Abstract
Mutation in the gene encoding microphthalmia-associated transcription factor (MITF) lead to Waardenburg syndrome 2 (WS2), an autosomal dominantly inherited syndrome with auditory-pigmentary abnormalities, which is clinically and genetically heterogeneous. Haploinsufficiency may be the underlying mechanism for WS2. However, the mechanisms explaining the genotypic and phenotypic variations in WS2 caused by MITF mutations are unclear. A previous study revealed that MITF interacts with LEF-1, an important factor in the Wnt signaling pathway, to regulate its own transcription through LEF-1-binding sites on the MITF promoter. In this study, four different WS2-associated MITF mutations (p.R217I, p.R217G, p.R255X, p.R217del) that are associated with highly variable clinical features were chosen. According to the results, LEF-1 can activate the expression of MITF on its own, but MITF proteins inhibited the activation. This inhibition weakens when the dosage of MITF is reduced. Except for p.R217I, p.R255X, p.R217G, and p.R217del lose the ability to activate TYR completely and do not inhibit the LEF-1-mediated activation of the MITF-M promoter, and the haploinsufficiency created by mutant MITF can be overcome; correspondingly, the mutants’ associated phenotypes are less severe than that of p.R217I. The dominant negative of p.R217del made it have a second-most severe phenotype. This study’s data imply that MITF has a negative feedback loop of regulation to stabilize MITF gene dosage that involves the Wnt signaling pathway and that the interaction of MITF mutants with this pathway drives the genotypic and phenotypic differences observed in Waardenburg syndrome type 2 associated with MITF mutations.
Similar content being viewed by others
Introduction
Waardenburg syndrome (WS) is the most common autosomal dominantly inherited syndromic hearing loss, and it is clinically and genetically heterogeneous [1,2,3]. The seven genes including MITF, PAX3, SOX10, SNAI2, EDN3, EDNRB, and KITLG are involved in this syndrome; the first four are transcription factors, and the latter three are signal molecules [4,5,6]. Clinically, WS is divided into four subtypes (WS1-4) based on the presence or absence of additional symptoms with associated auditory-pigmentary abnormalities due to a lack of melanin [7, 8]. WS2 is characterized by the absence of additional symptoms. In humans, WS2 is the most common subtype, and approximately 15% of WS2 is caused by heterozygous mutation of the gene encoding microphthalmia-associated transcription factor (MITF) [4, 9]. Rarely, MITF mutations lead to Tietz syndrome, which has a more severe phenotype of hearing loss and generalized, albinoid-like hypopigmentation of the skin and hair from birth instead of the patchy depigmentation observed in WS [3, 4, 10, 11]. Otherwise, mutations in MITF can also lead to different phenotypes such as melanoma [12].
MITF belongs to the Myc super family of b-HLH-Zip proteins, and there are at least nine different isoforms with differences in the promoters and first exons [6, 13]. The MITF-M isoform is exclusively expressed in melanocytes and melanoma cells [6, 9]. MITF is involved in the differentiation and development of neural crest cells (NCCs), and it plays a very important role in the survival, migration, differentiation, and development of melanocytes [14,15,16,17]. MITF can directly activate the expression of the target gene TYR, but it must interact with other proteins to regulate dopachrome tautomerase (DCT) expression [18, 19]. MITF mutations affect the transcription of TYR and DCT and lead to a reduction of melanin (haploinsufficiency) and the different genotypic variations in WS2, consistent with our previous report [12, 20,21,22].
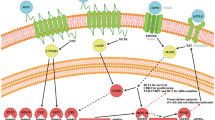
Studies have shown that mouse MITF acts as a transcription factor to regulate its own gene expression by recruiting LEF-1/β-catenin (two important factors in the Wnt signaling) to its promoter, where MITF and LEF-1 form a protein complex that enhances transcription from this promoter, TYR gene promoter and the DCT gene promoter [23,24,25,26,27]. The molecular association of Wnt, MITF, TYR, and DCT was shown in Fig. 1. However, it is unclear whether mutated human MITF affects Wnt signaling and influences the interaction between MITF and LEF-1 and the expression of MITF, and DCT to establish the different phenotypes of WS2.
We chose to study the MITF mutants p.R255X (R255X), p.R217I (R217I), p.R217G (R217G), and p.R217del (R217del) [28, 29], which are all observed in China, to explore the effects of mutant MITF on Wnt signaling and the interaction between MITF and LEF-1.
Materials and methods
Plasmid construction
Luciferase reporters containing the human TYR promoter (pGL3-TYR-Luc) and MITF-M promoter (pGL3-MITF-Luc) were kindly provided by Jiri Vachtenheim et al. [30] (Czech), Bondurand Nadege et al. [31] (French), respectively. The expression vector pCMV-MITF-Flag was described previously [22]. The mutants R255X, R217I, R217G, and R217del were generated using Quikchange II site-directed mutagenesis (GE Healthcare Chalfont St. Giles, Buckinghamshire, UK) from the template pCMV-MITF-Flag. The expression vector pCDNA3.1-LEF-1-HA containing full-length human LEF-1 complementary DNA (cDNA; GenBank Accession No: NM_016269.4) was described previously [27]. The luciferase reporter containing the human DCT gene promoter (PGL3-DCT-Luc) was generated by the Nanjing Genescript Biotechnology Company (China). All plasmids were confirmed by automatic sequencing analysis.
Mammalian cell culture, transfections, and luciferase reporter assays
The HEK293T (293T) cells and the human melanoma UACC903 cells were maintained in DMEM (high glucose) supplemented with 10% fetal bovine serum (FBS) and 100 u/ml of penicillin/streptomycin as described. Cells were grown at 37 °C in 5% CO2. All transient transfection assays were performed using Lipofectamine 2000 (Invitrogen Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s instructions. Cells were grown at an approximate 50% confluency in 24-well plates for approximately 24 h. Then, the cells were transfected with 5 ng of the reporter plasmids, 20 ng of expression vector, and 5 ng of pCMV-β-gal (BD Biosciences/Clontech, Palo Alto, CA, USA). The final DNA amount in each well was adjusted to 200 ng with empty vectors. At 48 h after transfection, cells were washed with 1× PBS and lysed with 1× reporter lysis buffer (Promega, Madison, WI, USA). The extracts were assayed for luciferase and β-galactosidase activity. Luciferase reporter assays were performed using a luciferase assay system (Promega, Madison, WI, USA)) according to the manufacturer’s protocol. Luciferase activity was measured using a SIRIUS luminometer (Berthold Detection Systems GmbH, Pforzheim, Germany) and normalized to β-galactosidase activity. Relative luciferase activity is shown as the ratio of each normalized luciferase activity to the value obtained with pGL3-TYR-Luc and empty vector. For competition assays, various amounts of MITF expression plasmids (1, 5, 10, 20, 50, 100 ng) were mixed with fixed amounts of LEF-1 (100 ng) and the reporter MITF plasmid (5 ng) for transfection. All reporter assays were conducted at least three times and performed in triplicate on different days using different batches of cells. The data were analyzed using GraphPad Prism 5 software (GraphPad software, Inc., San Diego, CA, USA).
In vitro co-immunoprecipitation assays
In all, 293T cells growing in a 100-mm plate were transfected at approximately 80% confluency with 6 µg of pcDNA-LEF-1-HA and 6 µg of pCMV-MITF-Flag or its mutants using Lipofectamine 2000 (Invitrogen Life Technologies, Carlsbad, CA, USA). At 36–48 h after transfection, the co-immunoprecipitation were performed as reported previously [32]. The immunoprecipitation (IP) samples were incubated with 1 µg anti-Flag M2 monoclonal antibody (Sigma, St. Louis, WA, USA, F1804), and the negative control was incubated with 1 µg normal mouse IgG as a negative control. The anti-HA rabbit antibody (1:1000 dilution, Cell Signaling Technology, Boston, MA, USA, 3724s) was used in the immunoblot analysis.
Biotinylated DNA affinity precipitation
The LEF-1-binding oligonucleotide derived from the promoter of the MITF gene, F: 5′-TTGGCCTTGATCTGACAGTGAGTTTGACTTTATAGCTCGTC-3′, was synthesized, biotinylated at the 5-terminus, and then annealed with its complementary strand to generate double-stranded oligonucleotides. The assays were performed as previously described [32]. In all, 293T cells were used with anti-HA rabbit antibody (1:1000 dilution, Cell Signaling Technology, Boston, MA, USA, 3724s).
Results
Mutations of MITF in WS2 and their phenotype
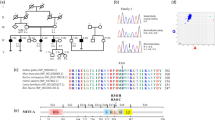
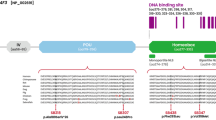
Four heterozygosis mutations, c.763 C>T (p.R255X), c.650 G>T (p.R217I), c.649 A>G (p.R217G), and c.647_649del (p.R217del), have been identified in the MITF gene in four WS2 cases in China [28, 29]. They all presented with deafness and heterochromia iridis due to a lack of melanin, but the severity varied. Their genotypes and phenotypes are shown in Supplementary Fig S1.
Transactivation of the TYR promoter by WT or mutant MITF
MITF plays a very important role in the survival, migration, differentiation, and development of melanocytes [14,15,16,17]. It is the key transcription factor and master regulator of melanin development. MITF can upregulate the transcription of the target genes TYR, TYRP1, and TYRP2/DCT through the E-box (CANNTG) located in their promoters and consequently increase the expression of the melanocyte-specific enzyme tyrosinase [19, 21, 33,34,35]. The study has shown the phenotypic heterogeneity by MITF gene mutation was associated with transcription activation [12]. To investigate whether mutant MITF affects the transcription activation of TYR, we co-transfected the wild-type or mutant MITF with the luciferase reporter containing the TYR promoter in HEK293T cells and UACC903 cells. As indicated in Fig. 2a, b, WT MITF enhances TYR promoter activity approximately sixfold in 293T cells and 25-fold in UACC903 cells. However, the mutants R255X, R217G, and R217del nearly abolished the transactivation of the TYR promoter. R217I retained partial transactivation of the TYR promoter in the two cell types, as reported previously [12, 22].
Effects of MITF and its mutants on TYR promoter activity. The luciferase reporter plasmid TYR-Luc was transiently transfected into 293T cells (a) or melanoma UACC903 cells (b) in combination with WT or mutant MITF expression vectors. The basal level of luciferase was set as 1. Data from all other transfections are presented as fold induction above this level. Luciferase activity was normalized to β-galactosidase activity. Each value shown is the mean ± SD of three replicates from a single assay. The results shown are representative of at least three independent experiments (**p < 0.01,***p < 0.001 compared with the basal activity, ###p < 0.001 compared with the WT MITF, unpaired Student’s t-test)
Functional interaction between LEF-1 and MITF-M on the MITF and DCT promoters
MITF and LEF-1 form a protein complex and interact with gene promoters to regulate the transcription of DCT and MITF-M [23,24,25]. Luciferase assays were performed to determine whether the mutant MITF would disrupt the interaction between MITF and LEF-1. As shown in Fig. 3a, decreased transcriptional activity from the DCT promoter was detected with MITF or LEF-1 alone, but when they were co-expressed in 293T cells, the DCT promoter activity was dramatically increased, which is consistent with a previous report [23]. Mutant MITF destroyed this synergistic transcriptional activation of the DCT promoter by LEF-1 and MITF. However, the synergistic transactivation of the DCT promoter by MITF and LEF-1 was not observed in UACC903 cells. LEF-1 alone can also transactivate the MITF promoter, but this activity is dramatically reduced in combination with MITF in 293T cells (Fig. 3c), and the extent of this reduction increased as the MITF dosage increased (Fig. 3c). We did not observe the same effect on MITF promoter transactivation with the R255X, R217G, and R217del mutants as seen with wild-type MITF, but the R217I mutant produced similar results (Fig. 3d, e).
Functional interaction between LEF-1 and WT/mutant MITF-M on the MITF promoter and DCT promoter based on luciferase reporter assays. a, b The transactivation of the DCT promoter by LEF-1 and WT/mutant MITF. In all, 293T cells (a) and UACC903 cells (b) were transiently co-transfected with DCT-Luc reporter plasmid, LEF-1, and WT/mutant MITF. c LEF-1-mediated activation of the MITF-M promoter was dramatically reduced in combination with MITF in 293T cells. The luciferase reporter plasmid MITF-Luc was transiently co-transfected into 293T cells in combination with the LEF-1 expression vector and increasing quantities of the MITF expression vector. d, e The transactivation of the MITF promoter by LEF-1 and WT/mutant MITF. In all, 293T cells (c) and UACC903 cells (d) were transiently co-transfected with the MITF-Luc reporter plasmid, LEF-1, and WT/mutant MITF. For all assays, relative luciferase activity is shown as the ratio of each normalized luciferase activity to the value obtained with each reporter plasmid and empty vector. The data shown are the means ± S.D. of three independent experiments. (*p < 0.05 ***p < 0.001 by one-way ANOVA with Dunnett’s multiple comparison tests compared with the basal activity; for a and b, ##p < 0.01; ###p < 0.001; ns: not significant; compared with the value from shown promoter together with LEF-1 and MITF by unpaired Student’s t-test; For c–e, ##p < 0.01; ###p < 0.001; ns: not significant; compared with the value from shown promoter together with LEF-1 by unpaired Student’s t-test)
Effects of mutant MITF on the interaction between LEF-1 and MITF
To investigate whether WS-associated mutations affect the interaction between LEF-1 and MITF, we performed a series of co-immunoprecipitation studies. As shown in Fig. 4, WT/mutant MITF and LEF-1 all co-immunoprecipitate when co-expressed in 293T cells. The mutant MITF did not affect the interaction between LEF-1 and wild-type MITF.
Interaction between LEF-1 and WT/mutant MITF. LEF-1 was tagged with HA, and WT MITF and mutants R255X, R217I, R217G, and R217del were tagged with FLAG. In all, 293T cells were transfected with LEF-1 and WT/mutant MITF expression plasmids as is shown in the figure. Cell extracts were subjected to immunoprecipitation with anti-Flag M2 antibody or with normal mouse IgG (NIG) as a negative control. Immune complexes were resolved by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and immunoblotted with anti-HA rabbit polyclonal antibody. Input was analyzed using monoclonal anti-HA antibody as indicated
Effects of mutant MITF on binding of LEF-1 at the MITF promoter
To determine whether the WT/mutant MITF would affect the binding of LEF-1 on the MITF promoter, we performed Biotinylated DNA affinity precipitation As shown in Fig. 5, WT and R217I mutant MITF disrupt the binding of LEF-1 on the MITF promoter, but R255X, R217G, and the R217del mutants do not disrupt the binding of LEF-1 on the MITF promoter.
Effects of mutant MITF on the DNA-binding capacity of LEF-1 on the MITF promoter. The lysates of 293T cells transfected with LEF-1 and WT/mutant MITF expression plasmids were incubated with or without biotinylated double-stranded oligonucleotides containing the LEF-1-binding region in the MITF promoter. The DNA/protein complex was pulled down with streptavidin-agarose beads. The precipitated proteins were separated on sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and analyzed by immunoblotting using anti-HA rabbit polyclonal antibodies
Discussion
WS is a melanin-associated disease that is clinically and genetically heterogeneous. We report four different WS2-associated MITF mutations, all found in China, that present highly variable clinical features. The mutant R217I is characterized by the most severe phenotype, and R217del is the next most severe. The phenotypes of R255X and R217G are milder.
As is shown in Fig. 1, MITF is a target of Wnt signaling; Wnt signaling upregulates MITF expression through functional LEF-1-binding sites on the M promoter [25]. MITF also functions as a nuclear mediator to interact with the c-terminal portion of LEF-1 through b-HLH-Zip domains to regulate its own expression through promoter activation (as a non-DNA-binding cofactor) [25] and to regulate DCT and TYR gene expression (as a DNA-binding coactivator) [23, 27]. MITF controls the synthesis of melanin by regulating the expression of TYR, DCT, and its own gene; MITF alone can activate the TYR gene promoter, but it must interact with LEF-1 to modulate its own gene promoter and the DCT gene promoter. We have shown that MITF alone or LEF-1 alone minimally activate the DCT promoter, but when they are co-expressed in 293T cells, the DCT promoter activity is dramatically increased, consistent with previous reports [23]. The mutant MITF destroyed the synergistic transcriptional activation of LEF-1 and MITF on the DCT promoter in two types of cells. However, the synergistic transactivation of the DCT promoter by MITF and LEF-1 was not observed in UACC903 cells, perhaps because they express high levels of LEF-1 (Fig. 3b). LEF-1 alone can activate the expression of MITF, but MITF proteins attenuate the LEF-1-mediated activation of the MITF-M promoter; this inhibition weakens when the dosage of MITF is reduced (Fig. 3c). This result was contradicting with previous literature [25] and it may be because of different cell type and need to be futher verified in vivo.
The MITF-M promoter contains three clustered LEF-1-binding sites. Functional interaction between LEF-1 and MITF-M on the MITF promoter depends on the binding of LEF-1 to the clustered LEF-1-binding sites in the nucleus [25]. None of the mutants affect the interaction between LEF-1 and wild-type MITF (Fig. 4). In our case, the mutant R217I localized in the nucleus similarly as wild-type MITF [3]. Therefore, R217I retained a partial ability to activate TYR (Fig. 2a, b) [12], but it rendered LEF-1 unable to bind to the MITF promoter (Fig. 5) and inhibited the activation of the MITF-M promoter by LEF-1 (Fig. 3d, e). The haploinsufficiency caused by R217I was thus further aggravated and associated with the most severe phenotype, even presenting as Tietz syndrome [11].
The phenotypic heterogeneity by MITF gene mutation was associated with DNA-binding activity and transcription activation [12]. The R217del, the R255X and the R217G all fail to bind DNA and activate expression from the melanocyte-specific promoters but show different phenotype [12]. The mutant R217del does not localize to the nucleus and remains cytoplasmic [36]; it does not affect LEF-1-binding to the MITF promoter (Fig. 5), and thus it cannot inhibit the LEF-1-mediated activation of the MITF-M promoter (Fig. 3d, e). However, it has a more severe phenotype and can even lead to Tietz syndrome, possibly because of its dominant negative effect [3, 37, 38].
R255X localizes in the cytoplasm and nucleus [39], and R217G localizes in the nucleus [29]. These mutants allow LEF-1 to retain its binding to the MITF promoter (Fig. 5) and do not inhibit the activation of the MITF-M promoter by LEF-1 (Fig. 3d, e). Thus, the haploinsufficiency caused by R255X and R217G can be overcome; they cannot activate the target gene TYR completely, but they have a milder phenotype.
The data imply that MITF may have a negative feedback loop of regulation involving the Wnt signaling pathway to stabilize MITF gene dosage. The haploinsufficiency of the mutant MITF can be overcome through the regulation of Wnt signaling pathway. The activation of target gene TYR and the synthesis of melain will be improved and the phenotype of WS2 will become milder. Our study shows that the Wnt signaling pathway is involved in the genotypic and phenotypic variations seen in Waardenburg syndrome type 2 caused by mutations of MITF in vitro and the haploinsufficiency is the main pathogenic mechanism of WS2. Our results provide a new molecular insights into how MITF mutations can lead to different phenotypes of WS2 through Wnt/β-catenin signaling pathway. Further in vivo studies need to be performed to justify our results.
References
Touraine RL, Attie-Bitach T, Manceau E, Korsch E, Sarda P, Pingault V, et al. Neurological phenotype in Waardenburg syndrome type 4 correlates with novel SOX10 truncating mutations and expression in developing brain. Am J Hum Genet. 2000;66:1496–503.
Jiang L, Chen H, Jiang W, Hu Z, Mei L, Xue J, et al. Novel mutations in the SOX10 gene in the first two Chinese cases of type IV Waardenburg syndrome. Biochem Biophys Res Commun. 2011;408:620–4.
Amiel J, Watkin PM, Tassabehji M, Read AP, Winter RM. Mutation of the MITF gene in albinism-deafness syndrome (Tietz syndrome). Clin Dysmorphol. 1998;7:17–20.
Pingault V, Ente D, Dastot-Le Moal F, Goossens M, Marlin S, Bondurand N. Review and update of mutations causing Waardenburg syndrome. Hum Mutat. 2010;31:391–406.
Ogawa Y, Kono M, Akiyama M. Pigmented macules in Waardenburg syndrome type 2 due to KITLG mutation. Pigment Cell Melanoma Res. 2017;30:501–4.
Hou L, Pavan WJ. Transcriptional and signaling regulation in neural crest stem cell-derived melanocyte development: do all roads lead to Mitf? Cell Res. 2008;18:1163–76.
Waardenburg PJ. A new syndrome combining developmental anomalies of the eyelids, eyebrows and nose root with pigmentary defects of the iris and head hair and with congenital deafness. Am J Hum Genet. 1951;3:195–253.
Read AP, Newton VE. Waardenburg syndrome. J Med Genet. 1997;34:656–65.
Tassabehji M, Newton VE, Read AP. Waardenburg syndrome type 2 caused by mutations in the human microphthalmia (MITF) gene. Nat Genet. 1994;8:251–5.
Tietz W. A syndrome of deaf-mutism associated with albinism showing dominant autosomal inheritance. Am J Hum Genet. 1963;15:259–64.
Smith SD, Kelley PM, Kenyon JB, Hoover D. Tietz syndrome (hypopigmentation/deafness) caused by mutation of MITF. J Med Genet. 2000;37:446–8.
Grill C, Bergsteinsdottir K, Ogmundsdottir MH, Pogenberg V, Schepsky A, Wilmanns M, et al. MITF mutations associated with pigment deficiency syndromes and melanoma have different effects on protein function. Hum Mol Genet. 2013;22:4357–67.
Hershey CL, Fisher DE. Genomic analysis of the Microphthalmia locus and identification of the MITF-J/Mitf-J isoform. Gene. 2005;347:73–82.
Meadows NA, Sharma SM, Faulkner GJ, Ostrowski MC, Hume DA, Cassady AI. The expression of Clcn7 and Ostm1 in osteoclasts is coregulated by microphthalmia transcription factor. J Biol Chem. 2007;282:1891–904.
Hughes MJ, Lingrel JB, Krakowsky JM, Anderson KP. A helix-loop-helix transcription factor-like gene is located at the mi locus. J Biol Chem. 1993;268:20687–90.
Hodgkinson CA, Moore KJ, Nakayama A, Steingrimsson E, Copeland NG, Jenkins NA, et al. Mutations at the mouse microphthalmia locus are associated with defects in a gene encoding a novel basic-helix-loop-helix-zipper protein. Cell. 1993;74:395–404.
Tachibana M, Perez-Jurado LA, Nakayama A, Hodgkinson CA, Li X, Schneider M, et al. Cloning of MITF, the human homolog of the mouse microphthalmia gene and assignment to chromosome 3p14.1-p12.3. Hum Mol Genet. 1994;3:553–7.
Yang SH, Han JS, Baek SH, Kwak EY, Kim HJ, Shin JH, et al. Construction of protein chip to detect binding of Mitf protein (microphthalmia transcription factor) and E-box DNA. Appl Biochem Biotechnol. 2008;151:273–82.
Goding CR. Mitf from neural crest to melanoma: signal transduction and transcription in the melanocyte lineage. Genes Dev. 2000;14:1712–28.
Nobukuni Y, Watanabe A, Takeda K, Skarka H, Tachibana M. Analyses of loss-of-function mutations of the MITF gene suggest that haploinsufficiency is a cause of Waardenburg syndrome type 2A. Am J Hum Genet. 1996;59:76–83.
Hemesath TJ, Steingrimsson E, McGill G, Hansen MJ, Vaught J, Hodgkinson CA, et al. Microphthalmia, a critical factor in melanocyte development, defines a discrete transcription factor family. Genes Dev. 1994;8:2770–80.
Zhang H, Luo H, Chen H, Mei L, He C, Jiang L, et al. Functional analysis of MITF gene mutations associated with Waardenburg syndrome type 2. FEBS Lett. 2012;586:4126–31.
Yasumoto K, Takeda K, Saito H, Watanabe K, Takahashi K, Shibahara S. Microphthalmia-associated transcription factor interacts with LEF-1, a mediator of Wnt signaling. EMBO J. 2002;21:2703–14.
Saito H, Yasumoto K, Takeda K, Takahashi K, Yamamoto H, Shibahara S. Microphthalmia-associated transcription factor in the Wnt signaling pathway. Pigment Cell Res. 2003;16:261–5.
Saito H, Yasumoto K, Takeda K, Takahashi K, Fukuzaki A, Orikasa S, et al. Melanocyte-specific microphthalmia-associated transcription factor isoform activates its own gene promoter through physical interaction with lymphoid-enhancing factor 1. J Biol Chem. 2002;277:28787–94.
Schepsky A, Bruser K, Gunnarsson GJ, Goodall J, Hallsson JH, Goding CR, et al. The microphthalmia-associated transcription factor Mitf interacts with beta-catenin to determine target gene expression. Mol Cell Biol. 2006;26:8914–27.
Wang X, Liu Y, Chen H, Mei L, He C, Jiang L, et al. LEF-1 regulates tyrosinase gene transcription in vitro. PLoS ONE. 2015;10:e0143142.
Chen H, Jiang L, Xie Z, Mei L, He C, Hu Z, et al. Novel mutations of PAX3, MITF, and SOX10 genes in Chinese patients with type I or type II Waardenburg syndrome. Biochem Biophys Res Commun. 2010;397:70–4.
Yang S, Dai P, Liu X, Kang D, Zhang X, Yang W, et al. Genetic and phenotypic heterogeneity in Chinese patients with Waardenburg syndrome type II. PLoS ONE. 2013;8:e77149.
Vachtenheim J, Novotna H, Ghanem G. Transcriptional repression of the microphthalmia gene in melanoma cells correlates with the unresponsiveness of target genes to ectopic microphthalmia-associated transcription factor. J Invest Dermatol. 2001;117:1505–11.
Bondurand N, Pingault V, Goerich DE, Lemort N, Sock E, Le Caignec C, et al. Interaction among SOX10, PAX3 and MITF, three genes altered in Waardenburg syndrome. Hum Mol Genet. 2000;9:1907–17.
Zhang H, Chen H, Luo H, An J, Sun L, Mei L, et al. Functional analysis of Waardenburg syndrome-associated PAX3 and SOX10 mutations: report of a dominant-negative SOX10 mutation in Waardenburg syndrome type II. Hum Genet. 2012;131:491–503.
Bentley NJ, Eisen T, Goding CR. Melanocyte-specific expression of the human tyrosinase promoter: activation by the microphthalmia gene product and role of the initiator. Mol Cell Biol. 1994;14:7996–8006.
Yasumoto K, Yokoyama K, Takahashi K, Tomita Y, Shibahara S. Functional analysis of microphthalmia-associated transcription factor in pigment cell-specific transcription of the human tyrosinase family genes. J Biol Chem. 1997;272:503–9.
Bertolotto C, Busca R, Abbe P, Bille K, Aberdam E, Ortonne JP, et al. Different cis-acting elements are involved in the regulation of TRP1 and TRP2 promoter activities by cyclic AMP: pivotal role of M boxes (GTCATGTGCT) and of microphthalmia. Mol Cell Biol. 1998;18:694–702.
Takebayashi K, Chida K, Tsukamoto I, Morii E, Munakata H, Arnheiter H, et al. The recessive phenotype displayed by a dominant negative microphthalmia-associated transcription factor mutant is a result of impaired nucleation potential. Mol Cell Biol. 1996;16:1203–11.
Shigemura T, Shiohara M, Tanaka M, Takeuchi K, Koike K. Effect of the mutant microphthalmia-associated transcription factor found in Tietz syndrome on the in vitro development of mast cells. J Pediatr Hematol/Oncol. 2010;32:442–7.
Izumi K, Kohta T, Kimura Y, Ishida S, Takahashi T, Ishiko A, et al. Tietz syndrome: unique phenotype specific to mutations of MITF nuclear localization signal. Clin Genet. 2008;74:93–5.
Chen H, Liao X, Liu Y, He C, Zhang H, Jiang L, et al. [Study of gene mutation and pathogenetic mechanism for a family with Waardenburg syndrome]. Zhonghua yi xue yi chuan xue za zhi . 2017;34:471–5.
Acknowledgements
We would like to thank all of the technical staff from the State Key Laboratory of Medical Genetics and Province Key Laboratory of Otolaryngology Critical Disease for their support. Our work was supported by the National Basic Research Program of China (2014CB541702), by grants from National Natural Science Foundation of China (No. 81470705 and No. 81771023) and the Youth Foundation of the First Affiliated Hospital of Zhengzhou University (No.YNQN2017001).
Author contributions
Project administration: X-PW, Z-JN, JS, L-YM, C-FH, Y-LL; resources: YF, HZ; supervision: YF, Y-LZ, HZ; validation: X-PW; visualization: X-PW; writing (original draft preparation): X-PW; writing (review and editing): X-PW
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Wang, XP., Liu, YL., Mei, LY. et al. Wnt signaling pathway involvement in genotypic and phenotypic variations in Waardenburg syndrome type 2 with MITF mutations. J Hum Genet 63, 639–646 (2018). https://doi.org/10.1038/s10038-018-0425-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-018-0425-z
This article is cited by
-
Identification of two variants in PAX3 and FBN1 in a Chinese family with Waardenburg and Marfan syndrome via whole exome sequencing
Functional & Integrative Genomics (2023)
-
PPP6C, a serine-threonine phosphatase, regulates melanocyte differentiation and contributes to melanoma tumorigenesis through modulation of MITF activity
Scientific Reports (2022)