Abstract
Among the many important biopolymers, DNA has been a key component in material sciences and nanotechnology. We have focused on the fabrication of metal nanoarchitectures using DNA as a template due to its intrinsic properties and advantages, such as a well-ordered structure, rich chemical functionality and programmable base-pairing interactions, as well as the availability of multiple enzymes for DNA manipulation. In this review, various methods for the fabrication of DNA-templated metal nanoarchitecture are introduced. The methods include DNA-mediated metal nanoparticle formation, DNA-templated conductive nanowire fabrication by metal depositions, sequence-selective metal deposition onto DNA for elaborate nanowire fabrication and DNA brushes as templates for use on solid substrates. DNA sequence-selective binding of metal ions and metal complexes and subsequent reduction to metals are fundamental issues for the fabrication of metal nanoarchitectures. The resultant metal nanoparticles and their assemblies can be used as functional nanomaterials in applications such as catalysts, conducting nanowires, optical nanomaterials and especially in metamaterials. This biopolymer-templating method can be applied not only to metal deposition but also to the assembly of functional molecules.
Similar content being viewed by others
Introduction
Thanks to the recent developments in nanotechnology there have been a number of important advances in the fields such as electronics, optics, informatics, chemistry, biology and medicine. Nanotechnology was first mentioned in a lecture by Richard Feynman in 1959 and has been expanded markedly to date through the aid of new nanoimaging techniques such as atomic force microscopy (AFM) and electron microscopy. Top-down microfabrication technology such as lithography has contributed to the miniaturization and further integration of electronic substrates. On the other hand, bottom-up technology has been mainly applied to the fabrication and subsequent assembly of fine particles of a few nanometers to several tens of nanometers in size, providing semiconductors or metal nanoparticles with unique optical or photonic properties. For example, semiconductor nanoparticles, called Qdots, emit strong fluorescence and noble metal nanoparticles (for example, gold, silver and so on) show surface plasmon resonance. Such unique properties on the nanostructures of semiconductors and metals have opened up new fields of research. In particular, metal nanostructures have attracted a great deal of attention due to their applications to optical or photonic devices and sensors based on surface plasmon resonance1, 2, 3, 4 and, of course, nanoelectrodes.5, 6, 7
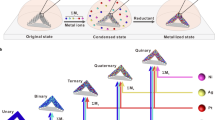
How to prepare more complex and extensive nanoarchitechtures is one of the most important issues in terms of broadening the range of practical applications. In general, top-down-type technology can provide freely designed two-dimensional structures down to several tens of nanometers in size. However, further miniaturization and three-dimensionalization remain challenges. Bottom-up type technology can produce structures of nanometer scale, although significant issues remain in terms of the fabrication and integration of complex structures. Bottom-up systems are prevalent in nature, especially in living bodies. It is a marvelous thing that bottom-up systems work in the production of sophisticated organic nanostructures and their assemblies in vivo. That is, bottom-up fabrication in a living system has managed to overcome the abovementioned issues. RNA and proteins are representatives of this type of fabrication. RNA is a biopolymer composed of four kinds of nucleotides and transcribed from a DNA template. When synthesized, RNAs form various structures according to their sequences and have essential roles in gene coding, decoding, regulation and expression. Proteins are translated from mRNA templates and have an amino-acid sequence as their primary structure. When synthesized, proteins form secondary and tertiary structures, such as alpha helices and beta sheets, according to their sequences. Further, more advanced functional organizations are constructed through assemblies with proteins or RNAs (quaternary structures). These elaborate biomaterials are constructed under extremely complicated conditions with various molecules and produce even longer DNA or proteins with high sequence accuracy, while organic syntheses of oligoDNA or peptides by chemists require relatively high concentration of substances (in the mm to m order) and a pure reaction solution free from impurities. These ideal fabrications in a living body are performed at controlled reaction site based on molecular recognition, templating and self-organization. Toward the mimicking of these good reaction sites using synthetic polymers, much attention has been paid to molecular imprinting methods, but it still remains challenging.8, 9 Therefore, bio-templated and, particularly, DNA-templated synthesis has been studied as a primary choice due to its controllability by sequence and simplicity in handling.10
DNA is an anionic polymer with excellent recognition ability that forms a double-stranded helical structure with quite rigid mechanical properties by Watson–Crick base pairing (A and T or G and C of different chains form complementary base pairs by specific hydrogen bonding). This double-helix formation, called hybridization, is reversible under high temperature and is highly sequence-specific. Because of the benefits of these characteristics, there are a huge number of reports on nanostructure fabrication using DNA hybridization. Mirkin et al.11 demonstrated that two kinds of DNA-modified gold nanoparticles (AuNPs), which have complementary sequences, could form ordered assemblies through hybridization. Shultz and colleagues reported that DNA-attached AuNPs could be set on a complementary DNA at a predetermined distance.12 DNA origami is a most impressive example of DNA nanotechnology first reported by Rothemund.13 In this method, long single-stranded DNA from the M13mp18 phage genome is used as a scaffold and folded into a predetermined nanostructure through hybridization with the aid of a number of appropriately designed short DNAs, called staple strands. Further, controlled bottom-up integration of metal nanoparticles was performed by the use of DNA origami or DNA assemblies as programmable scaffolds.14, 15, 16 Nanoarchitecture fabrications based on DNA hybridization are promising approaches and many well-summarized review papers already exist.17, 18, 19, 20, 21
On the other hand, DNA has other potential uses apart from hybridization in nanoarchitecture fabrications, such as specific binding to molecules, the formation of long rigid structures of easily controllable length, catalytic activities and chirality. Thus, in this review, we describe the recent advances in the fabrication of metal nanoarchitectures using DNA as a biotemplate with particular focus on the following topics: (1) DNA-mediated synthesis of metal nanoparticles; (2) metal depositions on DNAs for conductive nanowire; (3) sequence-selective metallization on DNA; and (4) DNA brushes as novel templates for metal nanoarchitectures to afford another approach to programmable assembly.
DNA-mediated synthesis of metal nanoparticles
DNA possess the ability to bind specifically to metal ions via several different modes of interaction dependent on nucleotide composition.22 Thus, the use of DNA as templates for the syntheses of inorganic nanoparticles to control size and morphology has been an active area of research. Petty and co-workers prepared Ag nanoclusters with a narrow size distribution by utilizing a specific DNA–Ag ion interaction.23, 24 The strong interaction of Ag ions to DNA enabled stoichiometrically controlled complex formation and subsequent chemical reduction with NaBH4 provided size-controlled Ag nanoclusters showing strong fluorescence. The fluorescence of the Ag nanoclusters could be tuned by the sequence of the oligonculeotides used as templates.25, 26 It is noteworthy that the plasmonic absorption of the prepared Ag nanoclusters with short oligonucleotides (short oligonucleotide-encapsulated Ag nanoclusters) showed induced circular dichroism.23 On the other hand, Kotlyar et al.27 reported a large circular dichroism response for silver nanoparticles (AgNPs) grown on the double-stranded long DNA of poly(dG)/poly(dC) as a chiral template of a helix structure at the surface plasmon frequency of AgNPs, suggesting that the chiral DNA template induced the growth of nanoparticles with chirality (Figure 1a).
(a) Circular dichroism spectra measured of gold nanoparticles (AgNPs) grown on the DNA (black), grown in solution without DNA (red) and adsorbed to the DNA (blue).27 (b) Absorption spectra of the suspension before (black) and after (red) ultraviolet irradiation for 5 min. Inset shows the size distribution, transmission electron microscopy (TEM) image and photograph of the dA20-AgNP solution. (c) Fluorescence spectrum and photograph of the dA20-AgNP dispersion under excitation by light at 360 nm. (d) TEM images of AgNPs prepared with dA20, dG20, dC20 or dT20. (e) Illustration (left) and TEM image (right) of the DNA hybridization-directed assembly between the dA20-AgNP and the dT20-AgNP.30 a was adapted with permission from Shemer et al.27 Copyright (2006) American Chemical Society. b–e were adapted from ref. 30 with permission from the Royal Society of Chemistry.
As some DNA shows catalytic activity in reactions with nucleic acid substrates, known as deoxyribozyme (DNAzyme), DNA-catalyzed or DNA-modulated synthesis is another focus of the work on active templates.28 Berti et al.29 utilized the photo-harvesting property of DNA to reduce Ag ions bound to DNA chains. Ultraviolet irradiation of the DNA–Ag ion complexes provided an AgNP plasmon absorbance peak, demonstrating AgNP formation. We have utilized DNA as a modulator for the photo-conversion of a lump of AgCl to functional AgNPs.30, 31, 32 Although silver halides, such as AgCl, are originally photo-reactive, the photo-conversion of AgCl to AgNPs is greatly accelerated in the presence of oligoDNA (dA20; 20 mer of deoxyadenosine phosphate). Prepared AgNPs were 18 nm in diameter with a relatively narrow size distribution and showed plasmonic absorption at around 400 nm in wavelength and strong fluorescence (Figure 1b and c).30 As with previous reports, AgNPs prepared by our method also showed significant sequence dependence on particle size (Figure 1d). Encapsulated dA20 in AgNPs worked as a stabilizer for dispersion in water and could also be utilized as a crosslinker to immobilize dT20-attached functional materials (Figure 1e). On the basis of the kinetic properties, DNA sequence dependence, and effects of solution conditions such as salt concentration and pH, we addressed the detailed mechanism of the DNA-mediated photo-transformation of AgCl to AgNPs.32 Interestingly, Ag/AgCl nanostructures, which are intermediate products of the photo-conversion process, showed remarkable photo-catalytic activity over two orders of magnitude greater than those of previously reported systems.31 Moreover, DNA-encapsulated Ag/AgCl nanoparticles of ca. 40 nm in size exhibited selective photo-catalytic activity in the decomposition of dye molecules; first-order kinetics for positively charged dye molecules and zero-order kinetics for negatively charged ones.
Metal depositions on DNAs for conductive nanowire
Because of the long linear structure with substantial rigidity of the double helix, rich chemical functionality and programmable nature based on high-molecular recognition, DNA molecules have been studied as potential templates for the fabrication of conductive nanowires of <10 nm in width via doping to DNA,33 electroless plating for the reduction of metal ions on DNA34, 35, 36 and so on.29, 37, 38, 39, 40, 41, 42 Braun and co-workers reported AgNP deposition on DNA, in which the DNA bridged the gap between microelectrodes via hybridization, and then silver ions adsorbed on the DNA via electrostatic interaction, followed by reduction with reducing reagents. A repeat of this cycle produced conductive silver nanowires of ca. 100 nm in width and several micrometers in length.43 As mentioned above, Berti et al.29 fabricated a silver nanowire through the photo-reduction of adsorbed silver ions catalyzed by DNA. We have studied a selective electroless deposition method, in which small metal were first attached to the intended sites as a catalyst followed by electroless deposition, which mainly proceeded at those sites with the assistance of the preadsorbed small metal particles. We used cisplatin, which is a platinum-based antineoplastic medicine that binds to DNA in preference to consecutive guanines, as a precursor metal. A silver nanowire of 50–100 nm in width and several micrometers in length was successfully prepared through the reduction of cisplatin to platinum metal, followed by extension and immobilization of DNA onto the substrate via the Langmuir–Blodgett method, and electroless plating of silver onto the platinum catalyst (Figure 2).36, 44 A high electric conductivity was observed on this silver nanowire over a length of 6 μm from the edge of a microelectrode using AFM with an electroconductive probe (conductive AFM measurement).36
On the other hand, there are many reports on the formation of DNA-templated conductive nanowire through coating with electroconductive polymers.45, 46 We have prepared polyaniline-coated DNA as a conductive nanowire through the Ru(bpy)32+-catalyzed photo-redox reaction of N-phenyl-p-phenylenediamine (as an aniline dimer); that is, Ru(bpy)32+-catalyzed photo-polymerization. Further, AuNP–polyaniline hybrid nanowires were prepared through a sequential assembly process consisting of the adsorption of positively charged AuNPs (~1.5 nm in diameter) and Ru(bpy)32+-catalyzed photo-polymerization of aniline dimers between the AuNPs on the DNA (Figure 3).47 Point-contact current imaging AFM experiments revealed that the DNA-templated polyaniline nanowire possessed Schottky emission conduction and AuNP–polyaniline hybrid nanowires had a potential room-temperature Coulomb blockade effect as a one-dimensional array of multiple tunnel junctions between the polyaniline and AuNP. As this Coulomb blockade has been the focus of next-generation electronic devices such as single-electron devices, numerous nanostructures within nanogap electrodes have been studied.48, 49, 50, 51 DNA-templated conductive nanowires could bring about great advances in the development of novel charge-transport devices. Recently, metal deposition was performed on DNA origami as well as on extended straight DNA, providing various-shaped metal nanostructures.6, 52, 53, 54
Schematic illustrations and typical atomic force microscopy images of the DNA-templated fabrication of polyaniline nanowires (a) and gold nanoparticle–polyaniline-alternated nanowires (b). The catalytic cycle of Ru(bpy)32+-catalyzed photo-polymerization of polyaniline is shown in c for reference.47 These figures were reproduced from ref. 47 with permission from the Royal Society of Chemistry. A full color version of this figure is available at Polymer Journal online.
Sequence-selective metallization of DNA
DNA-templated metallization was mostly applied to long naturally derived DNA such as λ-DNA, which has a random sequence, providing homogenous metal depositions. On the other hand, sequence-selective metallization would expand potential applications by enabling the fabrication of more elaborate metal nanostructures. Braun and co-workers reported sequence-specific patterning of DNA metal coating using RecA protein, which is related to DNA repair or homologous recombination (Figure 4).55 First, RecA proteins were assembled on single-stranded DNA to form a nucleoprotein filament, the nucleoprotein then bound to an aldehyde-derivatized double-stranded DNA (dsDNA) molecule at a homologous sequence, forming triple-stranded DNA together with RecA proteins. Next, the sample was incubated in an AgNO3 solution, with the Ag ions reduced by the DNA-bound aldehyde on the DNA but not on the RecA-binding position. RecA prevented Ag deposition by serving as a resist. As a result, a gap was created between the Ag-loaded DNA segments. Finally, subsequent electroless gold deposition was performed on the Ag aggregates, which work as a catalyst, producing two continuous gold wires separated by the predetermined gap. This procedure has been further developed by Braun’s and other research groups.37, 56 The use of RecA is a clever method of utilizing protein-binding ability and DNA molecular recognition. The production of various kinds of precise DNA-templated nanowire, however, requires a multitude of DNAs with adjusted lengths and sequences. Conventional synthetic DNA can only provide ca. 120 mer, in general, which is too short for application to nanowire production. Therefore, enzymatic polymerization provides another promising approach to prepare long DNAs with controlled lengths and sequences as a template. PCR is a widely used tool for amplifying specific dsDNA. Combining PCR and sequence-specific patterning using RecA enables various types of nanoscale programmable metallization.57
(a) Schematic illustrations of the homologous recombination reaction and molecular lithography. (b) Atomic force microscopy (AFM) image of a RecA-bound aldehyde-derivatized DNA. (c) AFM image of the sample after Ag deposition. (d) AFM image of the sample after gold metallization. Inset is a close-up image of the gap. (e) Scanning electron microscopy image of the wire after gold metallization. Scale bars in b through e are 0.5 μm; scale bar in inset d is 0.25 μm.55 These figures were reproduced from Karen et al.55 with permission from the American Association for the Advancement of Science. A full color version of this figure is available at Polymer Journal online.
As an alternative approach to sequence-specific metallization, we have focused on the guanine-selective binding of cisplatin. To utilize guanine-selective cisplatin binding for designed metal depositions, the local control of DNA guanine content is required. For the preparation of this DNA, we studied the unique polymerase reaction mediated by Klenow fragment exonuclease minus (KF−) of Escherichia coli DNA polymerase I. KF− can provide high-molecular-weight dsDNA molecules with a narrow molecular-weight distribution from a pair of complementary oligonucleotides, which has homo-sequence or doublet or triplet repeats, as the template primer through the slippage extension reaction explained by the strand-slippage model.58, 59, 60, 61 First, we synthesized poly(dG)/poly(dC)-poly[d(AT)] as a diblock copolymer composed of continuous G (guanine) and repetitive AT sequences by the slippage extension reaction of the repetitive sequences from three template-primers, dG20, dC10 and dC10d(AT)10.62 Sequence-selective metallization was successfully performed on this diblock copolymer with cisplatin. Although the length of the domains can be changed by adjusting reaction conditions, it has a critical limit in terms of the flexibility of domain control; for example, it is impossible to make more than three domains. Fortunately, nucleotide analogs with unnatural bases offer may allow us to overcome these drawbacks. Because of their utility, various kinds of nucleotide analogs have been developed and are now on the market. Catalogs now offer unnatural nucleotides with 7-deaza-purin bases (7z-dG and 7z-dA), which have been applied to the analysis of the interactions between DNA and proteins,63 or as a marker with a lower redox potential than natural bases.64, 65 As the cisplatin-binding site is the nitrogen atom at the 7-position of the guanine base, 7-deaza-guanine is a good candidate for a non-cisplatin-binding nucleotide, while the chemical structure is similar to the guanine of the nucleotide that cisplatin preferentially binds.66, 67, 68 This similarity in chemical structure also allows it to be used in the enzymatic polymerization by KF− as well as the sequential polymerization of poly(dG)/poly(dC) and poly(7z-dG)/poly(dC) from one simple pair of template-primer DNAs (oligo(dG) and oligo(dC)) as a living polymerization, as the complimentary base, cytosine, is the same (Figure 5).69 In that report, we demonstrated the preparation of a tri-block copolymer composed of poly(dG)/poly(dC) parts and a poly(7z-dG)/poly(dC) part with sequence-selective platinum metal deposition on the GC part of the DNA block copolymers, providing sophisticated nanowires with a nanogap structure. This extended method allows the length and number of segments of the DNA block polymer to be easily controlled by adjusting the reaction time or conditions, such as solution temperature or substrate concentration, as well as the number of reaction cycles. This research indicated that the effective arrangement of molecular recognition, in this case the critical recognition of the 7-position of guanine by cisplatin and a relatively blunt recognition of the nucleotide bases by the polymerase, could expand the possible applications of DNA to nanofabrications.
(a) Schematic illustration of the preparation of tri-block copolymer DNA using Klenow fragment exonuclease minus (exo-) and (b) sequence-selective metallization on this tri-block DNA. (c) Atomic force microscopy image (i) and bird’s-eye view (ii) of the metal-deposited tri-block DNA copolymer.69 These figures were adapted from Mitomo et al.69 with permission from WILEY-VCH Verlag GmbH & Co. KGaA.
Recently, Kotlyar and co-workers reported a novel sequence-selective approach to silver deposition.70 The incubation of oligonucleotide-coated AgNPs of 15 nm in a diameter with poly(dG)/poly(dC) yielded a uniform DNA nanowire of a few nanometer thickness, which is slightly thicker than the original DNA, while neither poly(dA)/poly(dT) nor the random sequence plasmid (pUC19) DNA underwent transition during incubation with the AgNPs. Although the detailed mechanism remains to be clarified, it is thought that the higher affinity of silver to G and C rather than to A and T leads to specific binding and the low ionization potential of guanine provides for the preferential oxidation of silver atoms in the NPs.
DNA brushes as new templates for metal nanoarchitectures
Although there are a large number of reports on DNA-templated nanostructures, which have enormous potential, the integration of these nanostructures on a macroscopic scale with precise control of positioning is required for their practical application. This is one of the critical issues regarding the bottom-up approach, including that for DNA nanotechnology. One possible solution to this issue is fabrication on a substrate. One example of this is nanoparticle assembly to form superlattices via DNA hybridization. Mirkin et al.71 set weak interactions between the AuNPs and the substrate, as well as between the AuNPs covered with DNAs, and then prepared crystals as extensive orientation- and thickness-controlled AuNP superlattices on the DNA-tethered substrate. DNA-tethered substrates, particularly short DNAs such as synthetic oligonucleotides, have been well developed and widely used for biosensing or for biomedical use on DNA microarrays.72, 73 On the other hand, polymer-tethered surfaces, particularly dense surfaces known as polymer brushes, have recently been attracting a good deal of attention in the fields of polymer science and surface chemistry.74, 75, 76, 77 Polymer brushes exhibit many novel and unique properties, such as wettability, adsorption and lubrication, due to the limited polymer chain structure under densely grafted conditions.78, 79, 80 Moreover, polymer brushes can be three-dimensional-patterned (Figure 6a).81, 82 These reports suggest that a DNA brush could provide a good platform as a template for metal nanoarchitectures.
(a) (a) Gray-scale and (b) bitmap images of the Monalisa. (c) Atomic force microscopy topographic image of the Monalisa obtained from PMETAC brushes.81 (b) Measurement of 1 kb DNA brush density and height. (i) Scheme of DNA brush along a density gradient. DNA ends labeled with a fluorophore (blue) excited by total internal reflection fluorescence (TIRF) and by standard epiFL. (ii, upper) The epiFL image of a DNA density gradient. (Lower) DNA density profile in arbitrary fluorescence units along the x axis averaged along the symmetric y axis at various NaCl concentrations (color code bar in ionic strength). Scale bar represents 10 μm. (iii, upper) TIRF image of a DNA density gradient. (Lower) TIRF profiles along the gradient taken as a function of salt (NaCl) concentration. The TIRF profiles support the salt-reflecting compression of height.84 a was adapted from Zhou et al.81 with permission from WILEY-VCH Verlag GmbH & Co. KGaA. A full color version of this figure is available at Polymer Journal online.
The characteristics of the dsDNA brush were reported by Bar-ziv et al.,83, 84 who reported that DNA brushes composed of long dsDNA (0.3–2.5 kb) over 500 chains per μm2 in density-exhibited polymers in an extended state under low-salt conditions. This density is much lower than that of polymer brushes composed of synthetic polymers (Figure 6b).84 This is speculated to result from the rigidity of the double-stranded helical coiled structure and the strongly charged property of the DNA. At a DNA density of 1200 chains per μm2, the interchain distance is estimated to be ca. 30 nm, which could allow the insertion of nanoparticles. Thus, we applied DNA brushes as a template for the immobilization of rod-shaped AuNGs (gold nanorods: AuNRs).85 To provide an appropriate electrostatic attraction, the AuNRs were modified with a mixture of cationic and nonionic surface ligands (chemical structures are shown in Figure 7a). Interestingly, the extinction spectra of the AuNRs adsorbed on the DNA brushes varied with the mixing ratio of the surface ligand; in other words, the extinction spectra were dependent on the electrostatic attraction (Figure 7b). In reverse proportion to the ratio of cationic ligands, the extinction peak for longitudinal surface plasmon resonance, which is located at around 800 nm, became weaker, while that for transverse surface plasmon resonance located at 530 nm did not show any significant change. This result indicates that weak electrostatic attraction leads to the vertical alignment of AuNRs on the substrates. Angle-dependent spectral changes and scanning electron microscopic images supported this notion (Figure 7c–g). Although there have already been several reports on the procedure for the vertical alignment of anisotropic nanoparticles, such as drop casting,86, 87 lithographic nanotemplate-assisted techniques88, 89 and interfacial assembly,90 the extensive self-assembly of rod-shaped nanoparticles with a controlled density remains challenging due to thermal fluctuations and diffusion. Our DNA brush-templated assembly could afford a breakthrough from this perspective. Further, as DNA brush conformation was rather flexible and can change depending on the external condition, such as salt concentration,84 much focus has been placed on a dynamic tuning of AuNR alignments on DNA brushes now.
(a) Schematic illustration of the vertical assembly of gold nanorods (AuNRs) with the assistance of a double-stranded DNA brush. (b) Extinction spectra of AuNRs adsorbed on a DNA brush in a buffer (pH=7.7). Various mixing ratios of surface ligands were used as shown. (c) Scheme showing the extinction of AuNRs under p- and s-polarized light with tilt angles. (d, f) Photos of the cuvette containing the adsorbed AuNRs through a polarizer (p-polarized for d and s-polarrized for f). (e, g) Extinction spectra of AuNRs (cationic:nonionic ligands=1:4) adsorbed on a DNA brush (1200 chains per μm2) under p-polarized (e) and s-polarized (g) light.85
Further, to expand the potential applications of DNA brushes, the fabrication of thicker three-dimensional-nanostructured DNA brushes is needed. In general, when longer polymer chains are applied to a substrate for the preparation of a polymer brush, known as the grafting-to method, the polymer density is lower due to the steric hindrance from the globule structure, leading to a poorly extended state. Therefore, in the field of polymer science, surface-initiated polymerization via the atomic transfer radical polymerization method, or the grafting-from method, has received a good deal of attention for the fabrication of dense polymer brushes composed of long polymers. For DNAs, there have been several reports on the preparation of single-stranded DNA brushes via surface-initiated enzymatic polymerization with a terminal deoxynucleotidyl transferase, or rolling-circle amplification method, as a grafting-from method.91, 92, 93 However, there are only a few reports on the preparation of dsDNA brushes via the grafting-from method, despite the marked interest.94 As mentioned above, we have studied slippage extension reaction by the Klenow fragment, providing long homo-sequence DNA.62, 69 A good living property at the terminus in the slippage extension reaction, such as the atomic transfer radical polymerization, allows the surface-initiated enzymatic polymerization of dsDNA on substrates. Double-stranded oligo(dG)/oligo(dC) was immobilized on the substrate via streptavidin–biotin interaction at one end, and then polymerized by KF− (Figure 8a).95 The length of the polymerized DNA showed good linearity to the reaction time, at least up to 10 kb (Figure 8b and c). The DNA density was constantly over 1500 chains per μm2 independent of the chain length. Photo-patterning by ultraviolet irradiation through a photomask provided two-dimensional-patterned DNA brushes (Figure 8d) and these DNA brushes showed environment-responsive morphologic changes on AFM measurement similar to those in previous reports84 (Figure 8e and f). Step-wise polymerization with various substances provides block copolymer-type DNA brushes. It is expected that successful combination with sequence-selective metallization will afford a three-dimensional-metal nanopatterning technique. Furthermore, although it is not related to metal nanofabrication, due to the specific biodegradability of DNA by nucleases, DNA brushes prepared by this procedure showed great potential as cell culture substrates, enabling the collecting of cells from the substrate by nuclease reaction without any damage.96
(a) Schematic illustration of double-stranded DNA (dsDNA) brush preparation on streptavidin substrates by surface-initiated enzymatic polymerization. (b) Absorption spectra of dsDNA brushes after 15, 30, 45 and 60 min polymerization on the substrate. (c) Agarose gel electrophoresis of the polymerized DNA after removal from the substrate. (d) Fluorescent image of a two-dimensional (2D)-patterned DNA brush stained with SYBR Green I. Scale bars represent 20 μm. (e) Atomic force microscope images (bird’s-eye view) of a 2D-patterned DNA brush and (f) its height in MilliQ water (Merck Millipore, Tokyo, Japan) (i), 1 mm NaCl aq. (ii), 10 mm NaCl aq. (iii), 100 mm NaCl aq. (iv) and under dried conditions (v).95A full color version of this figure is available at Polymer Journal online.
Summary and outlook
DNA has valuable characteristics of not only easy preparation, sequence controllability and sequence-specific binding in hybridization but also a long rigid structure with easily controllable length, recognition of various molecules, catalytic activity, chirality and so on. Thus, DNA has been attracting a good deal of attention as a template for the nanoarchitecture fabrication, particularly metal nanostructures. In this review, we have described DNA-templated metal nanoarchitecture fabrications with particular focus on DNA-mediated metal nanoparticle formation, DNA-templated conductive nanowire fabrication by metal deposition, sequence-selective metal deposition onto DNA for elaborate nanowire fabrication and DNA brushes as novel templates on solid substrates. Various techniques in relation to DNA have been developed based on a huge amount of research work. Now DNA nanotechnology has opened up a new world through its combination with metal nanostructures, which is expected to lead to practical applications in not only nanoelectronics but also nanophotonics and nanomedicine. Again, DNA is an extremely convenient and useful material. Therefore, it is expected that the use of DNAs as templates for metal nanoarchitectures can help us to identify valuable new products such as metamaterials, which are materials engineered to have properties not found in nature, based on their plasmonic properties.
References
Willets, K. A. & Van Duyne, R. P. Localized surface plasmon resonance spectroscopy and sensing. Annu. Rev. Phys. Chem. 58, 267–297 (2007).
Stewart, M. E., Anderton, C. R., Thompson, L. B., Maria, J., Gray, S. K., Rogers, J. A. & Nuzzo, R. G. Nanostructured plasmonic sensors. Chem. Rev. 108, 494–521 (2008).
Chen, H., Ming, T., Zhao, L., Wang, F., Sun, L.-D., Wang, J. & Yan, C.-H. Plasmon–molecule interactions. Nano Today 5, 494–505 (2010).
Mitomo, H., Horie, K., Matsuo, Y., Niikura, K., Tani, T., Naya, M. & Ijiro, K. Active gap SERS for the sensitive detection of biomacromolecules with plasmonic nanostructures on hydrogels. Adv. Opt. Mater. 4, 259–263 (2016).
Lu, W. & Lieber, C. M. Nanoelectronics from the bottom up. Nat. Mater. 6, 841–850 (2007).
Liu, J., Geng, Y., Pound, E., Gyawali, S., Ashton, J. R., Hickey, J., Woolley, A. T. & Harb, J. N. Metallization of branched DNA origami for nanoelectronic circuit fabrication. ACS Nano 5, 2240–2247 (2011).
Jin, Z., Sun, W., Ke, Y., Shih, C.-J., Paulus, G. L. C., Hua Wang, Q., Mu, B., Yin, P. & Strano, M. S. Metallized DNA nanolithography for encoding and transferring spatial information for graphene patterning. Nat. Commun. 4, 1663 (2013).
Whitcombe, M. J., Kirsch, N. & Nicholls, I. A. Molecular imprinting science and technology: a survey of the literature for the years 2004-2011. J. Mol. Recognit. 27, 297–401 (2014).
Alexander, C., Andersson, H. S., Andersson, L. I., Ansell, R. J., Kirsch, N., Nicholls, I. A., O’Mahony, J. & Whitcombe, M. J. Molecular imprinting science and technology: a survey of the literature for the years up to and including 2003. J. Mol. Recognit. 19, 106–180 (2006).
Li, X. & Liu, D. R. DNA-templated organic synthesis: Nature’s strategy for controlling chemical reactivity applied to synthetic molecules. Angew. Chem. Int. Ed. 43, 4848–4870 (2004).
Mirkin, C. A., Letsinger, R. L., Mucic, R. C. & Storhoff, J. J. A DNA-based method for rationally assembling nanoparticles into macroscopic materials. Nature 382, 607–609 (1996).
Alivisatos, A. P., Johnsson, K. P., Peng, X., Wilson, T. E., Loweth, C. J., Bruchez, M. P. & Schultz, P. G. Organization of ‘nanocrystal molecules’ using DNA. Nature 382, 609–611 (1996).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Le, J. D., Pinto, Y., Seeman, N. C., Musier-Forsyth, K., Taton, T. A. & Kiehl, R. A. DNA-templated self-assembly of metallic nanocomponent arrays on a surface. Nano Lett. 4, 2343–2347 (2004).
Sharma, J., Chhabra, R., Liu, Y., Ke, Y. & Yan, H. DNA-templated self-assembly of two-dimensional and periodical gold nanoparticle arrays. Angew. Chem. Int. Ed. 45, 730–735 (2006).
Kuzuya, A., Kaino, M., Hashizume, M., Matsumoto, K., Uehara, T., Matsuo, Y., Mitomo, H., Niikura, K., Ijiro, K. & Ohya, Y. Encapsulation of a gold nanoparticle in a DNA origami container. Polym. J. 47, 177–182 (2014).
Simmel, S. S., Nickels, P. C. & Liedl, T. Wireframe and tensegrity DNA nanostructures. Acc. Chem. Res. 47, 1691–1699 (2014).
Zhang, F., Nangreave, J., Liu, Y. & Yan, H. Structural DNA nanotechnology : state of the art and future perspective. J. Am. Chem. Soc. 136, 11198–11211 (2014).
Saccà, B. & Niemeyer, C. M. DNA origami: the art of folding DNA. Angew. Chem. Int. Ed. 51, 58–66 (2012).
Kuzuya, A. & Ohya, Y. Nanomechanical molecular devices made of DNA origami. Acc. Chem. Res. 47, 1742–1749 (2014).
Kuzuya, A. & Ohya, Y. DNA nanostructures as scaffolds for metal nanoparticles. Polym. J. 44, 452–460 (2012).
Izatt, R. M., Christensen, J. J. & Rytting, J. H. Sites and thermodynamic quantities associated with proton and metal ion interaction with ribonucleic acid, deoxyribonucleic acid, and their constituent bases, nucleosides, and and nucleotides. Chem. Rev. 71, 439–482 (1971).
Petty, J. T., Zheng, J., Hud, N. V. & Dickson, R. M. DNA-templated Ag nanocluster formation. J. Am. Chem. Soc. 126, 5207–5212 (2004).
Ritchie, C. M., Johnsen, K. R., Kiser, J. R., Antoku, Y., Dickson, R. M. & Petty, J. T. Ag nanocluster formation using a cytosine oligonucleotide template. J. Phys. Chem. C 111, 175–181 (2007).
Richards, C. I., Choi, S., Hsiang, J. C., Antoku, Y., Vosch, T., Bongiorno, A., Tzeng, Y. L. & Dickson, R. M. Oligonucleotide-stabilized Ag nanocluster fluorophores. J. Am. Chem. Soc. 130, 5038–5039 (2008).
Gwinn, E. G., O’Neill, P., Guerrero, A. J., Bouwmeester, D. & Fygenson, D. K. Sequence-dependent fluorescence of DNA-hosted silver nanoclusters. Adv. Mater. 20, 279–283 (2008).
Shemer, G., Krichevski, O., Markovich, G., Molotsky, T., Lubitz, I. & Kotlyar, A. B. Chirality of silver nanoparticles synthesized on DNA. J. Am. Chem. Soc. 128, 11006–11007 (2006).
Silverman, S. K. Deoxyribozymes: DNA catalysts for bioorganic chemistry. Org. Biomol. Chem. 2, 2701–2706 (2004).
Berti, L., Alessandrini, A. & Facci, P. DNA-templated photoinduced silver deposition. J. Am. Chem. Soc. 127, 11216–11217 (2005).
Wang, G., Nishio, T., Sato, M., Ishikawa, A., Nambara, K., Nagakawa, K., Matsuo, Y., Niikura, K. & Ijiro, K. Inspiration from chemical photography: accelerated photoconversion of AgCl to functional silver nanoparticles mediated by DNA. Chem. Commun. 47, 9426–9428 (2011).
Wang, G., Mitomo, H., Matsuo, Y., Shimamoto, N., Niikura, K. & Ijiro, K. DNA-templated plasmonic Ag/AgCl nanostructures for molecular selective photocatalysis and photocatalytic inactivation of cancer cells. J. Mater. Chem. B 1, 5899 (2013).
Wang, G., Mitomo, H., Matsuo, Y., Niikura, K., Maeda, M. & Ijiro, K. DNA-modulated photo-transformation of AgCl to silver nanoparticles: visiting the formation mechanism. J. Colloid Interface Sci. 452, 224–234 (2015).
Taniguchi, M. & Kawai, T. DNA electronics. Physica E Low Dimens. Syst. Nanostruct. 33, 1–12 (2006).
Deng, Z. & Mao, C. DNA-templated fabrication of 1D parallel and 2D crossed metallic nanowire arrays. Nano Lett. 3, 1545–1548 (2003).
Ford, W. E., Harnack, O., Yasuda, A. & Wessels, J. M. Platinated DNA as precursors to templated chains of metal nanoparticles. Adv. Mater. 13, 1793–1797 (2001).
Hashimoto, Y., Matsuo, Y. & Ijiro, K. Fabrication of silver nanowires by selective electroless plating of DNA stretched using the LB method. Chem. Lett. 34, 112–113 (2005).
Nishinaka, T., Takano, A., Doi, Y., Hashimoto, M., Nakamura, A., Matsushita, Y., Kumaki, J. & Yashima, E. Conductive metal nanowires templated by the nucleoprotein filaments, complex of DNA and RecA protein. J. Am. Chem. Soc. 127, 8120–8125 (2005).
Ma, Y., Zhang, J., Zhang, G. & He, H. Polyaniline nanowires on Si surfaces fabricated with DNA templates. J. Am. Chem. Soc. 126, 7097–7101 (2004).
Dong, L., Hollis, T., Connolly, B. A., Wright, N. G., Horrocks, B. R. & Houlton, A. DNA-templated semiconductor nanoparticle chains and wires. Adv. Mater. 19, 1748–1751 (2007).
Liqin, D., Hollis, T., Fishwick, S., Connolly, B. A., Wright, N. G., Horrocks, B. R. & Houlton, A. Synthesis, manipulation and conductivity of supramolecular polymer nanowires. Chem. A Eur. J. 13, 822–828 (2007).
Farha Al-Said, S. A., Hassanien, R., Hannant, J., Galindo, M. A., Pruneanu, S., Pike, A. R., Houlton, A. & Horrocks, B. R. Templating Ag on DNA/polymer hybrid nanowires: control of the metal growth morphology using functional monomers. Electrochem. Commun. 11, 550–553 (2009).
Burley, G. A., Gierlich, J., Mofid, M. R., Nir, H., Tal, S., Eichen, Y. & Carell, T. Directed DNA metallization. J. Am. Chem. Soc. 128, 1398–1399 (2006).
Braun, E., Eichen, Y., Sivan, U. & Ben-Yoseph, G. DNA-templated assembly and electrode attachment of a conducting silver wire. Nature 391, 775–778 (1998).
Ijiro, K., Matsuo, Y. & Hashimoto, Y. Fabrication of metal nanowires by electroless plating of DNA. e-J. Surf. Sci. Nanotechnol. 3, 82–85 (2005).
Houlton, A., Pike, A. R., Angel Galindo, M. & Horrocks, B. R. DNA-based routes to semiconducting nanomaterials. Chem. Commun. 1797–1806 (2009).
Watson, S. M. D., Pike, A. R., Pate, J., Houlton, A. & Horrocks, B. R. DNA-templated nanowires: morphology and electrical conductivity. Nanoscale 6, 4027–4037 (2014).
Wang, G. Q., Tanaka, H., Hong, L., Matsuo, Y., Niikura, K., Abe, M., Matsumoto, K., Ogawa, T. & Ijiro, K. Novel charge transport in DNA-templated nanowires. J. Mater. Chem. 22, 13691–13697 (2012).
Negishi, R., Hasegawa, T., Terabe, K., Aono, M., Ebihara, T., Tanaka, H. & Ogawa, T. Fabrication of nanoscale gaps using a combination of self-assembled molecular and electron beam lithographic techniques. Appl. Phys. Lett. 88, 1–4 (2006).
Romero, H. E. & Drndic, M. Coulomb blockade and hopping conduction in PbSe quantum dots. Phys. Rev. Lett. 95, 1–4 (2005).
Cho, C.-H., Kim, B.-H. & Park, S.-J. Room-temperature Coulomb blockade effect in silicon quantum dots in silicon nitride films. Appl. Phys. Lett. 89, 13116 (2006).
Azuma, Y., Hatanaka, T., Kanehara, M., Teranishi, T., Chorley, S., Prance, J., Smith, C. G. & Majima, Y. One by one single-electron transport in nanomechanical Coulomb blockade shuttle. Appl. Phys. Lett. 91, 1–4 (2007).
Pearson, A. C., Liu, J., Pound, E., Uprety, B., Woolley, A. T., Davis, R. C. & Harb, J. N. DNA origami metallized site specifically to form electrically conductive nanowires. J. Phys. Chem. B 116, 10551–10560 (2012).
Schreiber, R., Kempter, S., Holler, S., Schüller, V., Schiffels, D., Simmel, S. S., Nickels, P. C. & Liedl, T. DNA origami-templated growth of arbitrarily shaped metal nanoparticles. Small 7, 1795–1799 (2011).
Geng, Y., Liu, J., Pound, E., Gyawali, S., Harb, J. N. & Woolley, A. T. Rapid metallization of lambda DNA and DNA origami using a Pd seeding method. J. Mater. Chem. 21, 12126 (2011).
Keren, K., Krueger, M., Gilad, R., Ben-Yoseph, G., Sivan, U. & Braun, E. Sequence-specific molecular lithography on single DNA molecules. Science 297, 72–75 (2002).
Keren, K., Berman, R. S., Buchstab, E., Sivan, U. & Braun, E. DNA-templated carbon nanotube field-effect transistor. Science 302, 1380–1382 (2003).
Sharma, R., Davies, A. G. & Wälti, C. Nanoscale programmable sequence-specific patterning of DNA scaffolds using RecA protein. Nanotechnology 23, 365301 (2012).
Kotlyar, A. B., Borovok, N., Molotsky, T., Fadeev, L. & Gozin, M. In vitro synthesis of uniform poly (dG)– poly (dC) by Klenow exo À fragment of polymerase I. Nucleic Acid Res. 33, 525–535 (2005).
Karthikeyan, G., Chary, K. V. R. & Rao, B. J. Fold-back structures at the distal end influence DNA slippage at the proximal end during mononucleotide repeat expansions. Nucleic Acids Res. 27, 3851–3858 (1999).
Schlötterer, C. & Tautz, D. Slippage synthesis of simple sequence DNA. Nucleic Acids Res. 20, 211–215 (1992).
Ji, J., Clegg, N. J., Peterson, K. R., Jackson, A. L., Laird, C. D. & Loeb, L. A. In vitro expansion of GGC:GCC repeats: identification of the preferred strand of expansion. Nucleic Acids Res. 24, 2835–2840 (1996).
Tanaka, A., Matsuo, Y., Hashimoto, Y. & Ijiro, K. Sequence-specifically platinum metal deposition on enzymatically synthesized DNA block copolymer. Chem. Commun. 4270–4272 (2008).
Fletcher, T. M., Salazar, M. & Chen, S. Human telomerase inhibition by 7-deaza-2‘-deoxypurine nucleoside triphosphates †. Biochemistry 35, 15611–15617 (1996).
Pivoňková, H., Horáková, P., Fojtová, M. & Fojta, M. Direct voltammetric analysis of DNA modified with enzymatically incorporated 7-deazapurines. Anal. Chem. 82, 6807–6813 (2010).
Yang, I. V., Ropp, P. A. & Thorp, H. H. Toward electrochemical resolution of two genes on one electrode: Using 7-deaza analogues of guanine and adenine to prepare PCR products with differential redox activity. Anal. Chem. 74, 347–354 (2002).
Takahara, P. M., Rosenzweig, A., Frederick, C., C., Lippard, A. & S. J. Crystal structure of double-stranded DNA containing the major adduct of the anticancer drug cisplatin. Nature 377, 649–652 (1995).
Huang, H., Zhu, L., Reid, B. R., Drobny, G. P. & Hopkins, P. B. Solution structure of a cisplatin-induced DNA interstrand cross-link. Science 270, 1842–1845 (1995).
Onoa, G. B. B., Cervantes, G., Moreno, V. & Prieto, M. J. Study of the interaction of DNA with cisplatin and other Pd(II) and Pt(II) complexes by atomic force microscopy. Nucleic Acids Res. 26, 1473–1480 (1998).
Mitomo, H., Watanabe, Y., Matsuo, Y., Niikura, K. & Ijiro, K. Enzymatic synthesis of a DNA triblock copolymer that is composed of natural and unnatural nucleotides. Chem. Asian J. 10, 455–460 (2015).
Eidelshtein, G., Fardian-Melamed, N., Gutkin, V., Basmanov, D., Klinov, D., Rotem, D., Levi-Kalisman, Y., Porath, D. & Kotlyar, A. Synthesis and properties of novel silver-containing DNA molecules. Adv. Mater. 28, 4839–4844 (2016).
Senesi, A. J., Eichelsdoerfer, D. J., Macfarlane, R. J., Jones, M. R., Auyeung, E., Lee, B. & Mirkin, C. A. Stepwise evolution of DNA-programmable nanoparticle superlattices. Angew. Chem. Int. Ed. 52, 6624–6628 (2013).
Trevino, V., Falciani, F. & Barrera-saldaña, H. A. DNA microarrays : a powerful genomic tool for biomedical and clinical research. Mol. Med. 13, 527–541 (2007).
Stoughton, R. B. Applications of DNA microarrays in biology. Annu. Rev. Biochem. 74, 53–82 (2005).
Zoppe, J. O., Cavusoglu Ataman, N., Mocny, P., Wang, J., Moraes, J. & Klok, H.-A. Surface-initiated controlled radical polymerization: state-of-the-art, opportunities, and challenges in surface and interface engineering with polymer brushes. Chem. Rev. 117, 1105–1318 (2017).
Krishnamoorthy, M., Hakobyan, S., Ramstedt, M. & Gautrot, J. E. Surface-initiated polymer brushes in the biomedical field: Applications in membrane science, biosensing, cell culture, regenerative medicine and antibacterial coatings. Chem. Rev. 114, 10976–11026 (2014).
Azzaroni, O. Polymer brushes here, there, and everywhere: recent advances in their practical applications and emerging opportunities in multiple research fields. J. Polym. Sci. A Polym. Chem. 50, 3225–3258 (2012).
Ayres, N. Polymer brushes: applications in biomaterials and nanotechnology. Polym. Chem. 1, 769 (2010).
Brittain, W. J., Minko, S. & Brittain, W. A structural definition of polymer brushes. J. Polym. Sci. A Polym. Chem. 45, 3505–3512 (2007).
Kobayashi, M., Terayama, Y., Yamaguchi, H., Terada, M., Murakami, D., Ishihara, K. & Takahara, A. Wettability and antifouling behavior on the surfaces of superhydrophilic polymer brushes. Langmuir 28, 7212–7222 (2012).
Kreer, T. Polymer-brush lubrication: a review of recent theoretical advances. Soft Matter 12, 3479–3501 (2016).
Zhou, X., Wang, X., Shen, Y., Xie, Z. & Zheng, Z. Fabrication of arbitrary three-dimensional polymer structures by rational control of the spacing between nanobrushes. Angew. Chem. Int. Ed. 50, 6506–6510 (2011).
Zhou, X., Liu, X., Xie, Z. & Zheng, Z. 3D-patterned polymer brush surfaces. Nanoscale 3, 4929 (2011).
Shemer, G., Atsmon, Y., Karzbrun, E. & Bar-Ziv, R. H. Collective conformations of DNA polymers assembled on surface density gradients. J. Am. Chem. Soc. 134, 3954–3956 (2012).
Bracha, D., Karzbrun, E., Shemer, G., Pincus, P. A. & Bar-ziv, R. H. Entropy-driven collective interactions in DNA brushes on a biochip. Proc. Natl Acad. Sci. USA 110, 4534–4538 (2013).
Nakamura, S., Mitomo, H., Aizawa, M., Tani, T., Matsuo, Y., Niikura, K., Pike, A., Naya, M., Shishido, A. & Ijiro, K. DNA brush-directed vertical alignment of extensive gold nanorod array with controlled density. ACS Omega 2, 2208–2213 (2017).
Baker, J. L., Widmer-Cooper, A., Toney, M. F., Geissler, P. L. & Alivisatos, A. P. Device-scale perpendicular alignment of colloidal nanorods. Nano Lett. 10, 195–201 (2010).
Singh, A., Gunning, R. D., Ahmed, S., Barrett, C. A., English, N. J., Garate, J.-A. A. & Ryan, K. M. Controlled semiconductor nanorod assembly from solution: influence of concentration, charge and solvent nature. J. Mater. Chem. 22, 1562–1569 (2012).
Thai, T., Zheng, Y., Ng, S. H., Mudie, S., Altissimo, M. & Bach, U. Self-assembly of vertically aligned gold nanorod arrays on patterned substrates. Angew. Chem. Int. Ed. 51, 8732–8735 (2012).
Flauraud, V., Mastrangeli, M., Bernasconi, G. D., Butet, J., Alexander, D. T. L., Shahrabi, E., Martin, O. J. F. & Brugger, J. Nanoscale topographical control of capillary assembly of nanoparticles. Nat. Nanotechnol. 12, 73–80 (2017).
Kim, K., Han, H. S., Choi, I., Lee, C., Hong, S., Suh, S., Lee, L. P. & Kang, T. Interfacial liquid-state surface-enhanced Raman spectroscopy. Nat. Commun. 4, 2182 (2013).
Chow, D. C., Lee, W., Zauscher, S. & Chilkoti, A. Enzymatic fabrication of DNA nanostructures : extension of a self-assembled oligonucleotide monolayer on gold arrays. J. Am. Chem. Soc. 5, 14122–14123 (2005).
Chow, D. C. & Chilkoti, A. Surface-initiated enzymatic polymerization of DNA. Langmuir 23, 11712–11717 (2007).
Barbee, K. D., Chandrangsu, M. & Huang, X. Fabrication of DNA polymer brush arrays by destructive micropatterning and rolling-circle amplification. Macromol. Biosci. 11, 607–617 (2011).
Chi, Y. S., Jung, Y. H., Choi, I. S. & Kim, Y. Surface-initiated growth of poly d(A-T) by Taq DNA polymerase. Langmuir 21, 4669–4673 (2005).
Mitomo, H., Nakamura, S., Suzuki, Y., Matsuo, Y., Niikura, K. & Ijiro, K. Preparation and characterization of double-stranded DNA brushes via surface-initiated enzymatic polymerization. J. Nanosci. Nanotechnol. in press
Mitomo, H., Eguchi, A., Suzuki, Y., Matsuo, Y., Niikura, K., Nakazawa, K. & Ijiro, K. Fabrication of a novel cell culture system using DNA-grafted substrates and DNase. J. Biomed. Nanotechnol. 12, 286–295 (2016).
Acknowledgements
This work was supported in part by the ‘Dynamic Alliance for Open Innovation Bridging Human, Environment and Materials’ from the Ministry of Education, Culture, Sports, Science and Technology of Japan (MEXT). This work was performed under the Cooperative Research Program of the ‘Network Joint Research Center for Materials and Devices’. A part of this work was conducted at Hokkaido University, supported by the ‘Nanotechnology Platform’ Program of the MEXT, Japan. Support from the Noguchi Institute (HM) is also acknowledged.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Ijiro, K., Mitomo, H. Metal nanoarchitecture fabrication using DNA as a biotemplate. Polym J 49, 815–824 (2017). https://doi.org/10.1038/pj.2017.63
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pj.2017.63