Abstract
Nonhuman primate (NHP) experimental models have contributed greatly to human health research by assessing the safety and efficacy of newly developed drugs, due to their physiological and anatomical similarities to humans. To generate NHP disease models, drug-inducible methods, and surgical treatment methods have been employed. Recent developments in genetic and developmental engineering in NHPs offer new options for producing genetically modified disease models. Moreover, in recent years, genome-editing technology has emerged to further promote this trend and the generation of disease model NHPs has entered a new era. In this review, we summarize the generation of conventional disease model NHPs and discuss new solutions to the problem of mosaicism in genome-editing technology.
Similar content being viewed by others
Introduction
Laboratory animals have long played a useful role in bridging in vitro research and clinical medicine. Mice are widely recognized as the most popular experimental animal and various disease models have been generated over the years that have greatly contributed to the elucidation of human diseases. Mouse models are genetically well controlled and their fecundity is useful for easily and clearly understanding the relationships between gene abnormalities and diseases. However, the physiological differences between rodents and humans are obvious and, in some research areas, such as the development of new drugs, it is difficult to extrapolate the results from mouse models to humans. Nonhuman primates (NHPs) are suitable experimental animals for ameliorating the species differences between rodents and humans because they have high homology with humans in terms of physiological functions and drug metabolism. In the field of neuroscience research, in particular, there is a high possibility for novel findings in NHPs that are beyond the limitations of rodent research, in that, brain structure and functions of NHPs more closely resemble those of humans.
Until recently, the methods for generating NHP disease models were drug induction, disorders due to surgical procedures, the use of spontaneous disease models, and the generation of transgenic NHPs using developmental engineering approaches. These methods have contributed to the generation of many kinds of disease models; however, target gene knockout (KO), which is the most commonly used genetic modification method for producing mouse disease models, has not been accomplished in NHPs. With the advent of genome editing, a recent and innovative set of genetic modification technologies, it has become possible to generate disease models by modifying specific genes in NHPs. In this review, we summarize methods of genetic modification in NHPs, including the utilization of genome-editing technology, and discuss mosaicism, which is a major concern in NHP genome editing.
Development of genetically engineered NHPs
The development of transgenic NHPs began with a report that described the introduction of a green fluorescent protein (GFP) transgene into rhesus monkeys in 2001 [1]. In this study, a high titer of a pseudotyped replication-defective retroviral vector carrying the GFP gene was injected into the perivitelline space of rhesus monkey oocytes, which were then transplanted into the uteri of surrogate mothers after fertilization. In the analysis of the ensuing animals, insertion of the GFP transgene into somatic cells was confirmed; however, GFP expression was not observed in any accessible tissues. In the same year, the generation of genetically modified rhesus macaques was achieved using replication-defective lentiviral vectors carrying the GFP gene [2]. In this study, GFP expression was confirmed in the placenta and umbilical cord blood in two living offspring but GFP expression was not observed in somatic cells. In 2008, the production of rhesus macaques with the huntingtin (HTT) gene, the gene causing Huntington’s disease (HD), was reported [3]. In this study, a high titer of a lentivirus, carrying exon 1 of the human HTT gene with 84 CAG repeats and the GFP gene, was injected into the perivitelline space of rhesus monkey oocytes, which were then transplanted into the uteri of surrogate mothers after fertilization. Five transgenic macaques with the mutant HTT and GFP were obtained and four of the five candidate animals showed the hallmark features of HD. This study was the first report of an NHP disease model in which genetic modification and associated phenotypes were shown. Although the above three reports are pioneering and showed the potential contributions of genetically modified NHPs in medical research, none of the reports confirmed, at the time of publication, the transmission of the transgene to the next generation of animals for establishing colonies of genetically modified disease model organisms. Therefore, further technical innovation was necessary to stably maintain transgenic NHPs.
To solve this problem, our research group devised a new method of transgenic NHP production using lentiviral vectors and, in 2009, we reported the generation of a transgenic marmoset in which transgenes and their functions were transferred to the next generation [4]. In this study, two key technologies were used to acquire genetically modified animals with high efficiency. One was the artificial expansion of the perivitelline space. If the fertilized egg is suspended in PB1 medium containing 0.25 M sucrose before injecting the virus solution, the perivitelline space expands due to dehydration of the embryo. Consequently, it is possible to easily inject large amounts of viral vector into this space, and rehydrates the embryo with the virus solution (Fig. 1). The other method was the selection of embryos by fluorescent protein expression prior to transfer to surrogate mothers. When the fertilized egg, injected with the virus solution, is cultured for several days, GFP expression can be observed under a fluorescence microscope and fluorescent protein-expressing embryos can be selected for transplantation into surrogate mothers. Therefore, it is guaranteed that all animals will harbor the transgene. Indeed, in our study, by injecting lentiviral vectors carrying enhanced GFP (EGFP) into embryos, and selecting and transplanting based on fluorescence expression, EGFP expression was confirmed in infant tissues and placenta in all five founders. Furthermore, in vitro fertilization of wild-type marmoset oocytes with transgenic sperm resulted in healthy offspring that expressed EGFP. This was the first report of a transgenic NHP with germline transmission, and since its publication, F2 generation animals have been obtained (Fig. 2). In this study, the offspring that were obtained systemically expressed the GFP transgene, and the procedure succeeded in increasing the efficiency of generating transgenic individuals and reducing mosaicism. The use of this technology resulted in the generation of transgenic marmoset and cynomolgus monkey models [5, 6]. In 2016, a new transgenic marmoset line generated with a lentiviral vector carrying the fluorescent calcium sensor GCaMP gene, useful for evaluating nerve cell activity in the brain, was reported [7]. In this study, eight transgenic marmosets were obtained, stable and functional expression of GCaMP was confirmed, and transgene transmission in the gametes of mature individuals was achieved. It is thought that this model will greatly contribute to the future development of in vivo technology capable to monitor cranial nerve activity. Most recently, the generation of polyglutamine (PolyQ) disease model marmosets, for the purpose of analyzing pathological conditions in age-related neurodegenerative diseases, has been reported [6]. In this study, seven transgenic marmosets were obtained using a lentiviral vector carrying the human ataxin 3 gene with 120 CAG repeats encoding an elongated PolyQ stretch. In phenotypic analyses, three marmosets showed age-related neurological symptoms specific to PolyQ disease patients, and transmission of the transgene was also confirmed in the next generation derived from these individuals. This is expected to be a useful disease model, which will exceed the limitations of current rodent models, for obtaining new findings on the pathology and treatment of neurodegenerative diseases.
A new method for viral vector injection. Illustration of the new method for viral injection into embryos using 0.25 M sucrose-supplemented medium before viral injection. The embryos are slightly dehydrated and the perivitelline space expands 1.2-fold to 7.5-fold. Consequently, more viral solution can be injected into the embryos
In macaque monkeys, acquisition of the next generation of mutant HTT transgenic macaques described above was reported in 2015 [8]. In this study, sperm from the three founder generation animals was used for artificial insemination to generate F1 animals. As a result, two F1 animals harboring the transgene derived from the founder animal were obtained. Macaque offspring with the autism-associated methyl-CpG-binding protein 2 (MeCP2) transgene have been successfully acquired by intracytoplasmic injection of sperm, which was developed in xenografted testicular tissue in immunodeficient mice [9, 10]. Phenotypic analysis of these offspring showed that social interactions were reduced in F1 animals compared to the wild type. These improved technologies for generating transgenic NHPs will enable the use of many kinds of NHP models to bridge the gap between basic and clinical sciences.
Target gene modification in NHPs using genome-editing technologies
To analyze the functions of genes and their relevance to diseases in detail, it is necessary to disrupt (KO) the target gene or artificially replace (knock in; KI) the gene sequence. Gene targeting, which uses homologous recombination to replace an endogenous gene with a targeting cassette gene, is the most reliable genetic manipulation method for generating KI and KO animals [11]. For gene targeting in mice, targeting constructs containing the gene mutations of interest are introduced into mouse embryonic stem cells (ESCs). Mouse ESCs can contribute to both somatic and germ cells after ESC injection into mouse host blastocysts. Therefore, when endogenous genes in ESCs are manipulated through target gene KO or KI, these mutated genes will be introduced into the chimeric mouse tissues, including the germline, via ESCs. However, in animal species other than mice and rats, ESCs do not have the competency to generate chimeric animals. In fact, we reported the establishment of ESCs, the induction of pluripotent stem cells (iPSCs), and homologous recombination within ESCs from marmosets [12,13,14]. Nevertheless, ESCs derived from marmosets cannot contribute to somatic tissues or the germline of chimeric animals because they are not in a naive state, which is characteristic of mouse ESCs capable of contributing to the germline [15, 16]. This was originally thought to be a species difference but, in 2007, new stem cells, called epiblast stem cells (EpiSCs), were established from post-implantation embryos [17, 18]. EpiSCs have the ability to differentiate into three germ layers but do not have chimeric competency. Furthermore, the EpiSC colonies were flattened and required basic fibroblast growth factor to maintain an undifferentiated status, and X chromosomes were inactivated in female cell lines. These characteristics are similar to the ESCs of primates, including humans. From these findings, it is thought that primate ESCs are much more similar to mouse EpiSCs, which are derived from post-implantation embryos. Based on several studies, it was hypothesized that there are two different types of pluripotency, namely, a naive type, which has chimeric competency, and a primed type, which does not have chimeric competency [16]. The signaling differences involved in pluripotency between marmosets and mice have also been clarified by embryonic transcriptome analysis [19].
However, the recent development of innovative genome-editing technologies, such as zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs), and clustered regularly interspaced short palindromic repeat/CRISPR-associated protein 9 (CRISPR/Cas9), have enabled the generation of targeted KO or KI animals without the use of ESCs or iPSCs. In NHPs, several investigations have established new approaches for producing targeted KOs or KIs with genome editing tools [20,21,22]. In these approaches, artificially designed nucleases were used to induce specific double-strand breaks at targeted positions in the genome. Double-strand breaks initiate endogenous cellular repair mechanisms, either homology-directed repair (HDR) or nonhomologous end-joining (NHEJ). Through these repair processes, mutations of endogenous genes can be inserted. Importantly, genome editing by engineered nucleases can be applied to all model animals, as HDR and NHEJ are essential functions in all organisms. ZFN is an artificial restriction enzyme composed of a sequence-independent DNA cleavage domain derived from the type II restriction enzyme FokI and a specific DNA-binding domain consisting of a tandem zinc-finger-binding motif [20]. TALEN is a chimeric protein produced by the fusion of the transcription activator-like effector DNA-binding domain and the endonuclease FokI [23]. Furthermore, a highly active form called Platinum TALEN, in which the amino acids of the DNA-binding module have been modified, has been newly developed and genome editing is reportedly possible with high efficiency in various animals and plants [24, 25]. CRISPR/Cas9 is a genome editing system that utilizes the RNA-based adaptive immune system of bacteria and archaebacteria. A simple base pair complementarity between a single-guide RNA that recognizes a target sequence and a target DNA sequence guides the Cas9 nuclease [26, 27]. The innovative development of such genome editing tools has enabled the manipulation of specific genes in many organisms and the generation of NHP disease models has become realistic.
The first generation of genetically modified NHPs by genome editing was reported in cynomolgus monkeys using CRISPR/Cas9 [28]. In this study, simultaneous KO of three target genes (Nr0b1, Ppar-γ, and Rag1) was attempted. To generate the candidate animals, the CRISPR/Cas9 components Cas9 mRNA and single-guide RNAs were injected into one-cell-stage cynomolgus monkey embryos and 19 candidate animals were obtained. Genetic analysis confirmed that two of the target genes (Ppar-γ and Rag1) were modified in the ear tissues and the placenta in two of the 19 animals. However, these genetic modifications occurred in a mosaic manner and phenotypes associated with the modified genes were not confirmed. Although the efficiency of genetic modification was low in this study, it represents a milestone in proving that CRISPR/Cas9 can modify an NHP embryonic genome and that multiple genes can be altered in one step. In the same year, the generation of genetically modified rhesus and cynomolgus monkeys using TALEN technology was reported [29]. In this study, the MeCP2 gene, which is related to autism spectrum disorder, was modified by a TALEN. As a result, male rhesus and cynomolgus fetuses carrying mutations in the MeCP2 gene in various tissues were miscarried at mid-gestation, consistent with Rett syndrome, which is male embryonic lethal in humans. However, one live female cynomolgus monkey was obtained, with abundant MeCP2 mutations in peripheral tissues. Phenotypic analysis of this MeCP2 gene-modified cynomolgus monkey have continued and detailed analyses using eye tracking and magnetic resonance imaging showed similarities with the physiology, behavior, and brain structural characteristics of Rett syndrome in humans [30]. Furthermore, another research group also reported genome editing of the MeCP2 gene in cynomolgus monkeys using a TALEN in the same year [31]. TALEN mRNAs were injected into one-cell-stage cynomolgus monkey embryos and one male candidate cynomolgus monkey was obtained. Unfortunately, the animal failed to survive after birth but genetic analysis confirmed that MeCP2 gene was modified in several tissues in a mosaic manner.
In 2016, our research group focused on improving the efficiency of genome-editing technology in NHPs and reported the generation of immunodeficiency-related interleukin 2 receptor common gamma (IL2RG) KO marmosets using ZFN and TALEN [32]. In this study, we succeeded in the highly efficient generation of phenotypic individuals in the founder generation by carrying out detailed preliminary examinations using a genome editing tool. NHPs, generated by our research, showing symptoms of immunodeficiency are predicted to be a disease model contributing not only to the pathological analysis of immunodeficiency but also to the fields of regenerative medicine and neuroscience in the near future. In the same year, microcephaly-associated MCPH1 KO cynomolgus monkeys were produced using a TALEN. In this study, detailed tests of altered genomic activity using cultured cells and embryos were performed, and one cynomolgus monkey that was obtained as a result of this study recapitulated most of the clinical features observed in microcephalous patients [33].
Reducing mosaicism
Genome-editing technology is expected to continue as a powerful tool for generating NHP disease models but it will be necessary to improve the design of the experiments for targeting NHPs. In many species, including NHPs, phenotypes associated with genetic modification do not always appear due to target genes being modified in a mosaic pattern (Fig. 3). Mosaic mice can generate genetically modified animals in a relatively short period of time by backcrossing but it is difficult to obtain the next generation in NHPs owing to their long lifespan. Also, to maintain NHPs, it is necessary to obtain the maximum number of results with as few individuals as possible, for ethical reasons and due to breeding costs. Furthermore, unlike gene targeting in ESCs, genetic modification efficiency and genetic modification patterns are different in each experiment. To solve this problem, it is important to verify the genome editing tools in detail prior to producing NHP models. This point has been thoroughly studied in our research [32]. First, tools for use in genome editing in NHP embryos can be easily screened by evaluating genome modification activity in cultured cells (Fig. 4a). Next, it is desirable that the selected genome editing tool should allow detailed examination in embryos prior to generating individuals (Fig. 4b). Examining the embryos not only evaluates the efficiency of producing genetically modified individuals but can also evaluate the presence or absence of embryonic lethality due to genome editing tool toxicity and detect biallelic modification of the target gene or off-target effects. In addition, a method recommended for final evaluation, the blastomere splitting test, should be mentioned (Fig. 5). This method analyzes isolated blastomeres by incubating them to around the eight-cell stage after injection of the genome editing tool, making it possible to predict the proportion of mosaicism in the animals to be generated in advance. In 2017, a similar study was also performed in cynomolgus monkey embryos using CRISPR/Cas9 [34]. Recently, in genome editing using CRISPR/Cas9, a method of using Cas9 nuclease in the form of a protein to shorten the timing of target gene modification has become popular because Cas9 nucleases can escape translation from mRNA [35]. This method contributes greatly to a higher efficiency of genetic alteration but mosaicism also occurs, caused by the continued cleavage of the target site by Cas9, which has sustained activity in cells. This group utilized Cas9 protein tagged with an ubiquitin proteasome degradation signal, injected it into cynomolgus monkey embryos, and analyzed the blastomeres after culturing. The use of the mutant Cas9 reduced mosaicism and improved genome editing efficiency compared with that of wild-type Cas9. This research provides useful information for applying genome editing by CRISPR/Cas9 to NHPs with a long generation time.
Illustration of the correlation between genome editing and phenotype. When complete genome editing occurs in the one-cell-stage embryo with the smallest number of target genes, the wild-type gene sequence does not remain and the founder animal displays the expected phenotype (upper). On the other hand, when partial genome editing occurs in embryos that have developed, wild-type and mutant gene sequences are mixed across cells and, in many cases, the expected phenotypes do not appear (middle and lower)
Strategies for evaluating genome editing tools. The illustration of a functional method for selecting the genome editing tool with the highest genome modification activity. a After transfecting candidate genome editing tools into cultured cells, it is possible to roughly confirm the presence or absence of genome modifications by analyzing the genomic DNA of the cells for the targeted gene. b By analyzing the development of an embryo injected with the selected genome editing tool and the target gene using the collected embryonic genome, the ratio of the generation of founder animals to genetically modified animals can be predicted
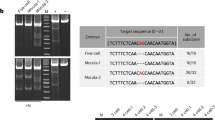
The blastomere splitting method for predicting mosaicism. Illustration of a useful method for predicting mosaicism, which predominantly affects the rate of obtaining phenotypic animals using genome editing. After injecting the candidate genome editing tool into the embryo, the homogeneity of genome modification can be confirmed by isolating the blastomeres of the embryo developed up to the eight-cell-stage and analyzing the target gene. When mutant genes are detected in all blastomeres, the appearance of the desired phenotype in the animals produced is highly likely (left). On the other hand, when the wild-type gene partially remains, it is suggested that the animals will become mosaic and the appearance of the desired phenotype cannot be expected (right). Photographs: Surveyor nuclease (Cel-1) assays visualizing the presence of genetic modifications. The yellow arrowheads are digestion bands showing genetic modifications
Future perspectives for genetically modified NHPs
In recent years, the generation of NHP disease models by genome-editing technology has rapidly developed and is attracting many researchers, especially in the neuroscience field. It is very innovative for the replacement of various studies carried out in conventional mouse disease models with NHPs, which have highly similar brain functions to humans. As a result, it is expected that new findings will be obtained for topics that are difficult to analyze in experimental animals, such as neurodegenerative diseases, psychiatric disorders, and cognitive function. Thus, in the future, further acceleration of the generation of various models in NHPs is expected and efficient generation of the desired animals is critical for maintaining the speed of research as well as for ethical considerations.
References
Chan AW, Chong KY, Martinovich C, Simerly C, Schatten G. Transgenic monkeys produced by retroviral gene transfer into mature oocytes. Science. 2001;291:309–12.
Wolfgang MJ, Eisele SG, Browne MA, Schotzko ML, Garthwaite MA, Durning M, Ramezani A, Hawley RG, Thomson JA, and Golos TG. Rhesus monkey placental transgene expression after lentiviral gene transfer into preimplantation embryos. Proc Natl Acad Soc USA. 2001; 98: 10728–32.
Yang SH, Cheng PH, Banta H, Piotrowska-Nitsche K, Yang JJ, Cheng EC, Snyder B, Larkin K, Liu J, Orkin J, et al. Towards a transgenic model of Huntington’s disease in a non-human primate. Nature. 2008;453:921–4.
Sasaki E, Suemizu H, Shimada A, Hanazawa K, Oiwa R, Kamioka M, Tomioka I, Sotomaru Y, Hirakawa R, Eto T, et al. Generation of transgenic non-human primates with germline transmission. Nature. 2009;459:523–7.
Seita Y, Tsukiyama T, Iwatani C, Tsuchiya H, Matsushita J, Azami T, Okahara J, Nakamura S, Hayashi Y, Hitoshi S, et al. Generation of transgenic cynomolgus monkeys that express green fluorescent protein throughout the whole body. Sci Rep. 2016;6:24868.
Tomioka I, Ishibashi H, Minakawa EN, Motohashi HH, Takayama O, Saito Y, Popiel HA, Puentes S, Owari K, Nakatani T, et al. Transgenic monkey model of the polyglutamine diseases recapitulating progressive neurological symptoms. eNeuro. 2017;4:0250–16.
Park JE, Zhang XF, Choi SH, Okahara J, Sasaki E, Silva AC. Generation of transgenic marmosets expressing genetically encoded calcium indicators. Sci Rep. 2016;6:34931.
Moran S, Chi T, Prucha MS, Ahn KS, Connor-Stroud F, Jean S, Gould K, Chan AW. Germline transmission in transgenic Huntington’s disease monkeys. Theriogenology. 2015;84:277–85.
Liu Z, Li X, Zhang JT, Cai YJ, Cheng TL, Cheng C, Wang Y, Zhang CC, Nie YH, Chen ZF, et al. Autism-like behaviours and germline transmission in transgenic monkeys overexpressing MeCP2. Nature. 2016;530:98–102.
Liu Z, Nie YH, Zhang CC, Cai YJ, Wang Y, Lu HP, Li YZ, Cheng C, Qiu ZL, Sun Q. Generation of macaques with sperm derived from juvenile monkey testicular xenografts. Cell Res. 2016;26:139–42.
Mansour SL, Thomas KR, Capecchi MR. Disruption of the proto-oncogene int-2 in mouse embryo-derived stem cells: a general strategy for targeting mutations to non-selectable genes. Nature. 1988;336:348–52.
Sasaki E, Hanazawa K, Kurita R, Akatsuka A, Yoshizaki T, Ishii H, Tanioka Y, Ohnishi Y, Suemizu H, Sugawara A, et al. Establishment of novel embryonic stem cell lines derived from the common marmoset (Callithrix jacchus). Stem Cells. 2005;23:1304–13.
Shiozawa S, Kawai K, Okada Y, Tomioka I, Maeda T, Kanda A, Shinohara H, Suemizu H, James Okano H, Sotomaru Y, et al. Gene targeting and subsequent site-specific transgenesis at the beta-actin (ACTB) locus in common marmoset embryonic stem cells. Stem Cell Dev. 2011;20:1587–99.
Tomioka I, Maeda T, Shimada H, Kawai K, Okada Y, Igarashi H, Oiwa R, Iwasaki T, Aoki M, Kimura T, et al. Generating induced pluripotent stem cells from common marmoset (Callithrix jacchus) fetal liver cells using defined factors, including Lin28. Genes Cells. 2010;15:959–69.
De Los Angeles A, Loh YH, Tesar PJ, Daley GQ. Accessing naive human pluripotency. Curr Opin Genet Dev. 2012;22:272–82.
Nichols J, Smith A. Naive and primed pluripotent states. Cell Stem Cell. 2009;4:487–92.
Brons IG, Smithers LE, Trotter MW, Rugg-Gunn P, Sun B, Chuva de Sousa Lopes SM, Howlett SK, Clarkson A, Ahrlund-Richter L, Pedersen RA, et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature. 2007;448:191–5.
Tesar PJ, Chenoweth JG, Brook FA, Davies TJ, Evans EP, Mack DL, Gardner RL, McKay RD. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature. 2007;448:196–9.
Boroviak T, Loos R, Lombard P, Okahara J, Behr R, Sasaki E, Nichols J, Smith A, Bertone P. Lineage-specific profiling delineates the emergence and progression of naive pluripotency in mammalian embryogenesis. Dev Cell. 2015;35:366–82.
Carroll D. Genome engineering with zinc-finger nucleases. Genetics. 2011;188:773–82.
Gaj T, Gersbach CA, Barbas CF III. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 2013;31:397–405.
Joung JK, Sander JD. TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol. 2013;14:49–55.
Cermak T, Doyle EL, Christian M, Wang L, Zhang Y, Schmidt C, Baller JA, Somia NV, Bogdanove AJ, Voytas DF. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 2011;39:e82.
Sakuma T, Hosoi S, Woltjen K, Suzuki K, Kashiwagi K, Wada H, Ochiai H, Miyamoto T, Kawai N, Sasakura Y, et al. Efficient TALEN construction and evaluation methods for human cell and animal applications. Genes Cells. 2013;18:315–26.
Sakuma T, Ochiai H, Kaneko T, Mashimo T, Tokumasu D, Sakane Y, Suzuki K, Miyamoto T, Sakamoto N, Matsuura S, et al. Repeating pattern of non-RVD variations in DNA-binding modules enhances TALEN activity. Sci Rep. 2013;3:3379.
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339:819–23.
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM. RNA-guided human genome engineering via Cas9. Science. 2013;339:823–6.
Niu Y, Shen B, Cui Y, Chen Y, Wang J, Wang L, Kang Y, Zhao X, Si W, Li W, et al. Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos. Cell. 2014;156:836–43.
Liu H, Chen Y, Niu Y, Zhang K, Kang Y, Ge W, Liu X, Zhao E, Wang C, Lin S, et al. TALEN-mediated gene mutagenesis in rhesus and cynomolgus monkeys. Cell Stem Cell. 2014;14:323–8.
Chen Y, Yu J, Niu Y, Qin D, Liu H, Li G, Hu Y, Wang J, Lu Y, Kang Y, et al. Modeling Rett syndrome using TALEN-edited MECP2 mutant cynomolgus monkeys. Cell. 2017;169:945–55 e910.
Liu Z, Zhou X, Zhu Y, Chen ZF, Yu B, Wang Y, Zhang CC, Nie YH, Sang X, Cai YJ, et al. Generation of a monkey with MECP2 mutations by TALEN-based gene targeting. Neurosci Bull. 2014;30:381–6.
Sato K, Oiwa R, Kumita W, Henry R, Sakuma T, Ito R, Nozu R, Inoue T, Katano I, Okahara N, et al. Generation of a nonhuman primate model of severe combined immunodeficiency using highly efficient genome editing. Cell Stem Cell. 2016;19:127–38.
Ke Q, Li W, Lai X, Chen H, Huang L, Kang Z, Li K, Ren J, Lin X, Zheng H, et al. TALEN-based generation of a cynomolgus monkey disease model for human microcephaly. Cell Res. 2016;26:1048–61.
Tu Z, Yang W, Yan S, Yin A, Gao J, Liu X, Zheng Y, Zheng J, Li Z, Yang S, et al. Promoting Cas9 degradation reduces mosaic mutations in non-human primate embryos. Sci Rep. 2017;7:42081.
Sung YH, Kim JM, Kim HT, Lee J, Jeon J, Jin Y, Choi JH, Ban YH, Ha SJ, Kim CH, et al. Highly efficient gene knockout in mice and zebrafish with RNA-guided endonucleases. Genome Res. 2014;24:125–31.
Acknowledgements
This work was supported by the Strategic Research Program for Brain Science, “Maintenance of Systems for Creation and Spread of Primate Model Animals;” a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan; a PRESTO grant from the Japan Science and Technology Agency to E.S., as well as a grant from the Funding Program for World-Leading Innovative R&D on Science and Technology (FIRST) “Strategic Exploitation of Neuro-Genetics for Emergence of the Mind” program from the Japanese Cabinet Office to E.S.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. If you remix, transform, or build upon this article or a part thereof, you must distribute your contributions under the same license as the original. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Sato, K., Sasaki, E. Genetic engineering in nonhuman primates for human disease modeling. J Hum Genet 63, 125–131 (2018). https://doi.org/10.1038/s10038-017-0351-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-017-0351-5
This article is cited by
-
Mapping sagittal-plane reference brain atlas of the cynomolgus macaque (Macaca fascicularis) based on consecutive cytoarchitectonic images
Brain Structure and Function (2024)
-
A piggyBac-based platform for genome editing and clonal rhesus macaque iPSC line derivation
Scientific Reports (2021)
-
Advanced model systems and tools for basic and translational human immunology
Genome Medicine (2018)