Abstract
Background
We aimed to determine the genetic relatedness between Staphylococcus epidermidis colonizing breast milk (BM) and BM-fed neonates during the first month of life.
Methods
S. epidermidis was isolated from the stool and skin swabs of 20 healthy term and 49 preterm neonates hospitalized in the neonatal intensive care unit and from the BM of mothers once a week and typed by multilocus variable-number tandem-repeat analysis. Virulence-related genes were determined by PCR.
Results
The gut (95%) and skin (100%) of term neonates were colonized with strains genetically similar to those in BM and carrying mecA and IS256 at low rate (both <6.7%). In preterm neonates, colonization with strains genetically similar to those in BM was low on the skin (34.7%) and in the gut in the first week of life (14.3%), but the prevalence of mecA (>90.6%) and IS256 (>61.7%) was high. By the fourth week, in the gut of preterm neonates the prevalence of mecA (73.8%) and IS256 (18.4%) decreased, but colonization with strains genetically similar to those in BM increased (83.7%).
Conclusion
During early life, the skin and gut of preterm neonates is colonized with S. epidermidis that is distinct from strains found in BM, but gradually the gut is enriched with strains genetically similar to those in BM, as in term neonates.
Similar content being viewed by others
Main
Coagulase-negative staphylococci (CoNS), particularly Staphylococcus epidermidis, are common colonizers of the neonatal skin and gut, but also of breast milk (BM) in mothers (1, 2). However, we recently showed that S. epidermidis strains in the BM of mothers of preterm neonates differ from those causing late-onset sepsis (LOS) in terms of lower prevalence of mecA, ica-operon, and IS256 (3). Therefore, the protective effect of BM on LOS has been hypothesized to be partially attributable to the introduction of less-pathogenic strains that could outcompete more virulent ones in the gut (4). However, the genetic similarity of CoNS strains colonizing the BM and neonatal gut has been demonstrated only in term neonates (2, 5) and not in preterm neonates.
CoNS are also the most common causative agents of LOS in preterm neonates (6). Although staphylococci are considered to enter the bloodstream from the skin through intravascular catheters, the latter are absent in one-third of LOS cases (7). Recent studies demonstrating the presence of subsequently invasive strains in the gut prior to the onset of LOS (4) suggest that translocation from the gut could be another cause of LOS due to CoNS. Moreover, LOS episodes in preterm neonates due to pathogens in unpasteurized BM, such as S. aureus (8), have been reported. We recently demonstrated that mothers of preterm neonates may acquire more pathogenic CoNS strains into their BM (3), thus potentially being a source of subsequently invasive CoNS. Although there could be multiple different sources of LOS caused by CoNS, no studies simultaneously comparing colonizing and LOS-causing strains in preterm neonates have been performed.
We aimed to describe the role of mother’s own BM on colonization of the gut and skin of preterm and term neonates by evaluating the genetic relatedness and the presence of virulence- and resistance-related genes of S. epidermidis in different body sites.
Methods
Study Design
Between January 2014 and December 2015, we recruited preterm neonates (gestational age (GA) <37 weeks) and their mothers if feeding with the mother’s own unpasteurized BM was initiated within the first week of life, from the third-level neonatal intensive care units (NICUs) of Tallinn Children’s Hospital or Tartu University Hospital. Term neonates (GA ≥37 weeks, birth weight (BW) ≥2,500 g) and their mothers were eligible if the neonate was exclusively breast-fed, the mother had not been hospitalized and had not received antibacterial agents between 3 months prior to and 1 month after delivery, and both were healthy. The use of probiotics was not allowed in the study participants. Proven LOS caused by CoNS was diagnosed if after the first 72 h of life at least one clinical (apnea or bradycardia spells, increased oxygen requirement, hyper- or hypothermia, lethargy and hypotonia, feeding intolerance, abdominal distension, hypotension, skin and subcutaneous lesions such as petechial rash, abscesses, sclerema) and one laboratory criterion of sepsis (white blood cell count <5 or >20 × 109 cells/l; immature to total neutrophil ratio >0.2; platelet count <100 × 109 cells/l; C-reactive protein >15 mg/l) was present in addition to at least two blood cultures drawn no more than 48 h apart, positive for the same species of CoNS, or CoNS were isolated from one blood culture with adequate antibacterial treatment given for 5 days or more.
Sample Collection
Stool and skin swabs from neonates and BM from mothers were sampled once a week in the first month of life. Stool samples were collected with a sterile spatula into a sterile container from diapers. A skin area of 1 cm2 of the right axilla was rubbed with a swab moistened in normal saline five times vertically and five times horizontally, and it was then placed into Amies transport medium without charcoal (Copan Italia spa, Brescia, Italy). BM was collected after washing the breast with soap and water, wiping with a clean towel, and cleaning with disinfectant. After drying, ~3 ml of BM was manually expressed into a sterile container after discarding the first drops. Stool samples and BM were stored at −20 °C until transported to −80 °C within 96 h. Skin swabs were stored at +4 °C and cultured within 72 h. Bloodstream isolates were obtained from neonates with clinical signs and symptoms of LOS from blood cultures drawn from an indwelling catheter or via venipuncture.
Isolation and Identification of Staphylococci
Ten-fold dilutions of 0.1 g of stool, 100 μl of BM, and skin swab solutions (prepared by vortexing swab for 1 min in 1 ml of normal saline) in normal saline were cultured in different sectors of mannitol-salt agar plates. The agar plates were incubated in ambient air at +37 °C for 48 h. The number of colonies with an appearance typical of staphylococcus was manually counted in a dilution containing visually separate colonies (about 10–100) and multiplied with the factor of dilution to obtain the total number of colonies. After counting the total number of colonies, five random colonies with an appearance typical of staphylococcus, each of distinct morphology, were chosen. All isolates, including those from blood cultures of neonates with LOS, were identified at the species level by a matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (Bruker Daltonics, Bremen, Germany). All staphylococci were stored in skimmed milk at −80 °C.
Multilocus Variable-Number Tandem-Repeat Analysis
From all S. epidermidis isolates, DNA was extracted by means of the boiling method (9) and subsequently typed by multilocus variable-number tandem-repeat analysis (MLVA), as described elsewhere (10, 11). MLVA loci Se4 and Se8 were excluded because the PCR products were too long to be distinguishable by agarose gel electrophoresis or had low typeability, respectively. The MLVA type (MT) was defined as a distinct MLVA profile that was arbitrarily assigned an integer. Simpson’s index of diversity (SID) and 95% confidence interval (CI) were computed using an online tool on the Comparing Partitions Website (http://www.comparingpartitions.info/; last accessed on 16 September 2016).
Multilocus Sequence Typing
At least one representative isolate of each MLVA type colonizing at least five neonates was selected for multilocus sequence typing, which was performed as described elsewhere (12). Newly detected alleles and sequence types (STs) were submitted to the multilocus sequence typing database (http://pubmlst.org/sepidermidis/).
Virulence- and Resistance-Related Genes
From neonatal stool samples and skin swabs and BM the first isolate of each MLVA type was analyzed for the presence of mecA (13), icaA (14), IS256 (14), and ACME (15), as described elsewhere.
Data Analysis
For statistical analysis, the software program R (version 3.2.2; 2015 The R Foundation for Statistical Computing) was used. Categorical variables were compared using Fisher’s exact test, and continuous variables were compared using the Mann–Whitney test. The Bray–Curtis similarity index was calculated using the EstimateS 9.10 software program (16).
Ethics
The study protocol was approved by the Research Ethics Committee of the University of Tartu. An informed consent form was signed by the mother on behalf of herself and the neonate.
Results
Study Population
A total of 49 preterm and 20 term neonate–mother pairs were enrolled in the study. All term neonates (median (IQR) GA 40 (39–40) weeks, BW 3,651 (3,324–3,970) g; 45% (n=7) were male) were delivered vaginally and discharged a median of (IQR) 2 (1.75–3) days after birth. The majority of preterm neonates were of GA <32 weeks (79.6%; 39/49); half were delivered by cesarean section, and more than three-quarters had a central venous or arterial catheter or required invasive respiratory support or antibacterial treatment (Table 1).
Colonization with Staphylococci
Staphylococci were detected in 248 of 251 (98.8%) BM samples, 275 of 296 (92.9%) stool samples, and 263 of 266 (98.9%) skin swabs. A total of 4,403 isolates, representing 12 staphylococcal species, were obtained. S. epidermidis colonized all neonates and BM and accounted for 66.6% of all isolates (n=2,933), which was followed by S. haemolyticus (n=676; 15.4%; Figure 1). Overall, preterm neonates compared with term neonates were less commonly colonized on the skin and/or in the gut with S. aureus (26.5% (13/49) vs. 85% (17/20); P<0.001), S. hominis (51% (25/49) vs. 90% (18/20); P=0.006), and S. lugdunensis (8.2% (4/49) vs. 60% (12/20); P<0.001), but they were more commonly colonized with S. haemolyticus (91.8% (45/49) vs. 55% (11/20); P=0.001) and S. capitis [55.1% (27/49) vs. 25% (5/20); P=0.045). The skin and/or gut of preterm neonates compared with the BM of their mothers were more commonly colonized with S. hominis (51% (25/49) vs. 26.5% (13/49); P=0.03), S. haemolyticus (91.8% (45/49) vs. 38.8% (19/49); P<0.001), and S. capitis (55.1% (27/49) vs. 6.1% (3/49); P<0.001); the skin and/or gut of term neonates compared with the BM of their mothers were more commonly colonized with S. aureus (85% (17/20) vs. 45% (9/20); P=0.02) and S. haemolyticus (55% (11/20) vs. 15% (3/20); P=0.02). The median (IQR) count of staphylococci in the BM of mothers of term neonates was significantly lower compared with that of mothers of preterm neonates (3.05 (2.40–3.53) vs. 4.61 (4.22–4.99) cfu/ml, respectively; P<0.001) with no changes during the first month of life. The count of staphylococci was a median (IQR) 7.57 (7.06–8.40) cfu/g in term and 7.15 (5.49–8.30) cfu/g in preterm neonates in the first week of life, decreasing significantly by the fourth week of life to a median (IQR) 7.18 (6.36–7.56) cfu/g in term neonates (P=0.01), but not in preterm neonates. The median (IQR) colony count of staphylococci on the skin was lower in term compared with that in preterm neonates (2.94 (2.70–3.17) vs. 5.06 (4.64–5.40) cfu/swab, respectively; P<0.001).
MLVA Types of S. epidermidis
Of 2,933 isolates of S. epidermidis, 2,930 were typeable by MLVA, yielding a total of 434 distinct MLVA types (SID 0.981 (95% CI (0.979–0.983)). The 23 most common MLVA types colonizing at least five neonates accounted for 44.4% (1,301/2,930) of isolates and were found with a similar frequency on the skin and in the gut of neonates. Nine of the twenty-three MLVA types colonized only preterm neonates and their mothers (Figure 2), which were hereafter designated as predominant NICU clones. Of six S. epidermidis isolates causing LOS, four were available for MLVA typing and belonged to the predominant NICU clones (MT96 (n=2), MT241 (n=1), and MT242 (n=1)).
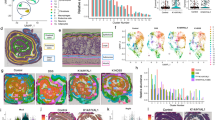
MLVA types (MT) colonizing at least five neonates on the skin and/or in the gut. (a) Prevalence of the MLVA types among preterm neonates (dark blue bars) and their mothers (light blue) and term neonates (dark red) and their mothers (light red). Corresponding sequence types (ST) or single-locus variants (SLVs) of the corresponding ST are shown. The asterisk (*) indicates MLVA types that caused late-onset sepsis. # Isolates of MT379 corresponded to several STs (ST35, ST54, ST142, SLV of ST143, ST487, and its SLV). Upward arrow (↑) indicates predominant NICU clones: i.e., 9 MLVA types of the 23 most common MLVA types that colonized only preterm neonates and their mothers, but not term neonates or their mothers. Note that the y-axis ranges from 0 to 60%. (b) The prevalence of the mecA, icaA, IS256, and ACME in the MLVA types. Each column of the heatmap corresponds to the MLVA types shown in (a). A darker color indicates a higher prevalence, ranging from 0% (in white) to 100% (in black).
In preterm neonates, the diversity of MLVA types decreased from the first to the fourth week of life on the skin (SID (95% CI) 0.92 (0.89–0.94) vs. 0.83 (0.77–0.89)), whereas it increased in the gut (SID (95% CI) 0.91 (0.88–0.94) vs. 0.96 (0.94–0.97)). In contrast, the diversity was higher and did not change during the first month of life in the gut (SID (95% CI) 0.97 (0.96–0.98)) and on the skin (SID (95% CI) 0.97 (0.97–0.98) of term neonates and in the BM of mothers of both term and preterm neonates (SID (95% CI) 0.98 (0.97–0.98) for both).
Relationship Between Colonization of the BM, Gut, and Skin
Genetically similar S. epidermidis MLVA types on the skin and in the gut of neonates were isolated from 95% (n=19) of term and 77.6% (n=38) of preterm neonates. Overall, genetically similar MLVA types in BM and in the gut of the neonate were isolated from 95% (n=19) of term and 83.7% (n=41) of preterm neonates, but the cumulative proportion of preterm neonates colonized with strains present in the BM increased over time from 14.3% (7/49) in the first to 83.7% (41/49) in the fourth week (P<0.001). Genetically similar MLVA types on skin and in the BM were isolated from the skin of all term neonates, but only from 34.7% (n=17) of preterm neonates (P<0.001).
In preterm neonates, genetically similar MLVA types on skin and in BM were isolated from the skin earlier than from BM (Figure 3a); genetically similar MLVA types in gut and in BM were isolated from BM earlier than from the gut; genetically similar MLVA types in gut and on skin were isolated at a similar time in the gut and on the skin. In contrast, in term neonates, genetically similar MLVA types in the gut and in BM were isolated earlier from the gut than from BM (Figure 3b) and genetically similar MLVA types in gut and on skin were isolated from the gut earlier than from the skin.
Violin plots of the age at the first isolation of genetically similar S. epidermidis MLVA types in breast milk and gut, breast milk and skin, and gut and skin in (a) preterm neonates and their mothers and (b) term neonates and their mothers. Each dot represents one mother/neonate colonized with the indicated MLVA type.
Colonization Dynamics of MLVA Types
The gut and skin of term neonates were colonized with MLVA types that were genetically similar to those in BM within the first 2 weeks of life, but in preterm neonates the colonization occurred gradually throughout the first month (Figure 3). According to the Bray–Curtis similarity index, S. epidermidis colonizing the gut of preterm neonates became more similar in terms of the MLVA types to those in BM and less similar to those on skin; the similarity between the skin and BM remained low throughout the first month of life (Figure 4). In term neonates and their mothers, however, all pairwise similarities of the MLVA types between the BM, gut, and skin decreased during the first month of life (Figure 4).
Similarity of S. epidermidis MLVA types in breast milk, gut, and skin in the first four weeks of life in preterm and term neonates and their mothers. The heatmap displays the Bray–Curtis similarity index based on the abundance of MLVA types among all isolates collected from the indicated sample type in the indicated week. Similarity increases from red (Bray–Curtis similarity index 0: i.e., completely dissimilar communities) to green (Bray–Curtis similarity index 1: i.e., identical communities). Overall, populations of S. epidermidis colonizing the skin and gut of preterm neonates or term neonates are genetically similar, but strains from term and preterm neonates are genetically distinct.
Virulence- and Resistance-Related Genes
The prevalence of mecA, icaA, IS256, and ACME was higher in S. epidermidis from hospitalized preterm neonates compared with that in healthy term neonates; mecA, IS256, and ACME were more common in preterm neonates compared with those in their mothers (Table 2). Predominant NICU clones (n=9) harbored mecA, IS256, and ACME (P<0.001 for all) more commonly than did 14 MLVA types that were also isolated from term neonates and their mothers and not considered NICU clones (Figure 2). In preterm neonates, in genetically similar MLVA types in BM and in gut the prevalence of mecA, IS256, and ACME and the proportion of predominant NICU clones were lower compared with that in genetically similar MLVA types in BM and on skin or gut and skin (Table 3).
Dynamics of Virulence- and Resistance-Related Genes
From the first to the fourth week of life, among S. epidermidis strains from the gut of preterm neonates, there was a decrease in the prevalence of mecA (from 93.6% (103/110) to 73.8% (104/141); P<0.001) and IS256 (from 69.1% (76/110) to 18.4% (26/141); P<0.001) as well as in the proportion of predominant NICU clones (from 62.7% (69/110) to 27% (38/141); P<0.001). On the skin of preterm neonates, the prevalence of icaA decreased (from 25.4% (36/142) to 11.1% (14/126); P=0.005), but the proportion of the predominant NICU clones increased (from 53.5% (76/142) to 66.7% (84/126); P=0.039). In the BM of mothers of preterm neonates, the prevalence of mecA increased (38.5% (70/182) vs. 50.3% (85/169); P=0.034). No changes were observed in the prevalence of the genes during the first month of life in term neonates and their mothers.
LOS in Preterm Neonates
A total of 14 episodes of LOS in 11 preterm neonates were caused by CoNS (6 S. epidermidis, 6 S. haemolyticus, 1 S. hominis, 1 mixed S. haemolyticus and S. hominis). Of six S. epidermidis isolates four were available for MLVA typing and belonged to MLVA types MT96 (n=2), MT241 (n=1), and MT242 (n=1). Prior to the onset of LOS, one neonate had LOS-causing strain in the gut, one had on the skin, and one had in both body sites; in the fourth neonate, no prior colonization with LOS-causing strain was detected. None of the mothers of the four preterm neonates with LOS had LOS-causing strain in their BM.
Discussion
To our knowledge, this is the first study describing the role of BM in neonatal colonization by determining genetic relatedness and the presence of virulence- and resistance-related genes of S. epidermidis colonizing the skin and gut of preterm and term neonates and the BM of their mothers. We demonstrated that in BM-fed term neonates the gut and skin of nearly all of them became colonized with strains genetically similar to those in the BM, but the presence of virulence- and resistance-related genes was low. In contrast, the skin and gut of BM-fed preterm neonates were colonized with S. epidermidis strains carrying mecA, icaA, IS256, and ACME genes at high rate and distinct from those colonizing BM. Nevertheless, during the first month of life, more virulent strains in the gut of preterm neonates were replaced by less-virulent S. epidermidis strains genetically similar to those in the mother’s BM despite antibacterial treatment and hospitalization in the NICU, but skin strains remained unchanged. Although BM did not contain subsequently invasive strains, these were detected both in the gut and on the skin of preterm neonates with LOS caused by S. epidermidis.
The difference between term and preterm neonates with respect to gut colonization with S. epidermidis strains genetically similar to those in BM may be attributable to several factors. First, early colonization with mecA and IS256 carrying S. epidermidis-predominant NICU clones, as found in this study, may impede subsequent acquisition of bacteria from other sources such as BM (17). Moreover, during their NICU stay, preterm neonates are continuously exposed to resistant and virulent nosocomial strains (18) that may have colonization advantages because of their adaptation to the hospital environment (19) conferred by, for example, IS256. Indeed, neonates hospitalized in the NICU become colonized with more virulent CoNS strains present on the hands of healthcare workers or in the air in the NICU (20). Second, in preterm neonates, the initiation of BM feeding is often delayed (21), as was in our study (median age of 2 days in preterm vs. 0 days in term neonates). Additionally, because of smaller birth weight the total daily volume of BM is lower than that in term neonates. Moreover, most preterm neonates receive formula or pasteurized donor milk (in our study 89.4% and 36.2%, respectively), which is supposed to be microbe-free and should not contribute to microbial colonization. Finally, the majority of preterm neonates require antibacterial treatment (98% in our study) within the first days of life (22), which could out-select antibiotic-resistant strains. Nevertheless, increasing cumulative amount of administered BM (from 13 ml/kg within the first 3 days to 762 ml/kg within the first 2 weeks of life) probably contributes to gradual colonization with S. epidermidis strains as shown in this study. Importantly, at the same time, the prevalence of mecA- and IS256-carrying strains and the proportion of predominant NICU clones decreased among S. epidermidis colonizing the gut of preterm neonates. Genetically similar MLVA types in BM and in the gut were isolated from BM earlier than from the gut and carried the virulence-related genes at a lower rate than did strains from neonates, suggesting that the mother’s own BM could be a source of less-pathogenic strains, as has been previously hypothesized (4).
In contrast to the gut, strains genetically similar to those in the BM colonized the skin of only one-third of preterm neonates, but of all term neonates. Similarly in another study, from the first minutes of life vaginally delivered healthy term neonates harbored bacterial communities on skin that are very similar to the mother’s microbiota (23). In contrast, preterm neonates are often delivered by cesarean section (44.9% in our study), and because of hospitalization in the NICU within the first hours of life they are separated from the mother shortly after birth; as a result, skin colonization with the mother’s strains is unlikely. Although mothers frequently visit their neonates in NICUs (24), skin-to-skin contact that promotes transfer of susceptible staphylococcal strains from the mother’s skin usually lasts only a few hours (25) and is probably not sufficient to induce colonization. On the contrary, our study suggests that the skin of preterm neonates could be a source of more pathogenic strains for the mother. This is supported by the high prevalence of mecA and IS256, and the high proportion of predominant NICU clones in genetically similar S. epidermidis strains in BM and on skin. Earlier colonization of skin than of BM with genetically similar strains further suggests transfer of microorganisms from the neonate to the mother. Similarly, previous studies have reported the acquisition of S. aureus strains causing outbreaks in the NICU into the microbiota of parents visiting their hospitalized neonates (26).
In the majority of preterm neonates the strains colonizing the skin and gut were genetically similar and the prevalence of virulence- and resistance-related genes was high, particularly within the first weeks of life. This coincides with the time when the incidence of LOS is highest (6), which supports that both body sites could be equally likely reservoirs of potentially invasive strains, previously shown for CoNS isolates resistant to antibiotics (1). This is corroborated by prior skin as well as gut colonization with S. epidermidis strains genetically similar to those in blood cultures of neonates with LOS in the current study. Previous studies have shown that Kangaroo care reduces colonization with methicillin-resistant staphylococci in preterm neonates (25). Our results suggest that gut colonization with virulent S. epidermidis strains could be reduced by the mother’s own BM, which promotes colonization with less-virulent strains and probably is not a source of LOS, as suggested by the absence of subsequently invasive strains in the BM of mothers of preterm neonates with LOS caused by S. epidermidis.
Some limitations of the study should be noted. First, we did not study the overall microbiota in the skin or gut or BM that could influence colonization with staphylococci (27). We aimed to concentrate on staphylococci as they are the most common causative agents of LOS (6). Second, we did not include preterm neonates not receiving BM. Therefore, we cannot exclude that the changes in gut-colonizing S. epidermidis toward lower pathogenicity occur independent of the administration of BM. Nevertheless, it is unlikely that preterm neonates fed with pasteurized donor milk or sterile formula could be exposed to a similarly high count of less-pathogenic staphylococci in unpasteurized BM (1,000 to 10,000 cfu/ml) (3). The importance of BM is supported by the high prevalence of colonization with S. epidermidis that is characteristic to breast-fed rather than to formula-fed neonates (2). Finally, we did not investigate the mother’s skin or NICU environment; as a result, we cannot exclude the probability that the source of less-pathogenic strains for gut was the mother’s skin or NICU environment. Nevertheless, because skin-to-skin contact with the mother promotes colonization of neonates with susceptible staphylococcal strains (25), lack of changes in the skin-colonizing S. epidermidis toward lower pathogenicity in preterm neonates in our study argues against the considerable transfer of staphylococci from the mother’s skin.
In conclusion, we demonstrate that BM has a prominent role in colonizing healthy term neonates with CoNS, whereas its role in hospitalized preterm neonates is less pronounced. Whereas the skin of preterm neonates remains colonized with virulent and resistant strains, the gut is gradually enriched with less-pathogenic strains likely originating from the mother’s BM.
References
Hira V, Kornelisse RF, Sluijter M et al. Colonization dynamics of antibiotic-resistant coagulase-negative Staphylococci in neonates. J Clin Microbiol 2013;51:595–7.
Jiménez E, Delgado S, Maldonado A et al. Staphylococcus epidermidis: a differential trait of the fecal microbiota of breast-fed infants. BMC Microbiol 2008;8:143.
Soeorg H, Metsvaht T, Eelmäe I et al. Coagulase-negative staphylococci in breast milk of mothers of preterm compared with term neonates. J Hum Lact 2017;33:329–340.
Soeorg H, Huik K, Parm U et al. Genetic relatedness of coagulase-negative staphylococci from gastrointestinal tract and blood of preterm neonates with late-onset sepsis. Pediatr Infect Dis J 2013;32:389–93.
Martín V, Maldonado-Barragán A, Moles L et al. Sharing of bacterial strains between breast milk and infant feces. J Hum Lact 2012;28:36–44.
Vergnano S, Menson E, Kennea N et al. Neonatal infections in England: the NeonIN surveillance network. Arch Dis Child Fetal Neonatal Ed 2011;96:F9–14.
Hira V, Sluijter M, Estevão S et al. Clinical and molecular epidemiologic characteristics of coagulase-negative staphylococcal bloodstream infections in intensive care neonates. Pediatr Infect Dis J 2007;26:607–12.
Kayıran PG, Can F, Kayıran SM, Ergonul O, Gürakan B . Transmission of methicillin-sensitive Staphylococcus aureus to a preterm infant through breast milk. J Matern Fetal Neonatal Med 2014;27:527–9.
Zhang K, Sparling J, Chow BL et al. New quadriplex PCR assay for detection of methicillin and mupirocin resistance and simultaneous discrimination of Staphylococcus aureus from coagulase-negative staphylococci. J Clin Microbiol 2004;42:4947–55.
Johansson A, Koskiniemi S, Gottfridsson P, Wiström J, Monsen T . Multiple-locus variable-number tandem repeat analysis for typing of Staphylococcus epidermidis. J Clin Microbiol 2006;44:260–5.
Cremniter J, Sivadon-Tardy V, Caulliez C et al. Genetic analysis of glycopeptide-resistant Staphylococcus epidermidis strains from bone and joint infections. J Clin Microbiol 2013;51:1014–9.
Thomas JC, Vargas MR, Miragaia M, Peacock SJ, Archer GL, Enright MC . Improved multilocus sequence typing scheme for Staphylococcus epidermidis. J Clin Microbiol 2007;45:616–9.
Kondo Y, Ito T, Ma XX et al. Combination of multiplex PCRs for staphylococcal cassette chromosome mec type assignment: rapid identification system for mec, ccr, and major differences in junkyard regions. Antimicrob Agents Chemother 2007;51:264–74.
Ziebuhr W, Krimmer V, Rachid S, Lössner I, Götz F, Hacker J . A novel mechanism of phase variation of virulence in Staphylococcus epidermidis: evidence for control of the polysaccharide intercellular adhesin synthesis by alternating insertion and excision of the insertion sequence element IS256. Mol Microbiol 1999;32:345–56.
Diep BA, Stone GG, Basuino L et al. The arginine catabolic mobile element and staphylococcal chromosomal cassette mec linkage: convergence of virulence and resistance in the USA300 clone of methicillin-resistant Staphylococcus aureus. J Infect Dis 2008;197:1523–30.
EstimateS: Statistical estimation of species richness and shared species from samples, Version 9. 2013. Available at http://purl.oclc.org/estimates. (accessed 26 July 2016).
Cong X, Xu W, Janton S et al. Gut microbiome developmental patterns in early life of preterm infants: impacts of feeding and gender. PLoS ONE 2016;11:e0152751.
Hira V, Sluijter M, Goessens WH et al. Coagulase-negative staphylococcal skin carriage among neonatal intensive care unit personnel: from population to infection. J Clin Microbiol 2010;48:3876–81.
Du X, Zhu Y, Song Y et al. Molecular analysis of Staphylococcus epidermidis strains isolated from community and hospital environments in China. PLoS ONE 2013;8:e62742.
Krediet TG, Jones ME, Janssen K, Gerards LJ, Fleer A . Prevalence of molecular types and mecA gene carriage of coagulase-negative Staphylococci in a neonatal intensive care unit: relation to nosocomial septicemia. J Clin Microbiol 2001;39:3376–78.
Klingenberg C, Embleton ND, Jacobs SE, O'Connell LA, Kuschel CA . Enteral feeding practices in very preterm infants: an international survey. Arch Dis Child Fetal Neonatal Ed 2012;97:F56–61.
Metsvaht T, Nellis G, Varendi H et al. High variability in the dosing of commonly used antibiotics revealed by a Europe-wide point prevalence study: implications for research and dissemination. BMC Pediatr 2015;15:41.
Dominguez-Bello MG, Costello EK, Contreras M et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci USA 2010;107:11971–5.
Garten L, Maass E, Schmalisch G, Bührer C . O father, where art thou? Parental NICU visiting patterns during the first 28 days of life of very low-birth-weight infants. J Perinat Neonatal Nurs 2011;25:342–8.
Lamy Filho F, de Sousa SH, Freitas IJ et al. Effect of maternal skin-to-skin contact on decolonization of methicillin-oxacillin-resistant Staphylococcus in neonatal intensive care units: a randomized controlled trial. BMC Pregnancy Childbirth 2015;15:63.
Conceição T, Aires de Sousa M, Miragaia M et al. Staphylococcus aureus reservoirs and transmission routes in a Portuguese neonatal intensive care unit: a 30-month surveillance study. Microb Drug Resist 2012;18:116–24.
Boix-Amorós A, Collado MC, Mira A . Relationship between milk microbiota, bacterial load, macronutrients, and human cells during lactation. Front Microbiol 2016;7:492.
Acknowledgements
We thank the study nurses Eve Kaur, Marika Zuihhina, and Tuuli Tammekunn, laboratory assistants Dagmar Hoidmets, Tiiu Rööp, and Sandra Sokmann, undergraduate student Magda Karakai, and all study participants.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Statement of financial support
This study was supported by Estonian Research Council (IUT34-24), European Regional Development Fund (Project SFOS WP1-NeuroAIDS), Archimedes Foundation (Project No. 3.2.1001.11-0032) and the European Society for Paediatric Infectious Diseases (ESPID Small Grant Award).
Rights and permissions
About this article
Cite this article
Soeorg, H., Metsvaht, T., Eelmäe, I. et al. The role of breast milk in the colonization of neonatal gut and skin with coagulase-negative staphylococci. Pediatr Res 82, 759–767 (2017). https://doi.org/10.1038/pr.2017.150
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pr.2017.150
This article is cited by
-
Genetic relatedness of Gram-negative bacteria colonizing gut and skin of neonates and mother’s own milk
Journal of Perinatology (2018)
-
Higher intake of coagulase-negative staphylococci from maternal milk promotes gut colonization with mecA-negative Staphylococcus epidermidis in preterm neonates
Journal of Perinatology (2018)
-
Temporal development of the gut microbiome in early childhood from the TEDDY study
Nature (2018)