Abstract
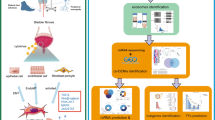
Interstitial cystitis (IC), also known as bladder pain syndrome, is a chronic inflammatory disease that affects the bladder. The symptoms of IC vary, including feeling an urgent need for immediate urination and of needing to urinate often, as well as bladder or pelvic pain. Despite its high incidence, no molecular diagnostic methods are available for IC, and the molecular pathogenesis is unknown. microRNAs (miRNA) can regulate expression of RNA transcripts in cells and aberrant expression of miRNAs is associated with several human diseases. Here, we investigated the molecular pathogenesis of IC based on miRNA expression signatures. RNA sequencing of miRNA levels in IC tissues and comparison with levels in normal bladder tissue and bladder cancer revealed dysregulated expression of 366 miRNAs (203 and 163 down- and upregulated miRNAs, respectively). In particular, miR-320 family miRNAs(miR-320a, miR-320b, miR-320c, miR-320d and miR-320e) had downregulated expression in IC tissues. Genome-wide gene expression analyses and in silico database analyses showed that three transcription factors, E2F-1, E2F-2 and TUB, are regulated by miR-320 family miRNAs. Immunostaining of IC tissues confirmed that these transcription factors are overexpressed in IC tissues. Novel approaches that identify aberrantly expressed miRNA regulatory networks in IC could provide new prognostic markers and therapeutic targets for this disease.
Similar content being viewed by others
Introduction
Interstitial cystitis (IC) is a non-specific chronic inflammatory bladder disease characterized by urinary frequency, urinary urgency, and pelvic pain induced by bladder filling [1, 2]. Patients with IC suffer significant declines in quality of life (QOL). The regional morbidity rate of IC varies, with 3–4 per 10 million in Japan, 60–70 per 10 million in the United States, and 18 per 10 million in Europe [3]. Based on cystoscopic findings, IC can be classified into 2 groups: Hunner type with ulcerative lesions (Hunner’s lesion), and non-Hunner type with bladder petechial oozing (glomerulation) after hydrodistension. Diagnosis and classification of IC is determined by cystoscopic findings and exclusion of other diseases based on subjective symptoms. Although several studies searching for diagnostic markers IC have been conducted, no definitive markers are known [4]. Moreover, no symptomatic treatments for IC such as bladder hydrodistension, drug therapy and food therapy, or curative therapy have been established [5]. Given the lack of consensus about IC etiology, a genomic approach to elucidate molecular mechanisms underlying IC pathogenesis is needed.
microRNA (miRNA) are small (19 to 22 nucleotide) non-coding RNAs that can finely control expression of protein-coding or non-coding RNAs [6]. A single miRNA can control sequence-dependent expression of an extremely large number of RNA transcripts [7]. Therefore, aberrantly expressed miRNAs can lead to dysregulation of intracellular RNA networks. Indeed, a large number of studies have shown aberrant miRNA expression in several diseases and cancers, thus highlighting the important role for miRNAs in human disease pathogenesis [8,9,10,11,12,13].
We have successfully identified several cancer networks that are regulated by antitumor miRNAs in various cancer types using genome-wide gene expression analyses and in silico database searches [14,15,16,17,18,19]. Here we adapted our miRNA analysis strategy to characterize the molecular pathogenesis of IC. First, we analyzed the miRNA expression signature of IC based on RNA sequencing of samples from clinical specimens of Hunner-type IC and non-Hunner type IC. We then examined dysregulated miRNA expression in IC tissues to identify novel molecular networks involved in IC pathogenesis.
Analyses of miRNA signature revealed that expression of 366 miRNAs (203 and 163 downregulated and upregulated, respectively) was dysregulated in IC tissues. Moreover, we identified several genes that are targeted by the dysregulated miRNAs in IC tissues. Novel approaches that identify aberrantly expressed miRNA regulatory networks in IC could provide new prognostic markers and therapeutic targets for this disease.
Materials and methods
Patients and IC clinical specimens
IC clinical specimens were obtained from patients who were diagnosed at Dokkyo Medical University Hospital and Chiba University Hospital. Written informed consent was obtained from each patient in advance of sample collection. All patients underwent bladder hydrodistension, and IC tissues were collected by bladder biopsy. Hunner type IC patients were diagnosed by the presence of lesions, whereas non-Hunner type IC was classified based on the presence of glomerulation during bladder hydrodistension. Patient characteristics and cystoscopic findings are summarized in Supplementary Table 1 and Fig. 1, respectively.
Cystoscopic findings of IC patients. Cystoscopic findings of IC patients used in deep sequencing analysis. No.1, 2, 6, 7 and 8 are Hunner type IC patients’ findings, indicating Hunner’s lesions. No. 3, 4 and 5 are non-Hunner type IC patients’ findings, showing bladder petechial oozing (glomerulation) after hydrodistension
RNA extraction and cell culture
We extracted total RNA using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol, and RNA quality was assessed as previously described [15, 17, 20, 21]. The human bladder cancer (BC) cell line T24 was obtained from the American Type Culture Collection (ATCC; Manassas, VA, USA) and maintained as previously described [15, 17, 20, 21].
Construction of miRNA expression signature for IC
To create a miRNA expression signature for IC, we performed small RNA sequencing using a HiSeq 2000 (Illumina, San Diego, CA, USA) for 8 IC samples (Supplementary Table 1). Samples from 5 normal and 5 cancer tissues of bladder that we described in a previous study were used as comparative specimens [17]. We carried out small RNA sequencing and data mining procedures as described previously [16, 17, 19, 22]. A false discovery rate (FDR) of less than 0.05 was considered to be significant.
Transfection with mature miRNA
We used Ambion Pre-miR miRNA precursor for hsa-miR-320b (assay ID: PM13132; Applied Biosystems, Foster City, CA, USA) and hsa-miR-320c (assay ID: PM13133; Applied Biosystems)as mature miRNA species. miRNA transfection of T24 cells and analysis of transfection efficiencies were conducted as described previously [15, 17, 20, 21].
Identification of putative genes regulated by the miR-320 family in T24 cells
To identify genes regulated by the miR-320-family, we combined in silico and genome-wide gene expression analyses as previously described [15, 17, 20, 21]. We used the TargetScanHuman 7.1 (June, 2016 release, http://www.targetscan.org/vert_71) database and an oligo microarray (Agilent Technologies; Human Ge 60 K) for gene expression analyses. Microarray data were deposited into the GEO database (https://www.ncbi.nlm.nih.gov/geo/).
Immunohistochemistry
We incubated tissue specimens overnight at 4 °C with anti-E2F-1 antibodies (1:50 dilution; Cat. no. sc-251; Santa Cruz Biotechnology, Santa Cruz, CA, USA), anti-E2F-2 antibodies (1:50; Cat. no. sc-9967; Santa Cruz Biotechnology) and anti-TUB antibodies (1:50; product no. HPA049019; Sigma-Aldrich, St. Louis, MO, USA). The procedures were conducted as described previously [15, 17, 20, 21].
Results
Small RNA sequences of specimens from normal bladder, bladder cancer and IC specimens
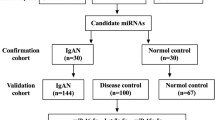
We performed deep sequencing of 18 small RNA libraries from specimens from 5 normal bladders, 5 bladder cancer cases, and 8 IC cases. Patient backgrounds for the specimens from normal and bladder cancer cases are described in our previous report [17], and those for IC specimens are summarized in Supplementary Table 1. BC and normal tissues were reanalyzed for this study. We obtained between 13,554,054 and 22,747,525 total reads and 4,837,296–19,218,202 mapped reads (Table 1). We focused on previously annotated miRNAs and detected 745–1,417 miRNAs for each sample. From comparisons of normal bladder samples, BC samples, and IC samples, we constructed miRNA expression signatures that included miRNAs exhibiting significantly up- or downregulated expression (Tables 2–5; Supplementary Tables 2–7; (|log2FC| > 1 and FDR < 0.05).
To extract miRNAs that had marked downregulation in IC tissues, we downselected candidate miRNAs according to the strategy shown in Supplementary Figure 1. Among the 203 miRNAs that were significantly downregulated in IC tissues compared to normal tissues, we excluded miRNAs that were also markedly upregulated or downregulated in comparisons of normal tissues with BC tissues. Of the 151 remaining miRNAs, we focused on the miR-320 family (miR-320a/b/c/d/e), which is composed of 5 miRNAs (miR-320a/b/c/d/e) having the same seed sequence, given the high possibility that these miRNAs will target similar genes (Supplementary Figure 2). In our profile, expression of all miR-320 family members was markedly downregulated in IC tissues.
Identification of molecular targets regulated by the miR-320 family
We hypothesized that genes regulated by miRNAs that have decreased expression in IC tissues could contribute to IC pathogenesis. Using the TargetScanHuman 7.1 database for initial analyses, we found 5158 genes that had candidate target sites for the miR-320 family. Next, genome-wide gene expression analyses were performed to narrow down candidate genes. This time, by transfection with miR-320b and miR-320c, which were mostly downregulated in IC, genes whose expression decreased in T24 cells were extracted. The microarray data were deposited into the GEO database (GEO accession number: GSE106791). These analyses identified 162 and 222 genes as candidates for regulation by miR-320b and miR-320c, respectively. Finally, we integrated these results, 85 genes remained as candidate genes for regulation by the miR-320 family (Table 6). The strategy for selection of target genes is shown in Supplementary Figure 3. Among putative targets of miR-320 family, several genes, e.g., E2F1, RAB23, ITGB3, and TERT, have been reported to be associated with bladder cancer [23,24,25,26]. However, as far as the PubMed database was searched, no reports were found about the relationship between these genes and IC. Additionally, we assigned genes to KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways using the GENECODIS program and identified 6 significantly enriched pathways (Supplementary Table 8).
E2F-1, E2F-2 and TUB expression in IC clinical specimens
We focused on 3 transcription factors (E2F-1, E2F-2 and TUB) and further analyses were performed. Previous studies showed that dysregulated expression or mutation of several transcription factors were deeply involved in human diseases and syndromes [27]. Function of E2F family is the transcriptional activation or repression of its target genes and pivotal role of cell proliferation, differentiation and apoptosis [28, 29]. The involvement of these genes and IC is obscure. We performed immunohistochemistry to assess protein expression of E2F-1, E2F-2, and TUB in IC clinical specimens. All three proteins were strongly expressed in IC urothelial cells compared with normal bladder and BC tissues (Fig. 2).
Discussion
Recently developed RNA sequencing technology is suitable for compiling miRNA expression signatures. We previously described several miRNA expression signatures of human cancers based on RNA sequencing and identified novel RNA networks regulated by antitumor miRNAs [14,15,16,17,18,19]. In this study, we presented the miRNA signature of IC determined by RNA sequencing. Identification of dysregulated miRNAs in IC tissues compared with normal bladder tissues or bladder cancer tissues will be useful for exploring molecular pathogenesis in IC cells.
Among the downregulated miRNAs in IC tissues, we focused on the miR-320 family because all members of this family showed downregulated expression of miR-320 in IC tissues compared with normal bladder tissues. In contrast to our study, a miRNA expression profile of IC generated by PCR-based microarray analysis in a previous study showed that several miRNAs (miR-449b, miR-500, miR-328 and miR-320) were upregulated in IC tissues [30]. As a factor that causes different results from our research, analysis methods and differences in patient background may be considered. This report also showed that IC patients had notably decreased expression of neurokinin-1 (tachykinin receptors) that was presumed to be due to upregulated miR-320 expression in IC tissues [30]. Downregulation of the miR-320 family and the antitumor activity of these miRNAs have been reported for several cancers [31,32,33,34,35]. In bladder cancer, previous studies showed that miR-320a and miR-320c acted as antitumor miRNAs by targeting integrin beta-3 (ITGB3) and cyclin-dependent kinase 6 (CDK6), respectively [25, 36].
To identify possible genes involved in IC pathogenesis, we searched molecular pathways containing genes targeted by the miR-320 family using genome-wide gene expression analyses and in silico database analyses. Using these approaches, we identified 85 genes as putative targets of miR-320 family regulation. Among these genes, we focused on transcription factors due to their potential to significantly affect downstream RNA networks. Several studies indicated that dysregulation of transcription factor activity is associated with various human diseases [37, 38]. More recently, the urothelial master transcription factors TP63, SHH and FOXA1 were reported to act as novel diagnostic markers and to have potential involvement in the molecular pathology of IC [39]. In this study, we focused on 3 transcription factors: E2F transcription factor 1 (E2F-1), E2F transcription factor 2 (E2F-2) and tubby bipartite transcription factor (TUB) and confirmed upregulated protein expression of all three in IC lesions.
The E2F family of transcription factors is involved in cell cycle regulation and synthesis of DNA in higher eukaryotes, and plays a pivotal role during the G1/S transition in mammalian cells [28, 29]. E2F-1, E2F-2, and E2F-3a function as transcription activators, whereas E2F-3b, E2F-4, E2F-5, E2F-6, E2F-7, and E2F-8 act as transcription suppressors [28]. E2Fs are associated with cancer and are major target genes for the Rb tumor suppressor protein (pRb). Inactivation of pRb causes overexpression of E2Fs that can act as a cancer promoter [40]. Another report described the involvement of E2F-1 and E2F-2 overexpression in inflammation associated with traumatic spinal cord injury (SCI) in a mouse model. Results from that report suggested that SCI-induced activation of E2F-1 and E2F-2 expression may result in movement disorder and hypersensitivity after trauma. Furthermore, E2F-1 and E2F-2 knockout significantly reduced the amount of neuron death, neuroinflammation and tissue damage in the SCI mouse model, together with an enhanced greater neuroprotective effect [41]. Based on these earlier findings, overexpression of E2F-1 and E2F-2 could promote inflammation associated with IC.
The tubby protein encoded by the TUB gene is a common upstream cell signalling protein in multicellular eukaryotes. The tubby protein acts as a signalling factor that potentially couples with G protein activity [42, 43]. Moreover, tubby proteins are involved in neuronal differentiation and development, whereas mutations in tubby genes in mammals are associated with delayed obesity, sensorineural hearing loss and retinal degeneration [43,44,45]. However, the functions of tubby proteins in humans are unclear and there is no report that they are involved in inflammatory diseases.
The expression changes in these three transcription factors and the networks induced by these changes may indicate that they play a significant role in IC onset and progression.
In conclusion, in this study we determined the miRNA expression signature in IC by RNA sequencing and successfully identified dysregulated expression of miRNAs. We found that all members of the miR-320 family were downregulated in IC tissues and that E2F-1, E2F-2, and TUB were putative targets of regulation by these miRNAs. Immunohistochemistry showing increased levels of E2F-1, E2F-2, and TUB proteins in IC lesions support the sequencing results. In order to accurately understand the RNA network within the cell, it is ideal to use cells derived from the disease. From the latest genome technology, it is important that IC derived cell lines can be established and analyzed for its RNA networks. The miRNA signature of IC generated by RNA sequencing and in silico analyses could provide a basis to develop novel therapeutic targets for IC.
References
van de Merwe JP, Nordling J, Bouchelouche P, Bouchelouche K, Cervigni M, Daha LK, et al. Diagnostic criteria, classification, and nomenclature for painful bladder syndrome/interstitial cystitis: an ESSIC proposal. Eur Urol. 2008;53:60–7.
Macdiarmid SA, Sand PK. Diagnosis of interstitial cystitis/ painful bladder syndrome in patients with overactive bladder symptoms. Rev Urol. 2007;9:9–16.
Dawson, TE & Jamison, J Intravesical treatments for painful bladder syndrome/ interstitial cystitis. Cochrane Database Syst Rev. 2007;4:Cd006113.
Kim HJ. Update on the pathology and diagnosis of interstitial cystitis/bladder pain syndrome: a review. Int Neurourol J. 2016;20:13–7.
Chancellor MB, Yoshimura N. Treatment of interstitial cystitis. Urology. 2004;63:85–92.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell . 2004;116:281–97.
Friedman RC, Farh KK, Burge CB, Bartel DP. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009;19:92–105.
Wiemer EA. The role of microRNAs in cancer: no small matter. Eur J Cancer. 2007;43:1529–44.
Nelson KM, Weiss GJ. MicroRNAs and cancer: past, present, and potential future. Mol Cancer Ther. 2008;7:3655–60.
Rebane A, Akdis CA. MicroRNAs: essential players in the regulation of inflammation. J Allergy Clin Immunol. 2013;132:15–26.
Ha TY. MicroRNAs in human diseases: from cancer to cardiovascular disease. Immune Netw. 2011;11:135–54.
Qu Z, Li W, Fu B. MicroRNAs in autoimmune diseases. Biomed Res Int. 2014;2014:527895.
Dissanayake E, Inoue Y. MicroRNAs in allergic disease. Curr Allergy Asthma Rep. 2016;16:67.
Fuse M, Kojima S, Enokida H, Chiyomaru T, Yoshino H, Nohata N, et al. Tumor suppressive microRNAs (miR-222 and miR-31) regulate molecular pathways based on microRNA expression signature in prostate cancer. J Hum Genet. 2012;57:691–9.
Goto Y, Kojima S, Nishikawa R, Kurozumi A, Kato M, Enokida H, et al. MicroRNA expression signature of castration-resistant prostate cancer: the microRNA-221/222 cluster functions as a tumour suppressor and disease progression marker. Br J Cancer. 2015;113:1055–65.
Goto Y, Kurozumi A, Arai T, Nohata N, Kojima S, Okato A, et al. Impact of novel miR-145-3p regulatory networks on survival in patients with castration-resistant prostate cancer. Br J Cancer. 2017;117:409–20.
Itesako T, Seki N, Yoshino H, Chiyomaru T, Yamasaki T, Hidaka H, et al. The microRNA expression signature of bladder cancer by deep sequencing: the functional significance of the miR-195/497 cluster. PLoS ONE. 2014;9:e84311.
Goto Y, Kurozumi A, Nohata N, Kojima S, Matsushita R, Yoshino H, et al. The microRNA signature of patients with sunitinib failure: regulation of UHRF1 pathways by microRNA-101 in renal cell carcinoma. Oncotarget . 2016;7:59070–86.
Koshizuka K, Nohata N, Hanazawa T, Kikkawa N, Arai T, Okato A, et al. Deep sequencing-based microRNA expression signatures in head and neck squamous cell carcinoma: dual strands of pre-miR-150 as antitumor miRNAs. Oncotarget . 2017;8:30288–304.
Arai T, Okato A, Kojima S, Idichi T, Koshizuka K, Kurozumi A, et al. Regulation of spindle and kinetochore-associated protein 1 by antitumor miR-10a-5p in renal cell carcinoma. Cancer Sci. 2017;108:2088–101.
Matsushita R, Seki N, Chiyomaru T, Inoguchi S, Ishihara T, Goto Y, et al. Tumour-suppressive microRNA-144-5p directly targets CCNE1/2 as potential prognostic markers in bladder cancer. Br J Cancer. 2015;113:282–9.
Yonemori K, Seki N, Idichi T, Kurahara H, Osako Y, Koshizuka K, et al. The microRNA expression signature of pancreatic ductal adenocarcinoma by RNA sequencing: anti-tumour functions of the microRNA-216 cluster. Oncotarget . 2017;8:70097–115.
Jin H, Xie Q, Guo X, Xu J, Wang A, Li J, et al. p63alpha protein up-regulates heat shock protein 70 expression via E2F1 transcription factor 1, promoting Wasf3/Wave3/MMP9 signaling and bladder cancer invasion. J Biol Chem. 2017;292:15952–63.
Jiang Y, Han Y, Sun C, Han C, Han N, Zhi W, et al. Rab23 is overexpressed in human bladder cancer and promotes cancer cell proliferation and invasion. Tumour Biol. 2016;37:8131–8.
Shang C, Zhang H, Guo Y, Hong Y, Liu Y, Xue Y. MiR-320a down-regulation mediates bladder carcinoma invasion by targeting ITGB3. Mol Biol Rep. 2014;41:2521–7.
Borah S, Xi L, Zaug AJ, Powell NM, Dancik GM, Cohen SB, et al. Cancer. TERT promoter mutations and telomerase reactivation in urothelial cancer. Science. 2015;347:1006–10.
Lee TI, Young RA. Transcriptional regulation and its misregulation in disease. Cell . 2013;152:1237–51.
Attwooll C, Lazzerini Denchi E, Helin K. The E2F family: specific functions and overlapping interests. EMBO J. 2004;23:4709–16.
Gaubatz S, Lindeman GJ, Ishida S, Jakoi L, Nevins JR, Livingston DM, et al. E2F4 and E2F5 play an essential role in pocket protein-mediated G1 control. Mol Cell. 2000;6:729–35.
Sanchez Freire V, Burkhard FC, Kessler TM, Kuhn A, Draeger A, Monastyrskaya K. MicroRNAs may mediate the down-regulation of neurokinin-1 receptor in chronic bladder pain syndrome. Am J Pathol. 2010;176:288–303.
Wu YY, Chen YL, Jao YC, Hsieh IS, Chang KC, Hong TM. miR-320 regulates tumor angiogenesis driven by vascular endothelial cells in oral cancer by silencing neuropilin 1. Angiogenesis. 2014;17:247–60.
Vishnubalaji R, Hamam R, Yue S, Al-Obeed O, Kassem M, Liu FF, et al. MicroRNA-320 suppresses colorectal cancer by targeting SOX4, FOXM1, and FOXQ1. Oncotarget. 2016;7:35789–802.
Shi C, Zhang Z. MicroRNA-320 suppresses cervical cancer cell viability, migration and invasion via directly targeting FOXM1. Oncol Lett. 2017;14:3809–16.
Pan C, Gao H, Zheng N, Gao Q, Si Y, Zhao Y. MiR-320 inhibits the growth of glioma cells through downregulating PBX3. Biol Res. 2017;50:31.
Okato A, Goto Y, Kurozumi A, Kato M, Kojima S, Matsushita R, et al. Direct regulation of LAMP1 by tumor-suppressive microRNA-320a in prostate cancer. Int J Oncol. 2016;49:111–22.
Wang X, Wu J, Lin Y, Zhu Y, Xu X, Xu X, et al. MicroRNA-320c inhibits tumorous behaviors of bladder cancer by targeting Cyclin-dependent kinase 6. J Exp Clin Cancer Res. 2014;33:69.
Whyte WA, Orlando DA, Hnisz D, Abraham BJ, Lin CY, Kagey MH, et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell. 2013;153:307–19.
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-Andre V, Sigova AA, et al. Super-enhancers in the control of cell identity and disease. Cell . 2013;155:934–47.
Kaga K, Inoue KI, Kaga M, Ichikawa T, Yamanishi T. Expression profile of urothelial transcription factors in bladder biopsies with interstitial cystitis. Int J Urol. 2017;24:632–8.
Chen HZ, Tsai SY, Leone G. Emerging roles of E2Fs in cancer: an exit from cell cycle control. Nat Rev Cancer. 2009;9:785–97.
Wu J, Sabirzhanov B, Stoica BA, Lipinski MM, Zhao Z, Zhao S, et al. Ablation of the transcription factors E2F1-2 limits neuroinflammation and associated neurological deficits after contusive spinal cord injury. Cell Cycle. 2015;14:3698–712.
Carroll K, Gomez C, Shapiro L. Tubby proteins: the plot thickens. Nat Rev Mol Cell Biol. 2004;5:55–63.
Mukhopadhyay S, Jackson PK. The tubby family proteins. Genome Biol. 2011;12:225.
Boggon TJ, Shan WS, Santagata S, Myers SC, Shapiro L. Implication of tubby proteins as transcription factors by structure-based functional analysis. Science. 1999;286:2119–25.
Noben-Trauth K, Naggert JK, North MA, Nishina PM. A candidate gene for the mouse mutation tubby. Nature. 1996;380:534–8.
Acknowledgements
This study was supported by KAKENHI grants 17K16779(B) and 15K10801(C).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Arai, T., Fuse, M., Goto, Y. et al. Molecular pathogenesis of interstitial cystitis based on microRNA expression signature: miR-320 family-regulated molecular pathways and targets. J Hum Genet 63, 543–554 (2018). https://doi.org/10.1038/s10038-018-0419-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-018-0419-x
This article is cited by
-
Biomarkers for Bladder Pain Syndrome/Interstitial Cystitis
Current Bladder Dysfunction Reports (2021)
-
Molecular pathogenesis of triple-negative breast cancer based on microRNA expression signatures: antitumor miR-204-5p targets AP1S3
Journal of Human Genetics (2018)