Abstract
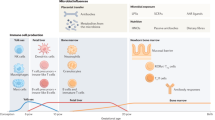
Limited microbial exposure is suggested to underlie the increase of allergic diseases in affluent countries, and bacterial diversity seems to be more important than specific bacteria taxa. Prospective studies indicate that the gut microbiota composition during the first months of life influences allergy development, and support the theory that factors influencing the early maturation of the immune system might be important for subsequent allergic disease. However, recent research indicates that microbial exposure during pregnancy may be even more important for the preventative effects against allergic disease. This review gives a background of the epidemiology, immunology, and microbiology literature in this field. It focuses on possible underlying mechanisms such as immune-regulated epigenetic imprinting and bacterial translocation during pregnancy, potentially providing the offspring with a pioneer microbiome. We suggest that a possible reason for the initial exposure of bacterial molecular patterns to the fetus in utero is to prime the immune system and/or the epithelium to respond appropriately to pathogens and commensals after birth.
Similar content being viewed by others
Low Microbial Exposure and Allergic Disease
Allergic diseases such as atopic eczema, food allergy, asthma, and allergic rhinoconjunctivitis are the most common chronic conditions in children today in affluent countries (1,2). Genetic factors are important in the pathogenesis of these diseases but cannot explain the rapid increase during the last century (3). Instead, the increase has been suggested to be caused by lifestyle changes such as increased stress, exposure to pollutants, drugs (e.g., paracetamol), and changes in nutrition (e.g., less consumption of fish and omega-3 fatty acids) (4). Limited exposure to microbes in developed countries has also been suggested to underlie the allergy development (4,5). Indeed, a decade ago, several studies employing cultivation methods or fluorescent in situ hybridization reported differences in the gut microbiota at a species level between allergic and nonallergic children (6,7,8,9,10). Allergic infants were colonized less with Bacteroides and bifidobacteria species (6,7,8,9), and more with Staphylococcus aureus (8) and Clostridium difficile (10). In some studies, the differences were observed before the onset of allergic symptoms (7,9), indicating that the differences were not simply a consequence of allergic inflammation. However, there is considerable inconsistency between the results from the different studies, and no specific microbes with a consistently harmful or allergy protective role have yet been identified (6,7,8,9,10). Recent reports indicate that a high intestinal microbial diversity during the first month of life is more important than the prevalence of specific bacterial taxa (11,12,13,14,15). The underlying rationale is that the gut immune system reacts to exposure to new bacterial antigens, and repeated exposures enhance the development of immune regulation. By employing barcoded 16S rDNA 454 pyrosequencing in stool samples, we have characterized the total microbiome in stool samples of infants during the first year of life, demonstrating that a low gut microbial diversity during the first month of life precedes the development atopic eczema and asthma in children at 2 (12) and 7 y of age, respectively (14).
Factors suggested to explain individual differences in gut microbiota composition and the risk of developing allergy in infancy are antibiotic treatment (16), caesarean section (17), diet (18), biodiversity in homes (mattresses, dust, etc.) (19,20), in surrounding environment (21), and in family members (skin, mouth, and gut) (22). Concomitant exposure to high levels of allergens and bacteria such as Firmicutes and Bacteroidetes in house dust in inner-city homes during the first year of life has been reported to protect against subsequent recurrent wheeze (23). Also, hygienic practices may influence the microbial diversity and allergy development (24). Recently, children whose parents “cleaned” their pacifier by sucking it were less likely to have asthma at 18 mo of age than children whose parents did not use this cleaning technique (25).
However, despite these findings, only relatively small differences in microbial diversity can be revealed within populations with a homogenous way of living. There is a dramatic difference in gut microbial diversity between wealthy countries such as the United States and less affluent countries such as Malawi (26). Though difficult to prove, a greater historical change in microbial diversity should have occurred during the last century in the industrialized world (27). When John Bostock described “catarrhus aestivus” or hay fever for the first time in 1828, he only succeeded to find 28 cases including himself (28), despite a comprehensive research in all policlinics in London. He writes, “It is remarkable, that all cases are in the middle and upper classes of society, some indeed of high rank… I have not heard of a single unequivocal case among the poor.”
Microbial Exposure During Pregnancy
Prospective studies indicate that it is the gut microbiota composition during the first months of life and not later in life that influences allergy development (12,13,14), supporting the theory that factors influencing the early maturation of the immune system might be especially important for subsequent allergic disease (29). Experimental piglet models indicate that the establishment and the development of a normal gut microbiota and immune system seem to require continuous microbial exposure during the first months of life, and that this process can be compromised under conditions of excessive hygiene in isolators (30). Furthermore, altering the intestinal microbiota during a critical developmental window in early life with low-dose antibiotics also had lasting metabolic effects in a murine model (31). However, recent research indicates that microbial exposure during pregnancy may be even more important for the allergy preventive effect (13,32,33). For example, the well-known protective effect of growing up on farms with livestock seems to be most pronounced in children whose mothers were exposed to stables during pregnancy (32). Atopic sensitization (OR 0.58; 95% CI, 0.39–0.86) was strongly determined by maternal exposure to stables during pregnancy, whereas current exposures to the child had no effects (OR 0.96; 95% CI, 0.63–1.46). Also, the gene expression of toll-like receptors (TLR-2, TLR-4, and CD14) was strongly determined by maternal exposure to stables during pregnancy (32). Antibiotic treatment during pregnancy is also associated with asthma in the offspring (34). Furthermore, prenatal supplementation seems to be crucial for the preventive effect of probiotics on infant eczema (13,33). The overall results from prevention studies published to date indicate that only studies employing the combined administration of the probiotic product prenatally to the mother and postnatally to the mother and/or infant reveal a significant effect on eczema development (35,36,37,38,39), while prenatal (40) or postnatal supplementation alone (41) seems to be ineffective. Thus, it seems that the window of opportunity for intervention begins prior to birth, and likely within the fetal period. Since previous intervention studies started prenatal probiotic supplementation during the last trimester of pregnancy (28,29,30,31,32), it would be interesting to evaluate if starting supplementation from the second trimester of pregnancy (when circulating fetal T cells have developed) will have a more pronounced effect on allergy (27). It would be especially interesting to see this impact on asthma development, which insofar probiotic interventions have failed to prevent (42,43).
Epigenetic Imprinting
A conundrum in allergy epidemiology is the temporal dissociation between the major changes in microbial exposure of children (clean water, fewer siblings, refrigerators, antibiotics, etc.), which occurred during the first half of the century and the onset of the accelerating increase in allergic disease, which happened during the second half ( Figure 1 ). One theory is that a stepwise disappearance of ancestral commensal microbes explains this delayed effect (44). Vertical transmission of maternal microbes is largely responsible for the initial colonization in the gut of the child (45,46). “Heirloom” microbes acquired from the mother’s vaginal and gut microbiota during vaginal delivery may be uniquely evolved to the offspring’s genotype, increasing the chance for optimal mutualism (46). When the microbial diversity diminished in a generation of children, there will be a loss of transmission to the next one, resulting in a cumulative birth cohort phenomenon. Thus, maybe it took more than a generation to reach the point when the microbial diversity was low enough for the allergy boom.
There is a temporal dissociation between the major changes in microbial exposure to children in westernized countries, which occurred during the first half of the century, and the onset of the accelerating increase in allergic diseases that happened during the second half of the century (3).
Epigenetic changes could also explain how the effect of an exposure during childhood is transferred to the next generation ( Figure 2 ). The close immunological interaction between the mother and her offspring during pregnancy provides tremendous opportunities for the maternal microbial environment to influence the offspring’s immune development, possibly via epigenetic mechanisms (33,47). Indeed, microbial exposure to pregnant mice resulted in epigenetic changes of the promoter regions of cytokines associated with an allergic phenotype in an experimental animal model of asthma (48). The pregnant dams were exposed to the gram-negative bacterium Acinetobacter lwoffii, which had previously been isolated from cowshed in farms. The exposure reduced the airway response in the offspring in an IFN-γ-dependent manner. Moreover, the IFN-γ promoter of CD4+ T cells isolated from the spleen of the offspring had high histone-4 acetylation, while the IL-4 promoter region had low histone-4 acetylations, which was closely associated with high Th1-associated IFN-γ and low Th1-associated IL-4, IL-5, and IL-13 cytokine expression (48). The protective effect was not a result of a postnatal colonization of the A. lwoffii in the progeny, since none of stool samples from the offspring were positive for this particular strain. Caesarean section, which is associated with an increased risk of allergic disease (17) is an example of a breach of the initial postnatal transmission of maternal bacteria to the newborn infant (45,49). However, recent reports have shown contradictory results. Elective, as compared with emergency, caesarean section has a weaker association with subsequent allergic disease in children (50,51), suggesting an additional role of prenatal factors and the indications for the caesarean section.
Epigenetic changes of the promoter region of immunoregulatory genes important for allergy development could explain how the effect of microbial exposure to the mother during or even before pregnancy is transferred to the next generation.
There is also evidence from human studies that prenatal microbial exposure results in immunoregulatory epigenetic modifications at birth. For instance, exposure to farms during pregnancy has been associated with increased DNA demethylation of the Foxp3 locus in cord blood cells and enhanced neonatal regulatory T cell function (52), while the promoter regions of the Th2-associated genes RAD50 and IL-13 were hypermethylated in neonates from farming as compared with nonfarming families (53). In these studies, the exposure was confined to pregnancy, but epigenetic changes due to exposures early in life can persist to fertile age (54) and may possibly be transferred to the next generation. Epigenetic analyses of individuals prenatally exposed to the Dutch famine in the winter of 1944–45 in the end of the World War II have confirmed that epigenetic changes lasted up to six decades after the initial exposure (54,55). In the absence of horizontal stimulatory microbial exposures (e.g., siblings in crowded and large families, contaminated drinking water and food with high bacterial counts), there would be cumulative epigenetic changes in every new birth cohort.
Microbial Exposure of the Fetus
A naturally occurring example how the host selects its commensal bacteria is in the fascinating symbiosis between the Hawaiian bobtail squid Euprymna scolopes and the marine luminous bacterium Vibrio fischerii (56). The squid is mono-colonized throughout life with V. fischerii. The host uses the light that is produced by the bacteria to avoid predators during their nocturnal behavior—the host emits luminescence from the ventral surface to mimic down-welling moonlight, thereby casting a disguised silhouette. By investigating the bacterial colonization at the apical surface of the epithelial cells during the first hours of life, several mechanisms underlying this highly complex selection process have been revealed (57). This includes the initial secretion of mucus containing antimicrobial biomolecules (58) and the host release of nitric oxide (59) in response to cell wall derivatives—creating a selective “cocktail” to kill other bacteria but priming the V. fischerii. More importantly, although exposed to the myriad of other microbes, the host responds to the few attaching V. fischerii with changes in expression of genes encoding proteins altering the environment to favor symbiont colonization (57). The question is how this selection is orchestrated in vertebrates. There is little doubt of a genetic component also present in humans. For instance, the gut microbiota in monozygotic twins is more similar than the gut microbiota in dizygotic ones (60). However, given the need of a complex gut microbiota adapted to sudden or long-term changes in environment and nutrition, the selection of bacteria in the human newborn have to be tremendously dynamic.
The “sterile womb” paradigm, postulated by Henry Tissier more than a century ago (61) whereby the sterile fetus first acquires bacteria through passing the birth canal, has been challenged by recent reports showing that infants acquire an initial microbiome already before birth (62,63,64,65). Bacterial DNA has been detected in the placenta (65), umbilical cord (64), amnion fluids (62,63), and the meconium (63) in newborn term infants delivered by sterile caesarean section. Microbial DNA has also been detected in meconium of premature infants born in gestational wk 23–32 (66). Differences in meconium microbiota have been associated with premature birth (67) and subsequent allergy related symptoms in childhood in term neonates (68). In a study with 29 pregnant women, microbial DNA was detected in all placental samples collected during elective caesarean sections. Lactobacillus was the predominant genus (100%) followed by Bifidobacterium (43%), and Bacteroides (34%) (62). Recently, these results were corroborated and extended in a large study with 320 pregnant women, employing16S ribosomal DNA-based and whole-genome shotgun metagenomic studies (69). Identified taxa and their gene carriage patterns were compared with other human body site niches, including the oral, skin, airway (nasal), vaginal, and gut microbiomes from nonpregnant controls. Interestingly, the placental microbiome profiles were most akin to the remote human oral microbiome, which is considered to have many taxa such as Lactobacillus and Streptococcus in common with the upper small intestine. For obvious reasons, the microbiome of the upper small intestine was not assessed in the study. Moreover, using a murine model, genetically labeled Enterococcus faecium strain was only detected in amnion fluid from pups of inoculated dams and not from the pups in the noninoculated control group (63). We have also previously shown an increased prevalence of the probiotic bacterium Lactobacillus reuteri in colostrum after supplementation to pregnant mothers (70). Thus, breastfeeding provides a secondary route of maternal microbial transmission (46). Paradoxically, exclusively breastfed infants have a lower gut microbial diversity than formula-fed ones (71), despite the well-known long-lasting preventive effect of breastfeeding on obesity and overall infection rate (72), further suggesting that the bacteria in the breast milk might be selectively matched to the infant mucosa and immune system.
How bacteria reach the placenta and the mammary glands is still poorly understood, but the genetic material of enteric bacteria has been revealed in maternal mononuclear cells in blood and breast milk from pregnant women. Bacterial translocation from the gut to mesenteric lymph nodes and mammary gland has been shown to occur during late pregnancy and lactation in mice (73). The intestinal epithelial barrier prevents microbial entry into the circulatory system under stable conditions. Dendritic cells, however, can actively penetrate the epithelium, sample microbes from the gut lumen (74) and possibly transport the bacteria to the placenta ( Figure 3 ). Interestingly, this microbial translocation is increased in the mesenteric lymph nodes in pregnant mice (73). Moreover, there is a significant change with a reduction in microbial diversity in the gut from the first to the third trimester in pregnant women (75), further suggesting that the changes in the host–microbial interactions taking place in pregnancy are a physiological phenomenon important for the development of the fetus.
A possible route for bacteria originating from the gut to reach the placental tissue during pregnancy. Dendritic cells (DCs) actively penetrate the epithelium, sample the microbes within the gut lumen, and transport them to the placenta, where they are presented to the fetal epithelium and immune system.
We speculate that the presentation of maternal bacterial components to the fetus during pregnancy is important for the maturation of the immune system and the induction of oral tolerance to allergens. Exposure to live bacteria is not a prerequisite for this phenomenon and this mechanism should not be confused with chorioamnionitis, which is a pathological event. Bacterial DNA in the amniotic fluid and placenta has been shown to affect the expression of TLR-related genes in meconium collected at elective caesarean section in a study on human newborns (62). In a mice asthma model, exposure to A. lwoffii during gestation resulted in a suppression of TLR expression in placental tissue. Furthermore, the asthma-preventive effect of the A. lwoffii treatment was completely abolished in the heterozygous offspring of TLR-2, -3, -4, -7, -9 (−/−) knockout dams (76). However, the previously described allergy prevention studies with probiotics indicated that the preventive effect required both prenatal and postnatal exposure to a certain bacterial strain (35,36,37,38,39). One explanation to these findings is that the initial exposure of bacterial molecular patterns to the fetus during the fetal period primes the immune system and/or the epithelium to respond appropriately to pathogens and commensals after birth. This may occur via epigenetic mechanisms to provide the necessary physiological adaptations for the postnatal environment (33,47). The prenatal programming of the immune system by the maternal bacterial components may be required for a tolerogenic immune response to the copious bacterial load occurring immediately after birth. As a consequence, tolerance to allergens could develop via bystander suppression. Murine models have demonstrated that the neonatal gut microbiota shapes not only intestinal but also systemic immune development, with long-lasting impact on circulating IgE levels (77) and airway hyperactivity (78). Also, the gut microbiota-derived short chain fatty acids butyrate and propionate can stimulate peripheral Treg generation via epigenetically mediated histone deacetylase inhibition, enhancing acetylation of the murine Foxp3 locus (79).
Conclusions
This review has focused on possible mechanisms underlying immune tolerance and allergy prevention in the offspring such as immune-regulated epigenetic imprinting and bacterial translocation to the placenta during pregnancy. Further reductionist models investigating the effect of maternal bacteria on the offspring’s immune and epithelial cells are warranted. The tremendous progress in epigenetic techniques is a promising avenue to address the key questions in the near future. By elucidating the perinatal mechanisms underlying immune tolerance in the infant, new nutritional and treatment strategies can be developed to protect the youngest segment of the population.
Statement of Financial Support
No sponsor has been involved in the writing of the manuscript or the decision to submit it for publication. T.A. and M.J. have received honoraria for lectures and funding for a clinical trial from Biogaia AB, Sweden.
References
Asher MI, Montefort S, Björkstén B, et al. Worldwide time trends in the prevalence of symptoms of asthma, allergic rhinoconjunctivitis, and eczema in childhood: ISAAC Phases One and Three repeat multicountry cross-sectional surveys. Lancet 2006;368:733–43.
Longo G, Berti I, Burks AW, Krauss B, Barbi E . IgE-mediated food allergy in children. Lancet 2013;382:1656–64.
Aberg N . Asthma and allergic rhinitis in Swedish conscripts. Clin Exp Allergy 1989;19:59–63.
Prescott SL . Early-life environmental determinants of allergic diseases and the wider pandemic of inflammatory noncommunicable diseases. J Allergy Clin Immunol 2013;131:23–30.
Holt PG, Sly PD, Björkstén B . Atopic versus infectious diseases in childhood: a question of balance? Pediatr Allergy Immunol 1997;8:53–8.
Björkstén B, Naaber P, Sepp E, Mikelsaar M . The intestinal microflora in allergic Estonian and Swedish 2-year-old children. Clin Exp Allergy 1999;29:342–6.
Björkstén B, Sepp E, Julge K, Voor T, Mikelsaar M . Allergy development and the intestinal microflora during the first year of life. J Allergy Clin Immunol 2001;108:516–20.
Watanabe S, Narisawa Y, Arase S, et al. Differences in fecal microflora between patients with atopic dermatitis and healthy control subjects. J Allergy Clin Immunol 2003;111:587–91.
Kalliomäki M, Kirjavainen P, Eerola E, Kero P, Salminen S, Isolauri E . Distinct patterns of neonatal gut microflora in infants in whom atopy was and was not developing. J Allergy Clin Immunol 2001;107:129–34.
Woodcock A, Moradi M, Smillie FI, Murray CS, Burnie JP, Custovic A . Clostridium difficile, atopy and wheeze during the first year of life. Pediatr Allergy Immunol 2002;13:357–60.
Adlerberth I, Strachan DP, Matricardi PM, et al. Gut microbiota and development of atopic eczema in 3 European birth cohorts. J Allergy Clin Immunol 2007;120:343–50.
Abrahamsson TR, Jakobsson HE, Andersson AF, Björkstén B, Engstrand L, Jenmalm MC . Low diversity of the gut microbiota in infants with atopic eczema. J Allergy Clin Immunol 2012;129:434–40, 440.e1–2.
Ismail IH, Licciardi PV, Tang ML . Probiotic effects in allergic disease. J Paediatr Child Health 2013;49:709–15.
Abrahamsson TR, Jakobsson HE, Andersson AF, Björkstén B, Engstrand L, Jenmalm MC . Low gut microbiota diversity in early infancy precedes asthma at school age. Clin Exp Allergy 2014;44:842–50.
Bisgaard H, Li N, Bonnelykke K, et al. Reduced diversity of the intestinal microbiota during infancy is associated with increased risk of allergic disease at school age. J Allergy Clin Immunol 2011;128:646–52.e1–5.
Alm B, Erdes L, Möllborg P, et al. Neonatal antibiotic treatment is a risk factor for early wheezing. Pediatrics 2008;121:697–702.
Bager P, Wohlfahrt J, Westergaard T . Caesarean delivery and risk of atopy and allergic disease: meta-analyses. Clin Exp Allergy 2008;38:634–42.
Nwaru BI, Takkinen HM, Kaila M, et al. Food diversity in infancy and the risk of childhood asthma and allergies. J Allergy Clin Immunol 2014;133:1084–91.
Ege MJ, Mayer M, Normand AC, et al. Exposure to environmental microorganisms and childhood asthma. N Engl J Med 2011;364:701–9.
Sjögren YM, Jenmalm MC, Böttcher MF, Björkstén B, Sverremark-Ekström E . Altered early infant gut microbiota in children developing allergy up to 5 years of age. Clin Exp Allergy 2009;39:518–26.
Hanski I, von Hertzen L, Fyhrquist N, et al. Environmental biodiversity, human microbiota, and allergy are interrelated. Proc Natl Acad Sci USA 2012;109:8334–9.
Song SJ, Lauber C, Costello EK, et al. Cohabiting family members share microbiota with one another and with their dogs. Elife 2013;2:e00458.
Lynch SV, Wood RA, Boushey H, et al. Effects of early-life exposure to allergens and bacteria on recurrent wheeze and atopy in urban children. J Allergy Clin Immunol 2014; e-pub ahead of print 28 May 2014.
Sherriff A, Golding J . Hygiene levels in a contemporary population cohort are associated with wheezing and atopic eczema in preschool infants. Arch Dis Child 2002;87:26–9.
Hesselmar B, Sjöberg F, Saalman R, Aberg N, Adlerberth I, Wold AE . Pacifier cleaning practices and risk of allergy development. Pediatrics 2013;131:e1829–37.
Yatsunenko T, Rey FE, Manary MJ, et al. Human gut microbiome viewed across age and geography. Nature 2012;486:222–7.
Schnorr SL, Candela M, Rampelli S, et al. Gut microbiome of the Hadza hunter-gatherers. Nat Commun 2014;5:3654.
Bostock J . Of the Catarrhus AEstivus, or Summer Catarrh. Med Chir Trans 1828;14:437–46.
Prescott SL . Early origins of allergic disease: a review of processes and influences during early immune development. Curr Opin Allergy Clin Immunol 2003;3:125–32.
Schmidt B, Mulder IE, Musk CC, et al. Establishment of normal gut microbiota is compromised under excessive hygiene conditions. PLoS One 2011;6:e28284.
Cox LM, Yamanishi S, Sohn J, et al. Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 2014;158:705–21.
Ege MJ, Bieli C, Frei R, et al. Prenatal farm exposure is related to the expression of receptors of the innate immunity and to atopic sensitization in school-age children. J Allergy Clin Immunol 2006;117:817–23.
Jenmalm MC, Duchén K . Timing of allergy-preventive and immunomodulatory dietary interventions—are prenatal, perinatal or postnatal strategies optimal? Clin Exp Allergy 2013;43:273–8.
Benn CS, Thorsen P, Jensen JS, et al. Maternal vaginal microflora during pregnancy and the risk of asthma hospitalization and use of antiasthma medication in early childhood. J Allergy Clin Immunol 2002;110:72–7.
Kalliomäki M, Salminen S, Arvilommi H, Kero P, Koskinen P, Isolauri E . Probiotics in primary prevention of atopic disease: a randomised placebo-controlled trial. Lancet 2001;357:1076–9.
Kukkonen K, Savilahti E, Haahtela T, et al. Probiotics and prebiotic galacto-oligosaccharides in the prevention of allergic diseases: a randomized, double-blind, placebo-controlled trial. J Allergy Clin Immunol 2007;119:192–8.
Abrahamsson TR, Jakobsson T, Böttcher MF, et al. Probiotics in prevention of IgE-associated eczema: a double-blind, randomized, placebo-controlled trial. J Allergy Clin Immunol 2007;119:1174–80.
Wickens K, Black PN, Stanley TV, et al. A differential effect of 2 probiotics in the prevention of eczema and atopy: a double-blind, randomized, placebo-controlled trial. J Allergy Clin Immunol 2008;122:788–94.
Rautava S, Kainonen E, Salminen S, Isolauri E . Maternal probiotic supplementation during pregnancy and breast-feeding reduces the risk of eczema in the infant. J Allergy Clin Immunol 2012;130:1355–60.
Boyle RJ, Ismail IH, Kivivuori S, et al. Lactobacillus GG treatment during pregnancy for the prevention of eczema: a randomized controlled trial. Allergy 2011;66:509–16.
Taylor AL, Dunstan JA, Prescott SL . Probiotic supplementation for the first 6 months of life fails to reduce the risk of atopic dermatitis and increases the risk of allergen sensitization in high-risk children: a randomized controlled trial. J Allergy Clin Immunol 2007;119:184–91.
Kalliomäki M, Salminen S, Poussa T, Isolauri E . Probiotics during the first 7 years of life: a cumulative risk reduction of eczema in a randomized, placebo-controlled trial. J Allergy Clin Immunol 2007;119:1019–21.
Abrahamsson TR, Jakobsson T, Björkstén B, Oldaeus G, Jenmalm MC . No effect of probiotics on respiratory allergies: a seven-year follow-up of a randomized controlled trial in infancy. Pediatr Allergy Immunol 2013;24:556–61.
Blaser MJ, Falkow S . What are the consequences of the disappearing human microbiota? Nat Rev Microbiol 2009;7:887–94.
Jakobsson HE, Abrahamsson TR, Jenmalm MC, et al. Decreased gut microbiota diversity, delayed Bacteroidetes colonisation and reduced Th1 responses in infants delivered by caesarean section. Gut 2014;63:559–66.
Funkhouser LJ, Bordenstein SR . Mom knows best: the universality of maternal microbial transmission. PLoS Biol 2013;11:e1001631.
Renz H, Brandtzaeg P, Hornef M . The impact of perinatal immune development on mucosal homeostasis and chronic inflammation. Nat Rev Immunol 2012;12:9–23.
Brand S, Teich R, Dicke T, et al. Epigenetic regulation in murine offspring as a novel mechanism for transmaternal asthma protection induced by microbes. J Allergy Clin Immunol 2011;128:618–25.e1–7.
Dominguez-Bello MG, Costello EK, Contreras M, et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci USA 2010;107:11971–5.
Tollånes MC, Moster D, Daltveit AK, Irgens LM . Cesarean section and risk of severe childhood asthma: a population-based cohort study. J Pediatr 2008;153:112–6.
Almqvist C, Cnattingius S, Lichtenstein P, Lundholm C . The impact of birth mode of delivery on childhood asthma and allergic diseases—a sibling study. Clin Exp Allergy 2012;42:1369–76.
Schaub B, Liu J, Höppler S, et al. Maternal farm exposure modulates neonatal immune mechanisms through regulatory T cells. J Allergy Clin Immunol 2009;123:774–82.e5.
Michel S, Busato F, Genuneit J, et al. Farm exposure and time trends in early childhood may influence DNA methylation in genes related to asthma and allergy. Allergy 2013;68:355–64.
Heijmans BT, Tobi EW, Stein AD, et al. Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci USA 2008;105:17046–9.
Tobi EW, Lumey LH, Talens RP, et al. DNA methylation differences after exposure to prenatal famine are common and timing- and sex-specific. Hum Mol Genet 2009;18:4046–53.
Nyholm SV, McFall-Ngai MJ . The winnowing: establishing the squid–vibrio symbiosis. Nat Rev Microbiol 2004;2:632–42.
Kremer N, Philipp EE, Carpentier MC, et al. Initial symbiont contact orchestrates host-organ-wide transcriptional changes that prime tissue colonization. Cell Host Microbe 2013;14:183–94.
Troll JV, Bent EH, Pacquette N, et al. Taming the symbiont for coexistence: a host PGRP neutralizes a bacterial symbiont toxin. Environ Microbiol 2010;12:2190–203.
Davidson SK, Koropatnick TA, Kossmehl R, Sycuro L, McFall-Ngai MJ . NO means ‘yes’ in the squid–vibrio symbiosis: nitric oxide (NO) during the initial stages of a beneficial association. Cell Microbiol 2004;6:1139–51.
Stewart JA, Chadwick VS, Murray A . Investigations into the influence of host genetics on the predominant eubacteria in the faecal microflora of children. J Med Microbiol 2005;54:Pt 12:1239–42.
Tissier H . Recherches sur la flore intestinale des nourrissons (état normal et pathologique). Paris: G. Carre and C. Naud, 1900.
Rautava S, Collado MC, Salminen S, Isolauri E . Probiotics modulate host–microbe interaction in the placenta and fetal gut: a randomized, double-blind, placebo-controlled trial. Neonatology 2012;102:178–84.
Jiménez E, Marín ML, Martín R, et al. Is meconium from healthy newborns actually sterile? Res Microbiol 2008;159:187–93.
Jiménez E, Fernández L, Marín ML, et al. Isolation of commensal bacteria from umbilical cord blood of healthy neonates born by cesarean section. Curr Microbiol 2005;51:270–4.
Satokari R, Grönroos T, Laitinen K, Salminen S, Isolauri E . Bifidobacterium and Lactobacillus DNA in the human placenta. Lett Appl Microbiol 2009;48:8–12.
Mshvildadze M, Neu J, Shuster J, Theriaque D, Li N, Mai V . Intestinal microbial ecology in premature infants assessed with non-culture-based techniques. J Pediatr 2010;156:20–5.
Ardissone AN, de la Cruz DM, Davis-Richardson AG, et al. Meconium microbiome analysis identifies bacteria correlated with premature birth. PLoS One 2014;9:e90784.
Gosalbes MJ, Llop S, Vallès Y, Moya A, Ballester F, Francino MP . Meconium microbiota types dominated by lactic acid or enteric bacteria are differentially associated with maternal eczema and respiratory problems in infants. Clin Exp Allergy 2013;43:198–211.
Aagaard K, Ma J, Antony KM, Ganu R, Petrosino J, Versalovic J . The placenta harbors a unique microbiome. Sci Transl Med 2014;6:237ra65.
Abrahamsson TR, Sinkiewicz G, Jakobsson T, Fredrikson M, Björkstén B . Probiotic lactobacilli in breast milk and infant stool in relation to oral intake during the first year of life. J Pediatr Gastroenterol Nutr 2009;49:349–54.
Abrahamsson TR, Jakobsson HE, Andersson AF, Björkstén B, Engstrand L, Jenmalm MC . Reply: gut microbiota diversity and atopic disease: does breastfeeding play a role? J Allergy Clin Immunol 2013;131:248–9.
Hörnell A, Lagström H, Lande B, Thorsdottir I . Breastfeeding, introduction of other foods and effects on health: a systematic literature review for the 5th Nordic Nutrition Recommendations. Food Nutr Res 2013;57.
Perez PF, Doré J, Leclerc M, et al. Bacterial imprinting of the neonatal immune system: lessons from maternal cells? Pediatrics 2007;119:e724–32.
Rescigno M, Urbano M, Valzasina B, et al. Dendritic cells express tight junction proteins and penetrate gut epithelial monolayers to sample bacteria. Nat Immunol 2001;2:361–7.
Koren O, Goodrich JK, Cullender TC, et al. Host remodeling of the gut microbiome and metabolic changes during pregnancy. Cell 2012;150:470–80.
Conrad ML, Ferstl R, Teich R, et al. Maternal TLR signaling is required for prenatal asthma protection by the nonpathogenic microbe Acinetobacter lwoffii F78. J Exp Med 2009;206:2869–77.
Atarashi K, Tanoue T, Shima T, et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 2011;331:337–41.
Olszak T, An D, Zeissig S, et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 2012;336:489–93.
Arpaia N, Campbell C, Fan X, et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 2013;504:451–5.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Abrahamsson, T., Wu, R. & Jenmalm, M. Gut microbiota and allergy: the importance of the pregnancy period. Pediatr Res 77, 214–219 (2015). https://doi.org/10.1038/pr.2014.165
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pr.2014.165
This article is cited by
-
Breastfeeding as a regulating factor of the development of the intestinal microbiome in the early stages of life
European Food Research and Technology (2022)
-
Lactobacillus rhamnosus Used in the Perinatal Period for the Prevention of Atopic Dermatitis in Infants: A Systematic Review and Meta-Analysis of Randomized Trials
American Journal of Clinical Dermatology (2022)
-
Prägung in der Kindheit und Bedeutung im weiteren Leben
Gastro-News (2021)
-
The Role of PPAR-γ in Allergic Disease
Current Allergy and Asthma Reports (2021)
-
The relationship of prenatal antibiotic exposure and infant antibiotic administration with childhood allergies: a systematic review
BMC Pediatrics (2020)