes. Captured libraries were then purified and re-amplified. These samples were pooled in groups of three and sequenced using a MiSeq V3 Reagent Kit. For patients 5, 6, 10–12, libraries were prepared using the SureSelect QXT system (Agilent Technologies, Santa Clara, CA, USA) and enrichment was performed using the ClearSeq Inherited Disease panel, which includes probes that allow for the capture of the coding regions of 2742 genes. Briefly, DNA was enzymatically fragmented and DNA fragments were purified, amplified, and subsequently hybridized to the panel. For each sample, index adapters were ligated to the 5′ and 3′ ends. DNA fragments were re-amplified by PCR and fragments from 300 to 500 bp were isolated. Samples were pooled in groups of five and sequenced using the MiSeq Reagent kit v2 (300 cycles). All libraries were sequenced using the MiSeq platform (Illumina).
Bioinformatics and in silico analysis
The sequencing data were uploaded to the Galaxy web platform, and we used the public server at usegalaxy.org to analyze the data [22]. For optimal manipulation within the server, FASTQ files were converted to a Sanger FASTQ format using the tool FASTQ Groomer. Sequence data was read and mapped using the Burrows-Wheeler Aligner (BWA)-MEM algorithm, and VarScan 2.3.6 was used for variant calling. The human assembly GRCh37 (hg19) was used as reference for mapping and variant calling. Two VCF files were produced for each sample, one for SNVs and one for INDELS. Each VCF file was reviewed and filtered using the Illumina VariantStudio 3.0 software.
Variants were annotated for minor allele frequencies in the dbSNP [23], 1000 genomes [24], Exome Variant Server (NHLBI GO Exome Sequencing Project), and Exome Aggregation Consortium (ExAC) databases. Heterozygous variants with minor allele frequencies >0.05 were filtered out.
For missense variants, Polymorphism Phenotyping v2 (PolyPhen-2) [25] and Sorting Intolerant From Tolerant (SIFT) [26] algorithms were employed to predict pathogenicity.
Sanger sequencing
Sanger sequencing was carried out on probands to confirm variants identified as pathogenic by NGS. Co-segregation analysis of candidate pathogenic variants was subsequently performed on DNA from available family members. Primers for polymerase chain reaction (PCR) were designed using the PrimerQuest® program (Integrated DNA Technologies, Coralville, IA, USA) and are available upon request. PCR products were isolated using 1.5% agarose gels, purified, and sequenced using the BigDye Terminator Cycle sequencing kit (Applied Biosystems Foster City, CA, USA). All samples were analyzed in a 3130 Genetic Analyzer (Applied Biosystems).
Results
The average number of variants identified in samples enriched with the TruSight One panel was 32,359 ± 4732 with 94.27% ± 0.34 of these variants having a vertical coverage >10 ×. For samples enriched with the ClearSeq inherited disease panel, the average number of variants identified was 11,880 ± 3372 with 90.58% ± 2.52 of these variants having a vertical coverage >10. Pathogenic mutations were demonstrated in 8 out of the 14 (60%) analyzed cases. Four out of 8 (57%) solved cases were familial cases while the remaining four molecularly solved cases were sporadic (Table 1).
Patient #1
A novel heterozygous pathogenic variant in OTX2, c.249-1G>A, was identified in Patient #1, a 7-year-old male affected with right microphthalmia and left anophthalmia (Fig. 1). His mother (right microphthalmia) and his maternal grandmother (right microphthalmia) were also affected with congenital eye malformations, suggesting an autosomal dominant inheritance (Fig. 1a, b). No other physical anomalies or learning difficulties were present in the propositus or in his affected relatives. The novel OTX2 variant affects a canonical splice signal at intron 2 and is predicted to alter the mRNA splicing process. This variant is absent in general population databases (dbSNP, 1000 Genomes, EVS, EXAC), and from 240 in-house exomes alleles from Mexican individuals without ocular malformations. Co-segregation analysis confirmed that both, the patient’s maternal grandmother and mother carried the OTX2 pathogenic splicing variant (Fig. 1c).
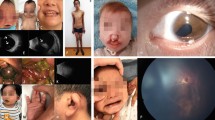
Familial microphthalmia/anophthalmia caused by a OTX2 pathogenic variant. a Simplified pedigree of patient #1 showing dominant transmission of congenital eye defects; patient #1, the proband, is indicated by an arrow. b Eye phenotypes in maternal grandmother (right microphthalmia), mother (right microphthalmia), and proband (microphthalmia OD/anophthalmia OS). c Partial Sanger sequencing of the OTX2 gene showing the heterozygous c.249-1 G > A splicing mutation (arrow) in affected family members
Patient #2
A homozygous c.667G>A (p.Gly223Arg) pathogenic variant in VSX2 was demonstrated in this case, a girl aged 1 year who was referred due to bilateral anophthalmia. She was product of a consanguineous marriage. Parents were healthy and no other relatives suffered from congenital eye defects. Extraocular anomalies were not demonstrated at physical examination and her developmental milestones were according to age. The p.Gly223Arg variant is absent in homozygous state from large-scale sequencing databases as ExAC and dbSNP, and also from a set of 240 in-house exomes alleles from Mexican individuals without ocular malformations. In silico analyses predicted pathogenicity of the VSX2 p.Gly223Arg variant.
Patient #3
A novel heterozygous c.6184C>T transition in the CHD7 gene (OMIM 608892), predicting a (p.Arg2062Trp) substitution, was identified in two siblings affected by anophthalmia/microphthalmia (Fig. 2). The proposita was a 24-year-old female with right microphthalmia and left anophthalmia (Fig. 2a). She did not exhibit additional physical anomalies and psychomotor development was normal. The patient had a 9-year-old brother with right microphthalmia and no anomalies on left eye (Fig. 2b). No additional somatic defects or intellectual disabilities were observed on clinical evaluation of these patients. Both parents were healthy, non-consanguineous, and had no ocular anomalies at ophthalmological examination.
Familial microphthalmia/anophthalmia due to a CHD7 pathogenic variant. a Microphthalmia (OD) and anophthalmia (OS) is observed in patient #3 while (b) right microphthalmia is evident in his brother. c–e Partial Sanger sequencing of the CHD7 gene showing the heterozygous c.6184C>T (p.Arg2062Trp) pathogenic variant. A smaller ‘T’ peak at the variant position is observed in genomic DNA from the unaffected mother (e), suggesting maternal mosaicism for the mutation
Patient #4
Compound heterozygous pathogenic variants in MFRP (OMIM 606227) were demonstrated in a 9-year-old boy with a diagnosis of bilateral simple microphthalmia (nanophthalmos). The identified variants were c.951C>A (exon 8), predicting a nonsense p.Tyr317* change, and c.1615 C>T (exon 13), predicting a missense p.Arg539Cys substitution. His 12-year-old brother was also affected by bilateral nanophthalmos, and genetic analysis by Sanger sequencing confirmed that he carried the same MFRP pathogenic variants. To exclude that both variants were located in the same allele (in cis), maternal DNA was analyzed demonstrating that she carried only the c.951C>A variant at exon 8. Paternal DNA was not available. No other ocular anomalies associated with MFRP defects as retinal dystrophy or optic disc drusen were observed in the patients.
Patient #5
In a 15-year-old boy with bilateral eye anterior segment dysgenesis, left microphthalmia and cataract, and glaucoma, a novel c.515C>T (p.Thr172Ile) heterozygous variant in VSX1 (OMIM 605020) was demonstrated (Fig. 3). This patient was adopted and no genetic analysis could be performed in his biological parents. He exhibited several facial anomalies including high nasal bridge, bulbous nasal tip, long, and smooth philtrum, wide mouth, micrognathia, and poorly developed antihelices (Fig. 3). Brain MRI disclosed hypoplastic corpus callosum as well as septum pellucidum absence. The patient had adequate verbal skills and normal height and weight measurements for his age.
Clinical and genetic features of patient #5. a Microphthalmia (OS) and bilateral anterior segment dysgenesis was observed. b Additional facial anomalies as convex and prominent nose, micrognathia, and poorly developed antihelices were also apparent. c Partial Sanger sequencing of the VSX1 gene indicating the heterozygous c.515C>T (p.Thr172Ile) variant (arrow). d Amino acid alignment of the VSX1 homeodomain from different species. The Thr172 residue is conserved in VSX1 proteins from different species
Patient #6
A previously reported c.1480C>T (p.Arg494*) nonsense heterozygous variant in CHD7 was identified in a 5-year-old girl with a history of bilateral microphthalmia, esophageal atresia, and psychomotor delay. Two days after her birth she underwent surgery for the esophageal defect. Bilateral optic disc coloboma were also noted. Parents were young, healthy, and denied a family history of congenital malformations. At examination, her stature and head circumference measurements were below the third centile. After NGS results, she received a final diagnosis of CHARGE syndrome (OMIM #214800). Parental DNA was not available for confirming a de novo origin of the CHD7 variant.
Patient #7
A novel heterozygous c.634G>A (p.Gly212Ser) variant in COL4A1 (OMIM 120130) was demonstrated in this 9-year-old boy who presented with bilateral microphthalmia and sclerocornea, and left eye glaucoma (Fig. 4a, b). At the age of 2 years he underwent enucleation of the right eye. He had a past history of convulsions that were treated with valproic acid, risperidone, and levetiracetam. His parents were healthy and there was no family history of congenital malformations or neurological anomalies. On brain MRI, white matter hyperintensities, suggestive of old infarcts, were observed (Fig. 4c, d). Based on genetic results and clinical reassessment, the final diagnosis of this patient was brain small vessel disease with leukoencelopathy and ocular anomalies (OMIM #607595). Parental DNA analysis showed wild type COL4A1 alleles, indicating a de novo origin of the pathogenic variant (Fig. 4e–g).
Phenotypic and genetic features of patient #7. a Enucleated OD and b microphthalmia and sclerocornea OS. c Brain MRI image at 7 years of age showing multiple periventricular white matter changes and extensive subcortical hyperintensity in the right parieto-occipital region. d Brain MRI at 8.5 years of age showing periventricular leukoencephalopathy, encephalomalacia around the right occipital horn, and secondary ventricular enlargement. e–g Partial Sanger sequencing of the COL4A1 gene demonstrating (e) the heterozygous c.634 G > A (p.Gly212Ser) variant (arrow) in DNA from the proband. Parental DNA analysis showed wild type COL4A1 sequences (f, g) indicating the de novo origin of the variant
Patient #8
A heterozygous c.701C>T, predicting p.Ser234Leu, in the PIEZO2 gene (OMIM 613629) was demonstrated in a 1-year-old girl with right eye sclerocornea and left eye microphthalmia, sclerocornea, and cornea plana. Her mother, 26 years of age, has a history of right sclerocornea and carried the same heterozygous variant in PIEZO2, suggesting a dominant transmission of the anomaly. At the age of 9 months, an ocular USG in the proposita demonstrated a funnel-like membrane arising from optic disc in the left eye, compatible with unilateral retinal detachment. No extraocular anomalies or intellectual disability were present in the proposita or in her affected mother. The p.Ser234Leu PIEZO2 variant was not observed by sanger sequencing in a set of 412 alleles from Mexican individuals without eye malformations or from a set of 140 in house exomes (280 alleles). In silico analyses of this variant using PredictSNP tool [27] indicated that while PolyPhen-1, PolyPhen-2, and SIFT classified it as deleterious, MAPP, PhD-SNP, and SNAP classified it as neutral. Additionally, Human Splicing Finder 3.1 predicts that the PIEZO2 c.701C>T variant affects a canonical splice site signal at exon 6.
Patients #9-#14
No obvious candidate variants were demonstrated after NGS of DNA samples from these 6 patients with congenital eye defects.
Discussion
Due to genetic heterogeneity associated to microphthalmia/anophthalmia of Mendelian basis, NGS approaches for simultaneous analysis of dozens of causal genes are a useful tool for genetic screening. The rate of molecularly solved cases differs from cohort to cohort and these differences are most probably due to factors as inclusion of familial vs sporadic cases, previous exclusion of mutations in major microphthalmia/anophthalmia genes (i.e., SOX2, RAX, OTX2, among others), and severity of included malformations. The occurrence of de novo mutations, mosaicism, and incomplete penetrance makes genetic counseling a challenge in these disorders [5].
In this work, a cohort of 14 Mexican patients with microphthalmia and/or anophthalmia were analyzed by NGS. A mutation detection rate of ~60% (8/14) was obtained demonstrating the utility of NGS for molecular diagnosis in these disorders. Seven out of 8 different mutations identified occurred in well-known eye defects genes (OTX2, VSX2, MFRP, VSX1) or in genes associated with syndromes that include ocular defects (CHD7, COL4A1) (including two instances of CHD7 pathogenic variants). A single pathogenic variant was identified in PIEZO2, a gene that was not previously associated with isolated ocular defects.
OTX2
Heterozygous mutations in OTX2 accounts for 2–8% of patients with anophthalmia/microphthalmia [18]. Orthodenticle homeobox 2 (OTX2) encodes a member of the bicoid subfamily of homeodomain-containing transcription factors. The encoded protein acts as a transcription factor and plays a role in brain, craniofacial, and sensory organ development. In a family with autosomal dominant segregation of anophthalmia/microphthalmia, a c.249-1G>A splicing mutation in OTX2 was identified. Mutations in OTX2 represents the second most common cause of anophthalmia/microphthalmia after SOX2 mutations [28]. To date, approximately 40 heterozygous OTX2 pathogenic variants have been identified in individuals with severe ocular malformations with or without brain and pituitary abnormalities [29]. Most pathogenic variants are frameshifting or introduce direct stop codons that predict a truncated protein product. To the best of our knowledge, this is the first demonstration of an OTX2 splicing mutation causing congenital eye malformations in humans. Intrafamilial clinical heterogeneity was observed in this family, as previously shown for other OTX2 pathogenic variants [30, 31].
VSX2
Visual system homeobox 2 gene (VSX2, previously known as CHX10) encodes a protein belonging to the paired-like (prd) class of homeobox transcription factor expressed in the optic vesicle and plays an important role in the development of the neural retina (NR) and retinal pigmented epithelium (RPE) in mammalian eye [32]. Pathogenic variants in VSX2 have been identified in a number of families with isolated microphthalmia or other ocular anomalies such as cataracts, colobomas or retinal dystrophy [33,34,35]. To date, approximately 20 biallelic VSX2 pathogenic variants have been identified in subjects with eye malformations, with missense mutations being the most frequently observed variations. In this work, NGS revealed a homozygous p.Gly223Arg VSX2 missense variant in a sporadic case of bilateral anophthalmia. The p.Gly223Arg mutation affects the evolutionarily conserved CVC motif that is needed for DNA binding and repression activities of VSX2. The VSX2 p.Gly223Arg variant has been previously identified in the heterozygous state in a bilateral microphthalmic patient carrying a truncating mutation in the other allele [12]. A similar homozygous variant, p.Gly223Ala, was identified by Reis et al. [36] in a Pakistani kindred with bilateral microphthalmia.
MFRP
Membrane frizzled-related protein (MFRP) is a type II transmembrane receptor possessing an extracellular frizzled-related cysteine-rich domain abundantly expressed in the RPE and ciliary body of the eye [37]. Mutations in MFRP are associated with nanophthalmos (simple microphthalmia) [38], posterior microphthalmia [39, 40], and a panocular phenotype including nanophthalmia, retinitis pigmentosa, foveoschisis, and optic disc drusen in humans [41, 42]. In a familial case of recessive simple microphthalmia studied here, compound heterozygosity for p.Tyr317* and p.Arg539Cys in MFRP was revealed in affected individuals. The Tyr317* variant was previously identified by our group in compound heterozygosity with c.498delC in two siblings of Mexican origin [43]. The p.Arg539Cys variant has been recently reported in at least two patients with nanophthamos and high hyperopia [44, 45].
VSX1
The VSX1 gene encodes a paired-like homeodomain transcription factor that is highly expressed in the embryonic craniofacial region, the granular layer of adult retina, and corneal tissue [46]. The 365-amino-acid encoded protein possesses a conserved homeobox DNA binding domain and a CVC domain (similar as those present in VSX2). Several VSX1 variants have been associated with the corneal disease keratoconus in subjects from different ethnicities [47, 48]. However, a definitive causal role of these variants has not yet been established because some of such variants can also occur in healthy individuals. Here, we identified a heterozygous p.Thr172Ile variant in VSX1 in a sporadic patient with anterior segment dysgenesis, left microphthalmia and craniofacial and brain malformations. The identified p.Thr172Ile variant is predicted to be deleterious by several in silico analyses, and it is absent from public databases as ExAC or dbSNP and also from a set of 280 alleles from Mexican individuals without eye malformations. In 2004, Mintz-Hittner et al. [49] described an African American family with dominant transmission of abnormal craniofacial features and anterior segment developmental anomalies in which a heterozygous p.Ala256Ser variant was demonstrated in all four affected relatives. The mutation was absent from control alleles and was predicted to result in a change of conformation and, therefore, activity of the protein. The Ala256 residue lies in the conserved CVC domain of VSX1. Thus, the sporadic patient described here represents the second instance of the syndrome of craniofacial anomalies and anterior segment dysgenesis (OMIM #614195). The Thr172 residue altered in our patient is located in the helix 1 of the homeodomain and it is invariably present in VSX1 proteins from vertebrates [50].
CHD7
The CHD7 gene (Chromo-Helicase-DNA binding protein 7) encodes a 2997 amino acid protein which is a member of the CHD family of ATP-dependent chromatin remodelers that employ the energy from ATP hydrolysis to mobilize or relocate nucleosomes. Thereby, such enzymes control DNA accessibility of chromatin and are critical for diverse processes such as transcription, DNA replication, and DNA repair (Reviewed in [51]). CHD7 mutations in humans result in CHARGE syndrome, a pleiotropic malformative disorder that includes cardiac, genital, and ocular anomalies, among others [52]. To date, more than 500 CHD7 mutations have been demonstrated in CHARGE syndrome patients, most of them corresponding to nonsense, frameshift, and splice mutations. CHD7 pathogenic variants have been also identified in about 5% of subjects with Kallmann syndrome (OMIM #612370), a clinical entity defined by the association of hypogonadotropic hypogonadism and olfactory deficiency [53]. Most CHD7 mutations in Kallmann syndrome are missense variants. In the present work, NGS approach allowed the identification of a heterozygous p.Arg2062Trp missense variant in CHD7 in a subject (patient #3) with anophthalmia/microphthalmia and in her similarly affected brother. Clinical examination of these patients excluded extraocular malformations, defective sense of smell, or intellectual disability. This variant is absent from public databases such as ExAC, 1000 Genomes, dbSNP and from more than 500 alleles from a cohort of Mexican individuals without eye malformations. Analysis of electropherogram traces from parental DNA indicated that the mother was a mosaic for the CHD7 p.Arg2062Trp variant. However, no additional maternal tissues were available for further analysis. Both lack of penetrance and germinal mosaicism have been previously described for CHD7 pathogenic variants [54, 55]. To our knowledge, this is the first demonstration of CHD7 pathogenic variants resulting in isolated ocular anomalies. Thus, our results provide a novel gene-phenotype correlation for CHD7. In an additional patient (#6) with ocular malformations and a history of esophageal atresia, a CHD7 p.Arg494* nonsense mutation was identified. This variant has been previously reported in CHARGE syndrome patients [56, 57]. Based on genetic results, the final diagnosis in this patient was CHARGE syndrome.
COL4A1
COL4A1 gene encodes the most abundant and ubiquitous basement membrane protein. In the eye, COL4A1 is widely expressed in conjunctiva, corneal epithelium and endothelium, and trabecular meshwork, among other localizations [58, 59]. Pathogenic variants in COL4A1 results in a wide spectrum of malformations that includes familial porencephaly [60], cerebral white matter small vessel disease [61], cataract, anterior segment dysgenesis, microcornea [62], microphthalmia [63], and Walker Warburg syndrome [64], among others.
In this work, a novel p.Gly212Ser COL4A1 missense mutation was identified in a patient referred due to microphthalmia, sclerocornea, and epilepsy. According to the genetic results, the final diagnosis of this patient was brain small vessel disease with leukoencelopathy and ocular anomalies (OMIM #607595). The p.Gly212Ser replacement occurs at the N-terminus of the triple-helix-forming domain and to the best of our knowledge it has not been previously reported in COL4A1-related diseases.
PIEZO2
Mechanotransduction involves a variety of processes that convert mechanical stimuli in biological signals. Mechanotransduction is important for processes of sensorial perception such as pain, touch, or hearing, but it has also been implicated in embryonic development of organs and tissues (reviewed in [65]). The PIEZO2 gene encodes a 2521 amino acid protein with several transmembrane domains, which functions as a mechanically activated cation channel involved in adapting mechanically activated currents in somatosensory neurons [66]. In humans, pathogenic variants in PIEZO2 are associated with diverse phenotypes mainly affecting craniofacial structures and limbs, including Marden-Walker syndrome (MIM #248700), and distal arthogryposes types 3 (OMIM #114300) and 5 (OMIM # 108145) [67]. In a familial case of microphthalmos/sclerocornea (mother and daughter affected) included in the present study, a heterozygous c.701C>T PIEZO2 variant was demonstrated in both affected patients. This variant results in the substitution of Serine 234, a polar uncharged amino acid residue of medium size, for leucine, a large hydrophobic amino acid, at the extreme end of a helix transmembrane domain (http://www.uniprot.org/). In rodents, it has been shown that PIEZO2 is expressed in corneal afferent neurons [68] and astrocytes in the optic nerve head [69]. Furthermore, recent work has demonstrated that PIEZO2 is extensively spliced, producing up to 17 different isoforms with specific cell type expression patterns in trigeminal ganglion neurons [70]. Therefore, since PIEZO2 relies heavily on splicing to produce so many isoforms in a tissue and cell specific manner, it is possible that the c.701 C>T change affects the processing of specific isoforms particularly expressed in optic tissue. It is interesting to note that, microphthalmia as well as corneal anomalies including keratoconus and keratoglobus have been reported in cases of Marden-Walker syndrome and distal arthrogryposis type 5 [71,72,73], two PIEZO2-related entities. Although it is tempting to argue that these data could support the association of corneal and/or eye defects with PIEZO2 variants, to our knowledge no additional patients with isolated congenital eye defects carrying a PIEZO2 variant have been reported. In this study, the c.701 C>T PIEZO2 variant segregates with the disease and was absent from more than 120,000 PIEZO2 alleles from ExAC database and from more than 650 PIEZO2 alleles from a cohort of Mexican individuals without eye malformations. A limitation of our study is that no functional studies on PIEZO2 were performed to support the causality of this gene variation. Thus additional reports and/or experimental data will be needed to confirm our findings. As two p.Ser234Leu PIEZO2 alleles are registered in gnomAD database (2/140,016 alleles) it is possible that this variant is associated with non-penetrance. It is also possible that the mutation we identified in this family is not related to their eye phenotype but is rather a coincidental finding.
Finally, in 6 out of 14 probands with microphthalmia and/or anophthalmia, including two autosomal recessive and one autosomal dominant cases, no obvious candidate pathogenic variants were recognized after NGS. Causal variants in negative cases could be located at regions not covered by NGS (deep intronic, regulatory sequences), could consist in structural variations as CNVs, missed by standard bioinformatic analysis of NGS results, or could be variants located at currently unknown ocular malformation genes. Obviously, some of these cases are not monogenic in nature.
NGS efficiently identified a monogenic etiology of microphthalmia/anophthalmia in ~60% of cases included in this cohort, the first from Mexican origin analyzed to date. The molecular defects identified through exome sequencing in this study expands the phenotypic spectra of CHD7-associated disorders and implicate PIEZO2 as a candidate gene for major eye developmental defects. Although our group of study is small, our results highlight the value of continued molecular screening of cohorts of individuals with developmental eye defects from distinct ethnic groups, as this will provide more realistic figures, not only for the global monogenic causes of congenital eye defects, but also for the contribution of particular genes to the phenotype.
References
Graw J. Eye development. Curr Top Dev Biol. 2010;90:343–86.
Sinn R, Wittbrodt J. An eye on eye development. Mech Dev. 2013;130:347–58.
Graw J, Löster J. Developmental genetics in ophthalmology. Ophthalmic Genet. 2003;24:1–33.
Horsford DJ, Hanson I, Freund C, McInnes RR, van Heyningen V. Transcription factors in eye disease and ocular development. In. Valle D, Beaudet AL, Vogelstein B, et al. editors. Chapter 240. The Online Metabolic & Molecular Bases of Inherited disease, McGraw-Hill, New York, USA. (https://ommbid.mhmedical.com, accesed on February, 2018).
Morrison D, FitzPatrick D, Hanson I, Williamson K, van Heyningen V, Fleck B, et al. National study of microphthalmia, anophthalmia, and coloboma (MAC) in Scotland: investigation of genetic aetiology. J Med Genet. 2002;39:16–22.
Slavotinek AM. Eye development genes and known syndromes. Mol Genet Metab. 2011;104:448–56.
Skalicky SE, White AJ, Grigg JR, Martin F, Smith J, Jones M, et al. Microphthalmia, anophthalmia, and coloboma and associated ocular and systemic features: understanding the spectrum. JAMA Ophthalmol. 2013;131:1517–24.
Freund C, Horsford DJ, McInnes RR. Transcription factor genes and the developing eye: a genetic perspective. Hum Mol Genet. 1996;5(Spec No):1471–88.
Graw J. The genetic and molecular basis of congenital eye defects. Nat Rev Genet. 2003;4:876–88.
Patel A, Sowden JC. Genes and pathways in optic fissure closure. Semin Cell Dev Biol. 2017; pii: S1084-9521(17)30149-0.
Verma AS, Fitzpatrick DR. Anophthalmia and microphthalmia. Orphanet J Rare Dis. 2007;2:47.
Chassaing N, Causse A, Vigouroux A, Delahaye A, Alessandri JL, Boespflug-Tanguy O, et al. Molecular findings and clinical data in a cohort of 150 patients with anophthalmia/microphthalmia. Clin Genet. 2014;86:326–34.
Williamson KA, FitzPatrick DR. The genetic architecture of microphthalmia, anophthalmia and coloboma. Eur J Med Genet. 2014;57:369–80.
Gonzalez-Rodriguez J, Pelcastre EL, Tovilla-Canales JL, Garcia-Ortiz JE, Amato-Almanza M, Villanueva-Mendoza C, et al. Mutational screening of CHX10, GDF6, OTX2, RAX and SOX2 genes in 50 unrelated microphthalmia-anophthalmia-coloboma (MAC) spectrum cases. Br J Ophthalmol. 2010;94:1100–4.
Gerth-Kahlert C, Williamson K, Ansari M, Rainger JK, Hingst V, Zimmermann T, et al. Clinical and mutation analysis of 51 probands with anophthalmia and/or severe microphthalmia from a single center. Mol Genet Genomic Med. 2013;1:15–31.
Riera M, Wert A, Nieto I, Pomares E. Panel-based whole exome sequencing identifies novel mutations in microphthalmia and anophthalmia patients showing complex Mendelian inheritance patterns. Mol Genet Genomic Med. 2017;5:709–19.
Richardson R, Sowden J, Gerth-Kahlert C, Moore AT, Moosajee M. Clinical utility gene card for: Non-Syndromic Microphthalmia Including Next-Generation Sequencing-Based Approaches. Eur J Hum Genet. 2017;25.
Jimenez NL, Flannick J, Yahyavi M, Li J, Bardakjian T, Tonkin L, et al. Targeted ‘next-generation’ sequencing in anophthalmia and microphthalmia patients confirms SOX2, OTX2 and FOXE3 mutations. BMC Med Genet. 2011;12:172.
Slavotinek AM, Garcia ST, Chandratillake G, Bardakjian T, Ullah E, Wu D, et al. Exome sequencing in 32 patients with anophthalmia/microphthalmia and developmental eye defects. Clin Genet. 2015;88:468–73.
Prokudin I, Simons C, Grigg JR, Storen R, Kumar V, Phua ZY, et al. Exome sequencing in developmental eye disease leads to identification of causal variants in GJA8, CRYGC, PAX6 and CYP1B1. Eur J Hum Genet. 2014;22:907–15.
Deml B, Reis LM, Lemyre E, Clark RD, Kariminejad A, Semina EV. Novel mutations in PAX6, OTX2 and NDP in anophthalmia, microphthalmia and coloboma. Eur J Hum Genet. 2016;24:535–41.
Afgan E, Baker D, van den Beek M, Blankenberg D, Bouvier D, Čech M, et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res. 2016;44(W1):W3–W10.
Smigielski EM, Sirotkin K, Ward M, Sherry ST. dbSNP: a database of single nucleotide polymorphisms. Nucleic Acids Res. 2000;28:352–5.
1000 Genomes Project Consortium, Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, et al. An integrated map of genetic variation from 1,092 human genomes. Nature. 2012;491:56–65.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009;4:1073–81.
Bendl J, Stourac J, Salanda O, Pavelka A, Wieben ED, Zendulka J, et al. PredictSNP: robust and accurate consensus classifier for prediction of disease-related mutations. PLoS Comput Biol. 2014;10:e1003440.
Schilter KF, Schneider A, Bardakjian T, Soucy JF, Tyler RC, Reis LM, et al. OTX2 microphthalmia syndrome: four novel mutations and delineation of a phenotype. Clin Genet. 2011;79:158–68.
Ragge NK, Brown AG, Poloschek CM, Lorenz B, Henderson RA, Clarke MP, et al. Heterozygous mutations of OTX2 cause severe ocular malformations. Am J Hum Genet. 2005;76:1008–22.
You T, Lv Y, Liu S, Li F, Zhao Y, Lv J, et al. Novel OTX2 mutation associated with congenital anophthalmia and microphthalmia in a Han Chinese family. Acta Ophthalmol. 2012;90:e501–2.
Somashekar PH, Shukla A, Girisha KM. Intrafamilial variability in syndromic microphthalmia type 5 caused by a novel variation in OTX2. Ophthalmic Genet. 2017;38:533–6.
Liu IS, Chen JD, Ploder L, Vidgen D, van der Kooy D, Kalnins VI, et al. Developmental expression of a novel murine homeobox gene (Chx10): evidence for roles in determination of the neuroretina and inner nuclear layer. Neuron. 1994;13:377–93.
Ferda Percin E, Ploder LA, Yu JJ, Arici K, Horsford DJ, Rutherford A, et al. Human microphthalmia associated with mutations in the retinal homeobox gene CHX10. Nat Genet. 2000;25:397–401.
Bar-Yosef U, Abuelaish I, Harel T, Hendler N, Ofir R, Birk OS. CHX10 mutations cause non-syndromic microphthalmia/ anophthalmia in Arab and Jewish kindreds. Hum Genet. 2004;115:302–9.
Iseri SU, Wyatt AW, Nurnberg G, Kluck C, Nurnberg P, Holder GE, et al. Use of genome-wide SNP homozygosity mapping in small pedigrees to identify new mutations in VSX2 causing recessive microphthalmia and a semidominant inner retinal dystrophy. Hum Genet. 2010;128:51–60.
Reis LM, Khan A, Kariminejad A, Ebadi F, Tyler RC, Semina EV. VSX2 mutations in autosomal recessive microphthalmia. Mol Vis. 2011;17:2527–32.
Kameya S, Hawes NL, Chang B, Heckenlively JR, Naggert JK, Nishina PM. Mfrp, a gene encoding a frizzled related protein, is mutated in the mouse retinal degeneration 6. Hum Mol Genet. 2002;11:1879–86.
Sundin OH, Leppert GS, Silva ED, Yang JM, Dharmaraj S, Maumenee IH, et al. Extreme hyperopia is the result of null mutations in MFRP, which encodes a Frizzled-related protein. Proc Natl Acad Sci USA. 2005;102:9553–8.
Matsushita I, Kondo H, Tawara A. Novel compound heterozygous mutations in the MFRP gene in a Japanese patient with posterior microphthalmos. Jpn J Ophthalmol. 2012;56:396–400.
Said MB, Chouchène E, Salem SB, Daoud K, Largueche L, Bouassida W, et al. Posterior microphthalmia and nanophthalmia in Tunisia caused by a founder c.1059_1066insC mutation of the PRSS56 gene. Gene. 2013;528:288–94.
Ayala-Ramirez R, Graue-Wiechers F, Robredo V, Amato-Almanza M, Horta-Diez I, Zenteno JC. A new autosomal recessive syndrome consisting of posterior microphthalmos, retinitis pigmentosa, foveoschisis, and optic disc drusen is caused by a MFRP gene mutation. Mol Vis. 2006;12:1483–9.
Mukhopadhyay R, Sergouniotis PI, Mackay DS, Day AC, Wright G, Devery S, et al. A detailed phenotypic assessment of individuals affected by MFRP-related oculopathy. Mol Vis. 2010;16:540–8.
Zenteno JC, Buentello-Volante B, Quiroz-González MA, Quiroz-Reyes MA. Compound heterozygosity for a novel and a recurrent MFRP gene mutation in a family with the nanophthalmos-retinitis pigmentosa complex. Mol Vis. 2009;15:1794–8.
Xu Y, Guan L, Xiao X, Zhang J, Li S, Jiang H, et al. Identification of MFRP mutations in chinese families with high hyperopia. Optom Vis Sci. 2016;93:19–26.
Velez G, Tsang SH, Tsai YT, Hsu CW, Gore A, Abdelhakim AH, et al. Gene therapy restores Mfrp and corrects axial eye length. Sci Rep. 2017;7:16151.
Semina EV, Mintz-Hittner HA, Murray JC. Isolation and characterization of a novel human paired-like homeodomain-containing transcription factor gene, VSX1, expressed in ocular tissues. Genomics. 2000;63:289–93.
Tanwar M, Kumar M, Nayak B, Pathak D, Sharma N, Titiyal JS, et al. VSX1 gene analysis in keratoconus. Mol Vis. 2010;16:2395–401.
Dash DP, George S, O’Prey D, Burns D, Nabili S, Donnelly U, et al. Mutational screening of VSX1 in keratoconus patients from the European population. Eye (Lond). 2010;24:1085–92.
Mintz-Hittner HA, Semina EV, Frishman LJ, Prager TC, Murray JC. VSX1 (RINX) mutation with craniofacial anomalies, empty sella, corneal endothelial changes, and abnormal retinal and auditory bipolar cells. Ophthalmology. 2004;111:828–36.
Chow RL, Snow B, Novak J, Looser J, Freund C, Vidgen D, et al. Vsx1, a rapidly evolving paired-like homeobox gene expressed in cone bipolar cells. Mech Dev. 2001;109:315–22.
Clapier CR, Cairns BR. The biology of chromatin remodeling complexes. Annu Rev Biochem. 2009;78:273–304.
Vissers LE, van Ravenswaaij CM, Admiraal R, Hurst JA, de Vries BB, Janssen IM, et al. Mutations in a new member of the chromodomain gene family cause CHARGE syndrome. Nat Genet. 2004;36:955–7.
Kim HG, Kurth I, Lan F, Meliciani I, Wenzel W, Eom SH, et al. Mutations in CHD7, encoding a chromatin-remodeling protein, cause idiopathic hypogonadotropic hypogonadism and Kallmann syndrome. Am J Hum Genet. 2008;83:511–9.
Jongmans MC, Hoefsloot LH, van der Donk KP, Admiraal RJ, Magee A, van de Laar I, et al. Familial CHARGE syndrome and the CHD7 gene: a recurrent missense mutation, intrafamilial recurrence and variability. Am J Med Genet A. 2008;146A:43–50.
Pauli S, Pieper L, Häberle J, Grzmil P, Burfeind P, Steckel M, Lenz U, Michelmann HW. Proven germline mosaicism in a father of two children with CHARGE syndrome. Clin Genet. 2009;75:473–9.
Legendre M, Gonzales M, Goudefroye G, Bilan F, Parisot P, Perez MJ, et al. Antenatal spectrum of CHARGE syndrome in 40 fetuses with CHD7 mutations. J Med Genet. 2012;49:698–707.
Janssen N, Bergman JE, Swertz MA, Tranebjaerg L, Lodahl M, Schoots J, et al. Mutation update on the CHD7 gene involved in CHARGE syndrome. Hum Mutat. 2012;33:1149–60.
Ishizaki M, Westerhausen-Larson A, Kino J, Hayashi T, Kao WW. Distribution of collagen IV in human ocular tissues. Invest Ophthalmol Vis Sci. 1993;34:2680–89.
Hann CR, Springett MJ, Wang X, Johnson DH. Ultrastructural localization of collagen IV, fibronectin, and laminin in the trabecular meshwork of normal and glaucomatous eyes. Ophthalmic Res. 2001;33:314–24.
Gould DB, Phalan FC, Breedveld GJ, van Mil SE, Smith RS, Schimenti JC, et al. Mutations in COL4A1 cause perinatal cerebral hemorrhage and porencephaly. Science. 2005;308:1167–71.
Gould DB, Phalan FC, vanMil SE, Sundberg JP, Vahedi K, Massin P, et al. Role of COL4A1 in small-vessel disease and hemorrhagic stroke. N Engl J Med. 2006;354:1489–96.
Coupry I, Sibon I, Mortemousque B, Rouanet F, Mine M, Goizet C. Ophthalmological features associated with COL4A1 mutations. Arch Ophthalmol. 2010;128:483–9.
Deml B, Reis LM, Maheshwari M, Griffis C, Bick D, Semina EV. Whole exome analysis identifies dominant COL4A1 mutations in patients with complex ocular phenotypes involving microphthalmia. Clin Genet. 2014;86:475–81.
Labelle-Dumais C, Dilworth DJ, Harrington EP, de Leau M, Lyons D, Kabaeva Z, et al. COL4A1 mutations cause ocular dysgenesis, neuronal localization defects, and myopathy in mice and Walker-Warburg syndrome in humans. PLoS Genet. 2011;7:e1002062.
Fernandez-Sanchez ME, Brunet T, Röper JC, Farge E. Mechanotransduction’s impact on animal development, evolution, and tumorigenesis. Annu Rev Cell Dev Biol. 2015;31:373–97.
Geng J, Zhao Q, Zhang T, Xiao B. In Touch with the mechanosensitive Piezo channels: structure, ion Permeation, and mechanotransduction. Curr Top Membr. 2017;79:159–95.
McMillin MJ, Beck AE, Chong JX, Shively KM, Buckingham KJ, Gildersleeve HI, et al. Mutations in PIEZO2 cause Gordon syndrome, Marden-Walker syndrome, and distal arthrogryposis type 5. Am J Hum Genet. 2014;94:734–44.
Bron R, Wood RJ, Brock JA, Ivanusic JJ. Piezo2 expression in corneal afferent neurons. J Comp Neurol. 2014;522:2967–79.
Choi HJ, Sun D, Jakobs TC. Astrocytes in the optic nerve head express putative mechanosensitive channels. Mol Vis. 2015;21:749–6.
Szczot M, Pogorzala LA, Solinski HJ, Young L, Yee P, Le Pichon CE, et al. Cell-type-specific splicing of Piezo2 regulates mechanotransduction. Cell Rep. 2017;21:2760–71.
Schrander-Stumpel C, de Die-Smulders C, de Krom M, Schyns-Fleuran S, Hamel B, Jaeken D, et al. Marden-Walker syndrome: case report, literature review and nosologic discussion. Clin Genet. 1993;43:303–8.
Sahni J, Kaye SB, Fryer A, Hiscott P, Bucknall RC. Distal arthrogryposis type IIB: unreported ophthalmic findings. Am J Med Genet A. 2004;127A:35–9.
Güell JL, Verdaguer P, Elies D, Gris O, Manero F. Corneal impairment in a patient with type 2 distal arthrogryposis. Eye Contact Lens. 2015;41:e5–8.
Acknowledgements
This work was partially supported by CONACYT grants 234413 and 233967. We would like to thank Daniel Moreno, M.D and Roger Fest-Parra, M.D for technical assistance, and Dr. Eduardo Perez-Campos for helpful observations on protocol organization.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Matías-Pérez, D., García-Montaño, L.A., Cruz-Aguilar, M. et al. Identification of novel pathogenic variants and novel gene-phenotype correlations in Mexican subjects with microphthalmia and/or anophthalmia by next-generation sequencing. J Hum Genet 63, 1169–1180 (2018). https://doi.org/10.1038/s10038-018-0504-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-018-0504-1
This article is cited by
-
Prenatal diagnosis of distal 13q deletion syndrome in a fetus with esophageal atresia: a case report and review of the literature
Journal of Medical Case Reports (2022)
-
Genetics of anophthalmia and microphthalmia. Part 1: Non-syndromic anophthalmia/microphthalmia
Human Genetics (2019)