Abstract
Poly(amino acids) and polypeptides have the potential to contribute significantly to a biomass-based and sustainable society, due to their biomass origin, functionality, and unique physical properties. To realize amino acid-based polymers as eco-friendly alternatives for petroleum materials, the synthesis of poly(amino acid)s/polypeptides through an environmentally friendly process is needed. In this focus review, the author summarizes the recent progress of chemo-enzymatic polymerization, which is a green and atom-economical reaction that provides new insight into the design of materials from polypeptides. Additionally, polypeptides can be designed to serve as functional and structural materials. The use of peptides as carriers of nucleic acids for delivery into target cells and organelles is one important application of such functional materials. Studies on polypeptides as structural materials are also reviewed.
Similar content being viewed by others
Poly(amino acid) as an eco-friendly material
Eco-friendly polymeric materials have been designed and processed primarily from bio-based polyesters such as poly(hydroxyalkanoate)1 and poly(lactic acid),2, 3 due to their plasticity, excellent workability, and biomass origin.4, 5, 6, 7, 8, 9, 10, 11, 12 Poly(hydroxyalkanoate) has several advantages such as the ability to be directly synthesized from various types of biomass, including lignin and carbon dioxide,13, 14, 15, 16, 17 and excellent biodegradability.18, 19 However, because of a lack of toughness, this biopolyester has limited use and application. Development of an engineered biopolymer such as a biomass-derived polyamide is an emerging interest in the field of sustainable chemistry and materials engineering. Biopolyamides (for example, nylon 4) have been investigated as candidates for biomass-based engineering plastics, but their narrow processing window for thermo-forming currently prevents them from being considered for practical purposes.20 Natural high-performance materials, including spider silk and mussel-derived adhesive (Figure 1), are mainly composed of poly(amino acid)s and a small amount of short peptides, suggesting that poly(amino acid)s could serve as an alternative to the petroleum-derived polymers in support of a sustainable society. Although spider dragline silk is a benchmark in terms of tough materials, scientific and technological advances still cannot produce an artificial dragline silk. To create and develop biomass-based tough polymers like this silk, the synthesis and design of materials from poly(amino acid)s is being investigated.
A poly(amino acid) is a polymer composed of amino acids as monomeric units. Structural and functional proteins, polypeptides, peptides and polymers derived from amino acids, that is, poly(β-alanine) and ɛ-poly(lysine), are classified as poly(amino acid)s. The use of poly(amino acid)s as functional materials has been widely studied, resulting in the development of polypeptides that are bioactive and that can have biological applications such as in tissue engineering, regenerative medicine and drug/gene delivery systems. Poly(amino acid)s are further endowed with remarkable biological functionality such as a target specificity, and their degradation products can be readily metabolized. Due to those functions, various hybrid poly(amino acid)s with synthetic polymers have also been studied as functional materials.21 In contrast, the use of poly(amino acid)s as structural materials is still limited and challenging, due to a lack of understanding of the structure–function relationship of poly(amino acid)-based materials that contain multiple types of water molecules. The role of water molecules in thermal stability and the biological and mechanical properties of poly(amino acid)s have been characterized using X-ray and differential scanning calorimetry.22 However, this information has not been sufficient to develop artificial poly(amino acid) materials with the desired thermal stability and toughness. Furthermore, a major drawback of poly(amino acid)-based materials is the limited repertoire of synthesis methods. To synthesize polypeptides and poly(amino acid)s, current synthesis techniques, such as solid-phase peptide synthesis and recombinant DNA techniques, still have limitations in their production capacity, atom economy and sequence regulation.23, 24, 25 Synthesis through an environmentally friendly process will be needed to establish poly(amino acid) as bio-based materials.
Chemo-enzymatic synthesis of poly(amino acid)s
To establish poly(amino acid)s as functional and structural materials, enantiomerically pure, high-yield, low-cost, atom-economical synthesis strategies are needed. Chemo-enzymatic synthesis is one possible solution for the efficient synthesis of poly(amino acid)s, with respect to yield, reaction time and atom economy (Figure 2a).26 Amino-acid ligases, as well as proteases, have been shown to mediate the formation of peptide bonds between α-amino acid monomers27, 28 and artificial enzymes that catalyze such a reaction may also be developed in the near future.29, 30 In the case of proteases, the amide-forming reaction is either a thermodynamically or a kinetically controlled process. The latter reaction is more selective in that the enzyme needs to react with an ester compound to form an acyl-enzyme intermediate. Competing with water molecules for hydrolysis to proceed, the enzyme will react with an amino acid-derived nucleophile and form a new peptide bond.31, 32 Compared with other synthetic methods, such as solid-phase peptide synthesis, ring-opening polymerization of the α-amino acid N-carboxyanhydride, and recombinant DNA methods,23, 24, 25 chemo-enzymatic synthesis demonstrates a relatively high yield without the need for multiple purifications, has a short reaction time, is atom-economical, and requires only mild reaction conditions. However, the disadvantage of chemo-enzymatic synthesis is that there is less control over the molecular weight and amino-acid sequence of the polypeptide/poly(amino acid) produced.
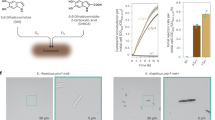
Chemo-enzymatic synthesis of peptide/poly(amino acid)s. (a) Schematic illustration of chemo-enzymatic synthesis of peptide/poly(amino acid)s from amino-acid esters using a protease as a catalyst. The reaction is performed in a buffer solution under a mild reaction temperature and pH. Successful syntheses of several types of homo, random, block and branched peptides have been reported. (b) Reaction schemes for the enzymatic synthesis of poly(L-tyrosine-r-3,4-dihydroxy-L-phenylalanine-r-L-lysine) (P(Tyr-DOPA-Lys)), via poly(L-tyrosine-r-L-lysine) (P(Tyr-Lys)), from L-tyrosine ethyl ester (Tyr-Et) and L-lysine ethyl ester (Lys-Et). Figures modified from Numata et al.53
Catalysts for the chemo-enzymatic polymerization of amino acids are generally proteases and peptidases, because of their chemical and physical stability. Papain, a cysteine protease that exhibits endo-peptidase, amidase and esterase activities and that originates from the latex of the tropical papaya fruit (Carica papaya), has been used as an enzymatic catalyst for chemo-enzymatic polymerization of amino acids.33, 34 Papain is stable and active under a wide range of conditions, namely, from pH 4 to pH 10 and at temperatures up to 80 °C.33, 34 The chemo-enzymatic synthesis of hydrophobic amino acids is essential to design and develop bulk materials from poly(amino acid)s. Papain is capable of polymerizing L-alanine ethyl ester, one of the major components of spider dragline silk, as well as a key amino acid to realize artificial structural poly(amino acid)s by chemo-enzymatic polymerization.35 The reactions carried out at an alkaline pH cause a higher degree of polymerization than those carried out at a neutral pH. Moreover, papain has demonstrated the ability to polymerize alkyl ester of L-tyrosine,36 L-glutamic acid,37 L-lysine,38 L-leucine and L-valine.39 In addition to the synthesis of homopolypeptide, copolymerization of amino acids is first achieved by papain using L-glutamic acid ester and various amino acid esters as co-substrates.40, 41 Interestingly, L-aspartic acid diethyl ester and alanine ethyl ester were not subjected to the papain-catalyzed homopolymerization; rather, they were copolymerized in the presence of L-glutamic acid and papain.42 Diblock and random co-oligopeptides of L-lysine and L-alanine were successfully synthesized by papain.43 Alternating oligopeptides were synthesized by papain-mediated catalysis using the dipeptide monomer, alanine-glycine ethyl ester.44 Papain-catalyzed oligomerization of L-lysine protected at the side chain with a tert-butoxycarbonyl or carboxybenzyl group ensures a higher yield and a simpler purification process owing to the precipitation-driven reaction derived from the hydrophobicity of the protected group.45 Thus, papain is powerful, and the most-studied protease for chemo-enzymatic synthesis of various types of peptides.
As described above, papain, an endo-type cysteine protease, is widely used for chemo-enzymatic polymerization of peptides, whereas serine proteases should be studied as catalysts to understand the reaction mechanism and to develop the chemo-enzymatic synthesis of peptide/poly(amino acid). Proteinase K, originating from Tritirachium album Limber, is a highly active serine protease that shows very broad substrate specificity, allowing the cleavage of peptide bonds next to the carboxyl group of aliphatic and aromatic amino acids as well as polyesters.46 Its hydrolytic activity is maintained up to 60 °C, as well as across a broad pH range from 7.5 to 12.0.46 These stable biochemical properties of proteinase K are ideal for chemo-enzymatic reactions. Until now, proteinase K-catalyzed peptide synthesis has been reported in linear and branched oligo(L-phenylalanine),31 oligo(L-lysine) and oligo(L-arginine), using tris(2-aminoethyl)amine as the branching terminator.47 In addition to cysteine proteases, exo-type proteases are ideal catalysts for polymerization. This is because exo-type protease-mediated hydrolysis proceeds only from the chain ends, which can reduce the hydrolysis as well as enhance aminolysis that results in peptides and not hydrolysis of synthesized peptides.
To synthesize and prepare poly(amino acid)-based materials via chemo-enzymatic polymerization, the author and co-workers focused on the blue mussel (Mytilus edulis) foot protein 5 (Mefp-5), one of the adhesive proteins derived from the mussel, which can be found at the surface of adhesive plaques and is mainly composed of L-glycine, L-lysine and 3,4-dihydroxy- L-phenylalanine (DOPA) (Figure 2b).48, 49 According to the previous studies on recombinant proteins and synthetic polymers containing DOPA, the redox chemistry of DOPA is primarily responsible for adhesion.50, 51, 52 In addition to DOPA, Mefp-5 is composed of L-lysine (approximately 20 mol%), which is necessary to form a network of structures between catechol and amine groups.50, 52 Accordingly, we synthesized adhesive peptide containing DOPA and L-lysine via two enzymatic reactions, namely, chemo-enzymatic synthesis of L-tyrosine and L-lysine by papain as well as tyrosinase-mediated conversion from tyrosine to DOPA.53 On the basis of adhesion tests using the synthesized peptides consisting DOPA, L-lysine and L-tyrosine at various pHs, with different protonation/deprotonation states, we proposed a mechanism whereby deprotonated DOPA can interact with surface materials, thus functioning as an adhesive molecule, while on the other hand, the primary amine group of lysine induces molecular networks under deprotonated conditions. Thus, the combination of multiple enzymatic reactions, including chemo-enzymatic polymerization, increases the efficiency of synthesizing new types of peptide-based materials such as Mefp-5-mimicking adhesive peptides.
Poly(amino acid)/polypeptide as functional materials: target-specific gene carriers
Gene delivery into mammalian cells
As a functional material, poly(amino acid)s including polypeptides and peptides can be applied to various value-added materials, that is, stimuli-responsive, self-assembling, nano-scale and self-healing materials. Another example that demonstrates the advantage of peptides is target-specific delivery of bioactive molecules into plant/animal cells and organelles. Such delivery systems would be even more advantageous if they were biodegradable, biocompatible and mechanically durable, and could be prepared and processed under ambient aqueous conditions to avoid the loss of bioactivity of the molecules to be delivered. Poly(amino acid)s have the potential to meet these demands because their robustness and stability can be controlled by varying their amino-acid sequences and secondary structures, such as β-sheet and alpha-helix structures.54 Currently, chemo-enzymatic synthesis of peptides is not applicable to this type of materials.
Silk, a structural protein, can help address these needs, due to its properties of self-assembly, mechanical toughness, processing flexibility, biodegradability and biocompatibility, and therein presents considerable utility for a number of human therapeutic interventions.55 Spider dragline silk-based block copolymers have been designed and prepared via genetic engineering and used for gene delivery and tissue engineering (Figure 3a).56, 57, 58, 59, 60 The addition of cationic components into silk-based copolymers allows efficient gene delivery due to membrane destabilizing, DNA condensing and pH buffering properties. Molecular transporters such as cell penetrating peptides (CPPs) are 10–50 amino-acid sequences that vary significantly in sequence, electrical charge, hydrophobicity, polarity, and have a remarkable capacity for membrane translocation. Signal peptides and partial sequences of antibody are capable of delivering molecules and chemicals to specific targets. We previously reported various recombinant silk-like peptides containing ligand molecules that facilitated selective delivery to target cells.56, 61, 62, 63, 64, 65, 66 Complexes of recombinant silk molecules containing one of the CPPs, ppTG1 peptide, with plasmid DNA (pDNA) were designed for pDNA delivery.62 The peptide composed of dimeric ppTG1 and a cationic sequence formed a globular pDNA complex of <100 nm in hydrodynamic diameter that was as efficiently transfected as the transfection regent Lipofectamine 2000. Additionally, the silk-based pDNA complexes with greater β-sheet structure demonstrated higher DNase resistance and efficient release of the pDNA by enzymes that degrade the silk proteins. Accordingly, the secondary structure of the peptide in the pDNA complexes is capable of controlling the enzymatic degradation rates of the complexes, and hence can regulate the release profile of genes from the complexes.
A peptide-based gene delivery system for mammalian cells. (a) Schematic of the formation of the pDNA complexes with recombinant silk protein SL-F3, a Histag-silk-poly(L-lysine)-tumor homing peptide F3 block copolymer. (b) In vitro transfection results in the loading of pDNA complexes with recombinant silk proteins and Lipofectamine 2000 in MDA-MB-435, MDA-MB-231 and MCF10A cells. Lipofectamine 2000 was used as a positive control. Data are shown as means±standard deviations (n=4). *A difference between two groups was considered as statistically significant at P<0.05. (c) In vivo transfection results in the loading of pDNA complexes with SL-mF3 in MDA-MB-231 tumor cells in mice (i, ii, iii) and in control experiments without tumor cells (iv, v, vi). Typical bioluminescent images at 3 days (i, iv), 1 week (ii, v) and 4 weeks (iii, vi) of luciferase expression in mice. Figures modified from Numata et al.64 A full color version of this figure is available at Polymer Journal online.
In the case of gene ddelivery for cancer treatment, recombinant silk molecules with tumor-homing peptides have also been prepared to add target specificity into the silk-based gene delivery systems.63, 64 The pDNA complexes of recombinant silk with F3 tumor-homing peptide showed target-specific transfection of tumorigenic cell lines MDA-MB-231 and MDA-MB-435, whereas they did not show useful transfection efficiency in healthy cells, MCF10A (Figure 3b). In vivo transfection experiments were also carried out using the same complex, which showed the highest transfection efficiency in MDA-MB-231 tumor cells. Bioluminescent images of live tumor-bearing mice at 3 days, 1 week and 4 weeks after injection of the pDNA complexes were captured to evaluate the luciferase expression (Figure 3c). The expression level increased slightly for 4 weeks, while the control mice did not show significant expression. More importantly, no significant toxicity was observed in the mice. On the basis of these limited in vivo results, the pDNA complexes containing F3 tumor-homing peptide have the potential to treat tumors specifically, without toxic side effects. Clearly, a more comprehensive study will be required to clarify the potential of this new system and the longevity of the transfection. However, it is also clear that the bioengineered silk-based gene delivery vehicles with functional peptides, such as CPP and the tumor-homing peptides, have the potential for controlled-release and target-specific gene delivery.
Gene delivery into plant cells
Plant gene delivery has been a less studied topic in comparison with gene delivery to treat human diseases. Although several methods to introduce genes into intact plant and cultured plant cells have been reported, each technique has its disadvantages, such as a limitation in the applicable plant types and low transfection efficiency.67 To establish a new gene delivery system for plants, we developed a peptide-based plant gene delivery system, which has ionic complexes of pDNA with designed peptide carriers, each of which combined a CPP with a polycation (Figure 4a).68 The fusion peptides consisting of nona-arginine (R9) and histidine-lysine (KH)9 polycationic sequences as well as CPP such as BP100 and Tat2 were mixed with pDNA encoding Renilla luciferase (RLuc) to form ionic complexes.68 The complexes were injected into leaves of model plants, namely 3-week-old Arabidopsis thaliana and Nicotiana benthamiana, and RLuc gene expression was quantified for up to 144 h. The best complexes had globular shapes with hydrodynamic diameters ranging from 300 to 400 nm and negatively charged surfaces, and showed the highest RLuc activity at 12 h after infiltration to A. thaliana and N. benthamiana for all the fusion peptides. This system, incorporating designed peptides, demonstrated rapid and efficient transfections in intact leaf cells of N. benthamiana and A. thaliana without protoplast preparations.
A peptide-based gene delivery system for plant cells and organelles. (a) The negatively charged pDNA and peptides designed with a polycation and CPP form an ionic complex. The pDNA complex penetrates through the cell wall and the cell membrane, and then the pDNA is expressed throughout the cell, including in the nucleus. (b) A combination of MTP and CPP as a vector for delivery of pDNA into leaves localized to the mitochondria. (c) Confocal laser scanning microscope observation of epidermal cells of a leaf previously infiltrated with the pDNA complexes containing MTP and CPP. Enlarged images of several mitochondria with GFP expression are shown in the extreme right panel. Scale bars indicate 10 μm. (d) Western blot analysis of crude mitochondrial extracts from leaves previously infiltrated with naked pDNA (i), or complexes containing only MTP (ii) or containing CPP and MTP with pDNA (iii). An arrowhead denotes a band corresponding to the molecular weight of GFP. Lane (M) is a molecular weight marker. Figures modified from Lakshmanan et al.68 and Chuah et al.71
In addition to pDNA, other nucleic acids such as double-stranded RNA (dsRNA) and linear DNA can be delivered to the plant cytosol via the peptide-based gene carrier.69 Fast and facile transient RNA interference is one of the most valuable biotechnologies for analyzing plant gene functions. Furthermore, the introduction of RNA without any degradation is an essential technique to realize an efficient genome editing system, such as the CRISPR (clustered regularly interspaced short palindromic repeats) associated Cas9 system. To establish a novel dsRNA delivery system for plants, we developed an ionic complex of synthetic dsRNA with a carrier peptide in which a CPP is fused with a polycation sequence as a gene carrier. Infiltration of the complex into intact leaf cells of A. thaliana as well as the poplar tree successfully induced rapid and efficient downregulation of exogenous and endogenous genes such as yellow fluorescent protein and chalcone synthase.70 This method allowed quick and local gene silencing in specific tissues and/or organs in plants. With this dsRNA delivery system, a new, efficient genome editing system is under investigation and will be reported soon.
Gene delivery to specific organelles
Organelles have important roles in cellular metabolic pathways at various life stages. Specifically, trafficking of biological compounds between organelles is a key mechanism of cellular activity that must be understood to create artificial metabolic pathways. However, the limitation of current transfection techniques for organelles prevents an understanding of the function and role of organelle genes, as well as prevents the use of organelles as a reaction site for producing value-added biomolecules. Major challenges in mitochondrial transfection are the double membrane, small size, and surprisingly motility of mitochondria, in addition to its large population size in cells. To date, no routinely successful method has been developed for introducing foreign genes into the mitochondria of intact, vascular plants. We recently reported the intracellular delivery of exogenous DNA localized to the mitochondria of A. thaliana by a combination of mitochondria-targeting peptide (MTP) and CPP (Figure 4b).71 Low concentrations of peptides were sufficient to deliver pDNA into the mitochondria and expression of imported pDNA reached detectable levels within 12 h. We found that electrostatic interaction with the cell membrane is not a critical factor for complex internalization. Instead, improved intracellular penetration of mitochondria-targeted complexes significantly enhanced gene transfer efficiency. The combination of CPP and MTP mediated significant levels of GFP expression (Figure 4c), as evident from the 27-kDa band corresponding to GFP (Figure 4d). Confocal laser scanning microscopy provided visual confirmation of GFP expression, localized in the mitochondria of epidermal cells of leaves transfected using the peptide–pDNA complex. These results delineate a simple and reproducible peptide-based method as a starting point for the development of more sophisticated and practical plant mitochondrial transfection strategies. The present peptide-based gene delivery system will be a key biotechnology in the production of various bio-fuel and bio-products, as well as functional food and pharmaceutical compounds using plants as a factory.
Poly(amino acid)/polypeptide as structural materials: crystal to micro-structures consisting of structural proteins
Formation of β-sheet crystals with a key role of stabilizing structures
Spider draglines and webs are well-known and typical poly(amino acid)-based structural materials. In the crystalline region of silk fibers, the anti-parallel β-pleated sheet is a fundamental and predominant secondary structure that has a key role in stabilizing these proteins via physical cross-links. In terms of β-sheet structure, silk fibrils were reported to be similar to amyloid fibrils on a molecular level,72, 73 and to enhance amyloidosis of amyloid protein due to cross-seeding effects as a disease mechanism.74 However, silk nanofibrils and nanofilaments composed of β-sheets were reported to show no significant cytotoxicity in neuronal cells in vitro.75 Contrary to amyloid fibrils, silk β-sheets composed of Gly and Ala did not express significant cytotoxicity, most likely because of a lack of electrical charge. Nano-assembly of silk β-sheets produced microfibrils after incubation for 72 h (Figure 5a), and formation of the nano-assembly was faster than that of Aβ(12–28), Aβ(28–42) and full-length Aβ(1–42) because of the greater hydrophobicity of the silk β-sheet.76, 77 The control of silk β-sheet formation is key the assembly needed to create artificial tough polymeric materials; however, this control has not yet been established because β-sheets assemble via hydrophobic interactions too quickly to regulate in solution. Even in solid state, such as the spinning of silk, the control of β-sheet formation remains a challenging component to the design and creation of artificial tough materials composed of poly(amino acid)s.
Structural characteristics of silk fibers and β-sheet crystals. (a) Atomic force microscopy (AFM) height images of nano-assemblies and nanofibrils of spider silk β-sheets before (i) and after incubation for 24 (ii), 48 (iii) and 72 h (iv). The peptides were prepared by solid-phase synthesis. Each scale bar denotes 5 μm. The color scale represents a 100-nm height. (b) The storage modulus (G ′) and the loss modulus (G ″) during gelation of the silk solution (38 g l−1) induced with ethanol were measured at 37 °C. The value of G ′ was under the detection limit of the rheometer below 570 s. (c) Differential interference contrast images of the silk hydrogel with a homogeneous network structure prepared at a silk concentration of 32 g l−1. Scale bars denote 10 μm. (d) Typical AFM amplitude image of silk crystals before the enzymatic degradation. The white broken line denotes nanofibrils composed of the crystals. (e) Line profile data of the crystals are indicated by the white solid line in (d). (f) Model of enzymatic degradation of crystalline regions of silk fibroin. (a) Crystalline region of silk fibroin. Tight and loose chain-packing regions coexist in the crystalline region. (b) Crystalline nanofibril composed of the tight chain-packing regions after degradation of the loose-chain packing region by protease XIV. (c) Nanofilament composed of β-sheets and soluble fragments containing a β-sheet structure after enzymatic degradation by protease XIV. (d) Crystalline region and soluble fragments without β-sheet structure after degradation of the edges and ends of the loose-chain packing region by alpha-chymotrypsin. Figures modified from Numata et al.22, 75, 77 A full color version of this figure is available at Polymer Journal online.
The formation of silk fibrils in concentrated solution results in the gelation of a silk solution with micro-fibrillar network formation. An increase in the β-sheet content, which can be induced by a pH change, sonication, shear stress or hydrophobic interactions, was reported as the major factor causing the gelation of silk solution.78, 79, 80, 81, 82, 83 We also developed a quick and easy method to prepare silk hydrogels using ethanol to induce phase separation, and analyzed the gelation behavior, state of water, secondary structure and mechanical properties of the resulting hydrogel.22 The sample prepared at a silk concentration of 7 g l−1 (silk solution/ethanol=1/9) showed a slight decrease in loss modulus G″ just before the gelation (Figure 5b), which is unusual and distinct from that of synthetic polymers.84 Such a rheological property indicates the aggregation of silk molecules through the formation of β-sheet structures and the subsequent gelation of aggregated silk molecules. On the basis of the structural and morphological characteristics of the silk hydrogels by Fourier transform infrared spectroscopy (FTIR), wide-angle X-ray scattering and differential interference contrast microscopy (Figure 5c), the silk molecules were assembled to form fibrillar and heterogeneous networks with β-sheet structures, which are prerequisites for gelation. Additionally, in differential scanning calorimetry measurements, it is noteworthy that the peaks originating from the bound water seemed to contain two peaks, implying that there might be two types of bound water in the silk hydrogel, namely water bound to micro-scale networks and water intercalated in the β-sheet assembly. The state of water with silk molecules is known to affect the mechanical and biological properties of materials including not only silk but also poly(amino acid)s. The state of water in the silk hydrogel was controllable by adjusting the β-sheet content and network density, which is one of the quantitative studies demonstrating a clear relationship between water and the structural properties of poly(amino acid)s.22
To enhance the mechanical properties of silk hydrogels with physical fibrillar network structures, we hybridized silk fibrils with silk nanoparticles and pectin, that is, a charged hydrophilic polysaccharide.85, 86 The silk-pectin hydrogel is composed of a heterogeneous network, which is different from the fiber-reinforced, interpenetrated networks and double-network hydrogels. A silk concentration of over 15 wt% is critical in constructing interacting silk-pectin networks, which result in strong, elastic hydrogels. Furthermore, the release of water during compression, which was observed in silk-pectin as well as silk hydrogels, is not characteristic of synthetic polymer-based hydrogels, but of tissues found in nature, for example, cartilage and plant cell walls. The silk-pectin hydrogel is therefore considered to have a network structure that contains water molecules and that is different from the network structures of pre-existing synthetic polymer-based hydrogels. The hydrogel of hydrophobic structural proteins demonstrates a different water state in comparison with synthetic polymer-based hydrogels that makes it more difficult to design and create a tough material containing both free and bound water molecules.
Structures revealed by degradation of β-sheet crystals
Contrary to β-sheet formation, an insight into the mechanisms of anti-parallel β-pleated sheet degradation will clarify how the β-sheet crystals influence structural profiles, biological remodeling and physical properties of materials. We have proposed a mechanism for enzymatic degradation of anti-parallel β-pleated sheets, and showed the series of steps involved in digesting Bombyx mori silk structures into fibrils and subsequently into nanofibrils (2 nm thick and 160 nm long). The degradation of the fibril-like polymer crystals by enzymes has been reported in the case of lamellar crystals of poly(hydroxyalkanoic acid).87, 88, 89, 90, 91, 92 In a crystalline biopolyester, tight and loose chain-packing regions form lamellar crystals. The loose chain-packing region is a disordered crystalline region that degrades faster than the highly crystalline domains. In the case of silk β-sheet crystals, we proposed a model of enzymatic degradation of silk crystalline regions due to protease activity based on atomic force microscopy observations, infrared radiation and wide-angle X-ray measurements (Figure 5f). The tight chain-packing regions and looser chain-packing regions exist randomly in the silk crystalline region, and the crystal can be degraded into nanofibrils. This report was the first to observe and recognize nanofibrils around 2 nm thick and 160 nm long, which consisted of a few β-sheet layers according to the crystal structure,93 in the crystalline region of silk fibroin films. The nanofibrils assemble and form fibrils, playing a role as nucleators of the crystalline regions, a novel and important feature of the system that can be exploited to design silk-based structural materials with predictable biodegradability and mechanical properties. This structure–function relationship including the nanofibrils is fundamental in structural material design, where the sizes of crystals can define the mechanical properties of bulk polymers. Thus, the degradation model provides new insights concerning both the fundamental mechanisms of β-sheet structure formation, as well as for the formation of silk-based structural materials.
Summary
Poly(amino acid)s/polypeptides have the potential to accelerate a biomass-based and sustainable society, since these naturally high-performance materials can replace some petroleum-based materials. Biopolymers composed of amino acids have been recognized as bioactive and functional materials according to a large number of studies conducted around the world; however, the use of those biopolymers as structural materials is still challenging. As mentioned in this focus review, one of the major drawbacks of poly(amino acid)-based materials is the limited synthesis methods that are currently available. It is my view that chemo-enzymatic polymerization has the potential to address this problem, but we still need a breakthrough to control the molecular weight and amino-acid sequence of the poly(amino acid)s. Furthermore, to realize poly(amino acid)-based structural materials, we need to understand the solid-state structures of the biopolymers with water molecules, especially during the deformation process. The role of water molecules in the thermal stability, mechanical properties, optical character and biological behavior of structural proteins and poly(amino acid)s is a challenging research target that must also be clarified to use these biopolymers as tough structural materials. We will continue to perform research aimed at clearing these critical challenges to use poly(amino acid)s more widely for the development of bio-materials.
References
Lenz, R. W. & Marchessault, R. H. Bacterial polyesters: biosynthesis, biodegradable plastics and biotechnology. Biomacromolecules 6, 1–8 (2005).
Numata, K., Finne-Wistrand, A., Albertsson, A. C., Doi, Y. & Abe, H. Enzymatic degradation of monolayer for poly(lactide) revealed by real-time atomic force microscopy: effects of stereochemical structure, molecular weight, and molecular branches on hydrolysis rates. Biomacromolecules 9, 2180–2185 (2008).
Numata, K., Srivastava, R. K., Finne-Wistrand, A., Albertsson, A. C., Doi, Y. & Abe, H. Branched poly(lactide) synthesized by enzymatic polymerization: Effects of molecular branches and stereochernistry on enzymatic degradation and alkaline hydrolysis. Biomacromolecules 8, 3115–3125 (2007).
Chuah, J. A., Yamada, M., Taguchi, S., Sudesh, K., Doi, Y. & Numata, K. Biosynthesis and characterization of polyhydroxyalkanoate containing 5-hydroxyvalerate units: effects of 5HV units on biodegradability, cytotoxicity, mechanical and thermal properties. Polym. Degrad. Stab 98, 331–338 (2013).
Insomphun, C., Mifune, J., Orita, I., Numata, K., Nakamura, S. & Fukui, T. Modification of beta-oxidation pathway in Ralstonia eutropha for production of poly(3-hydroxybutyrate-co-3-hydroxyhexanoate) from soybean oil. J. Biosci. Bioeng. 117, 184–190 (2014).
Numata, K. & Doi, Y. Biosynthesis of polyhydroxyalkanaotes by a novel facultatively anaerobic Vibrio sp. under marine conditions. Mar. Biotechnol 14, 323–331 (2012).
Numata, K., Morisaki, K., Tomizawa, S., Ohtani, M., Demura, T., Miyazaki, M., Nogi, Y., Deguchi, S. & Doi, Y. Synthesis of poly- and oligo(hydroxyalkanoate)s by deep-sea bacteria, Colwellia spp., Moritella spp., and Shewanella spp. Polym. J. 45, 1094–1100 (2013).
Numata, K., Motoda, Y., Watanabe, S., Tochio, N., Kigawa, T. & Doi, Y. Active intermediates of polyhydroxyalkanoate synthase from Aeromonas caviae in polymerization reaction. Biomacromolecules 13, 3450–3455 (2012).
Tsuge, T., Yano, K., Imazu, S., Numata, K., Kikkawa, Y., Abe, H., Taguchi, S. & Doi, Y. Biosynthesis of polyhydroxyalkanoate (PHA) copolymer from fructose using wild-type and laboratory-evolved PHA synthases. Macromol. Biosci. 5, 112–117 (2005).
Ushimaru, K., Motoda, Y., Numata, K. & Tsuge, T. Phasin proteins activate Aeromonas caviae polyhydroxyalkanoate (PHA) synthase but not Ralstonia eutropha PHA synthase. Appl. Environ. Microbiol. 80, 2867–2873 (2014).
Yamada, M., Takahashi, S., Okahata, Y., Doi, Y. & Numata, K. Monitoring and kinetic analysis of the molecular interactions by which a repressor protein, PhaR, binds to target DNAs and poly[(R)-3-hydroxybutyrate]. AMB Express 3, 6 (2013).
Chuah, J.-A., Tomizawa, S., Yamada, M., Tsuge, T., Doi, Y., Sudesh, K. & Numata, K. Characterization of site-specific mutations in a short-chain-length/medium-chain-length polyhydroxyalkanoate synthase: in vivo and in vitro studies of enzymatic activity and substrate specificity. Appl. Environ. Microbiol. 79, 3813–3821 (2013).
Numata, K. & Morisaki, K. Screening of marine bacteria to synthesize polyhydroxyalkanoate from lignin: contribution of lignin derivatives to biosynthesis by Oceanimonas doudoroffii. ACS Sustain. Chem. Eng. 3, 569–573 (2015).
Numata, K., Motoda, Y., Watanabe, S., Osanai, T. & Kigawa, T. Co-expression of two polyhydroxyalkanoate synthase subunits from Synechocystis sp. PCC 6803 by cell-free synthesis and their specific activity for polymerization of 3-Hydroxybutyryl-Coenzyme A. Biochemistry 54, 1401–1407 (2015).
Osanai, T., Numata, K., Oikawa, A., Kuwahara, A., Iijima, H., Doi, Y., Tanaka, K., Saito, K. & Yokota Hirai, M. Increased bioplastic production with an RNA polymerase sigma factor SigE during nitrogen starvation in Synechocystis sp PCC 6803. DNA Res. 20, 525–535 (2013).
Osanai, T., Oikawa, A., Numata, K., Kuwahara, A., Iijima, H., Doi, Y., Tanaka, K., Saito, K. & Yokota Hirai, M. Pathway-level acceleration of glycogen catabolism by a response regulator in the Cyanobacterium Synechocystis Species PCC 6803. Plant Physiol. 164, 1831–1841 (2014).
Tomizawa, S., Chuah, J. A., Matsumoto, K., Doi, Y. & Numata, K. Understanding the limitations in the biosynthesis of polyhydroxyalkanoate (PHA) from lignin derivativesACS Sustain. Chem. Eng. 2, 1106–1113 (2014).
Numata, K., Abe, H. & Doi, Y. Enzymatic processes for biodegradation of poly(hydroxyalkanoate)s crystals. Can. J. Chem. 86, 471–483 (2008).
Numata, K., Abe, H. & Iwata, T. Biodegradability of poly(hydroxyalkanoate) materials. Materials 2, 1104–1126 (2009).
Tachibana, K., Urano, Y. & Numata, K. Biodegradability of nylon 4 film in a marine environment. Polym. Degrad. Stab. 98, 1847–1851 (2013).
Vandermeulen, G. W. M. & Klok, H.-A. Peptide/protein hybrid materials: enhanced control of structure and improved performance through conjugation of biological and synthetic polymers. Macromol. Biosci. 4, 383–398 (2004).
Numata, K., Katashima, T. & Sakai, T. State of water, molecular structure, and cytotoxicity of silk hydrogels. Biomacromolecules 12, 2137–2144 (2011).
Arnau, J., Lauritzen, C. & Pedersen, J. Cloning strategy, production and purification of proteins with exopeptidase-cleavable His-tags. Nat. Protoc. 1, 2326–2333 (2006).
Merrifield, R. B. Solid-phase peptide synthesis. Adv. Enzymol. Relat. Areas Mol. Biol. 32, 221–296 (1969).
Shen, Y., Fu, X. H., Fu, W. X. & Li, Z. B. Biodegradable stimuli-responsive polypeptide materials prepared by ring opening polymerization. Chem. Soc. Rev. 44, 612–622 (2015).
Yazawa, K. & Numata, K. Recent advances in chemoenzymatic peptide syntheses. Molecules 19, 13755–13774 (2014).
Tabata, K., Ikeda, H. & Hashimoto, S. ywfE in Bacillus subtilis codes for a novel enzyme, L-amino acid ligase. J. Bacteriol. 187, 5195–5202 (2005).
Kullmann, W. Proteases as catalysts for enzymic syntheses of opioid peptides. J. Biol. Chem. 255, 8234–8238 (1980).
Overberger, C. G., Stpierre, T., Vorchhei., N & Yaroslavsky, S. Esterolytic catalysis of poly-4(5)-vinylimidazole and poly-5(6)-vinylbenzimidazole. J. Am. Chem. Soc. 85, 3513–3515 (1963).
Wong, Y. M., Hoshino, Y., Sudesh, K., Miura, Y. & Numata, K. Optimization of poly(N-isopropylacrylamide) as an artificial amidase. Biomacromolecules 16, 411–421 (2015).
Ageitos, J. M., Baker, P. J., Sugahara, M. & Numata, K. Proteinase K-catalyzed synthesis of linear and star oligo(L-phenylalanine) conjugates. Biomacromolecules 14, 3635–3642 (2013).
Chaiken, I. M., Komoriya, A., Ohno, M. & Widmer, F. Use of enzymes in peptide-synthesis. Appl. Biochem. Biotech. 7, 385–399 (1982).
Kamphuis, I. G., Drenth, J. & Baker, E. N. Thiol proteases. Comparative studies based on the high-resolution structures of papain and actinidin, and on amino acid sequence information for cathepsins B and H, and stem bromelain. J. Mol. Biol. 182, 317–329 (1985).
Berger, A. & Schechter, I. Mapping the active site of papain with the aid of peptide substrates and inhibitors. Philos. Trans. R. Soc. Lond. B Biol. Sci. 257, 249–264 (1970).
Baker, P. J. & Numata, K. Chemoenzymatic synthesis of poly(L-alanine) in aqueous environment. Biomacromolecules 13, 947–951 (2012).
Fukuoka, T., Tachibana, Y., Tonami, H., Uyama, H. & Kobayashi, S. Enzymatic polymerization of tyrosine derivatives. Peroxidase- and protease-catalyzed synthesis of poly(tyrosine)s with different structures. Biomacromolecules 3, 768–774 (2002).
Li, G., Vaidya, A., Viswanathan, K., Cui, J., Xie, W., Gao, W. & Gross, R. A. Rapid regioselective oligomerization of L-glutamic acid diethyl ester catalyzed by papain. Macromolecules 39, 7915–7921 (2006).
Qin, X., Xie, W. C., Su, Q., Du, W. Z. & Gross, R. A. Protease-catalyzed oligomerization of L-lysine ethyl ester in aqueous solution. ACS Catal. 1, 1022–1034 (2011).
Baker, P. J., Patwardhan, S. V. & Numata, K. Synthesis of homopolypeptides by aminolysis mediated by proteases encapsulated in silica nanospheres. Macromol. Biosci. 14, 1619–1626 (2014).
Uyama, H., Fukuoka, T., Komatsu, I., Watanabe, T. & Kobayashi, S. Protease-catalyzed regioselective polymerization and copolymerization of glutamic acid diethyl ester. Biomacromolecules 3, 318–323 (2002).
Li, G., Raman, V. K., Xie, W. C. & Gross, R. A. Protease-catalyzed co-oligomerizations of L-leucine ethyl ester with L-glutamic acid diethyl ester: sequence and chain length distributions. Macromolecules 41, 7003–7012 (2008).
Viswanathan, K., Schofield, M. H., Teraoka, I. & Gross, R. A. Surprising metal binding properties of phytochelatin-like peptides prepared by protease-catalysis. Green Chem. 14, 1020–1029 (2012).
Fagerland, J., Finne-Wistrand, A. & Numata, K. Short one-pot chemo-enzymatic synthesis of L-lysine and L-alanine diblock co-oligopeptides. Biomacromolecules 15, 735–743 (2014).
Qin, X., Khuong, A. C., Yu, Z., Du, W., Decatur, J. & Gross, R. A. Simplifying alternating peptide synthesis by protease-catalyzed dipeptide oligomerization. Chem. Commun. 49, 385–387 (2013).
Qin, X., Xie, W., Tian, S., Ali, M. A., Shirke, A. & Gross, R. A. Influence of N-epsilon-protecting groups on the protease-catalyzed oligomerization of L-lysine methyl ester. ACS Catal. 4, 1783–1792 (2014).
Ebeling, W., Hennrich, N., Klockow, M., Metz, H., Orth, H. D. & Lang, H. Proteinase K from Tritirachium-Album Limber. Eur. J. Biochem. 47, 91–97 (1974).
Ageitos, J. M., Chuah, J. A. & Numata, K. Chemo-enzymatic synthesis of linear and branched cationic peptides: evaluation as gene carriers. Macromol. Biosci. (e-pub ahead of print 31 March 2015; doi:10.1002/mabi.201400487)
Harrington, M. J., Masic, A., Holten-Andersen, N., Waite, J. H. & Fratzl, P. Iron-clad fibers: a metal-based biological strategy for hard flexible coatings. Science 328, 216–220 (2010).
Silverman, H. G. & Roberto, F. F. Understanding marine mussel adhesion. Mar. Biotechnol. 9, 661–681 (2007).
Burzio, L. A. & Waite, J. H. Cross-linking in adhesive quinoproteins: studies with model decapeptides. Biochemistry 39, 11147–11153 (2000).
Burzio, L. A. & Waite, J. H. The other Topa: formation of 3,4,5-trihydroxyphenylalanine in peptides. Anal. Biochem. 306, 108–114 (2002).
Waite, J. H. & Qin, X. Polyphosphoprotein from the adhesive pads of Mytilus edulis. Biochemistry 40, 2887–2893 (2001).
Numata, K. & Baker, P. J. Synthesis of adhesive peptides similar to those found in blue mussel (Mytilus edulis) using papain and tyrosinase. Biomacromolecules 15, 3206–3212 (2014).
Wadhwa, M. S., Collard, W. T., Adami, R. C., McKenzie, D. L. & Rice, K. G. Peptide-mediated gene delivery: influence of peptide structure on gene expression. Bioconjug. Chem. 8, 81–88 (1997).
Altman, G. H., Diaz, F., Jakuba, C., Calabro, T., Horan, R. L., Chen, J., Lu, H., Richmond, J. & Kaplan, D. L. Silk-based biomaterials. Biomaterials 24, 401–416 (2003).
Numata, K. & Kaplan, D. L. Silk-based delivery systems of bioactive molecules. Adv. Drug Deliv. Rev. 62, 1497–1508 (2010).
Gomes, S., Numata, K., Leonor, I. B., Mano, J. F., Reis, R. L. & Kaplan, D. L. AFM study of morphology and mechanical properties of a chimeric spider silk and bone sialoprotein protein for bone regeneration. Biomacromolecules 12, 1675–1685 (2011).
Qin, G., Lapidot, S., Numata, K., Hu, X., Meirovitch, S., Dekel, M., Podoler, I., Shoseyov, O. & Kaplan, D. L. Expression, cross-linking, and characterization of recombinant chitin binding resilin. Biomacromolecules 10, 3227–3234 (2009).
Rajkhowa, R., Gil, E. S., Kluge, J., Numata, K., Wang, L., Wang, X. & Kapaln, D. L. Reinforcing silk scaffolds with silk particles. Macromol. Biosci. 10, 599–611 (2010).
Sengupta, S., Park, S.-H., Seok, G. E., Patel, A., Numata, K., Lu, C.-L. & Kaplan, D. L. Quantifying osteogenic cell degradation of silk biomaterials. Biomacromolecules 11, 3592–3599 (2010).
Numata, K., Hamasaki, J., Subramanian, B. & Kaplan, D. L. Gene delivery mediated by recombinant silk proteins containing cationic and cell binding motifs. J. Control. Release 146, 136–143 (2010).
Numata, K. & Kaplan, D. L. Silk-based gene carriers with cell membrane destabilizing peptides. Biomacromolecules 11, 3189–3195 (2010).
Numata, K., Mieszawska-Czajkowska, A. J., Kvenvold, L. A. & Kaplan, D. L. Silk-based nanocomplexes with tumor-homing peptides for tumor-specific gene delivery. Macromol. Biosci. 12, 75–82 (2012).
Numata, K., Reagan, M. R., Goldstein, R. H., Rosenblatt, M. & Kaplan, D. L. Spider silk-based gene carriers for tumor cell-specific delivery. Bioconjug. Chem 22, 1605–1610 (2011).
Numata, K., Subramanian, B., Currie, H. A. & Kaplan, D. L. Bioengineered silk protein-based gene delivery systems. Biomaterials 30, 5775–5784 (2009).
Numata, K. & Doi, Y. Dual biosyntheses of poly[(R)-3-hydroxybutyric acid] and silk protein for the fabrication of biofunctional bioplastic. Polym. J. 43, 642–647 (2011).
Chilton, M. D. Adding diversity to plant transformation. Nat. Biotechnol. 23, 309–310 (2005).
Lakshmanan, M., Kodama, Y., Yoshizumi, T., Sudesh, K. & Numata, K. Rapid and efficient gene delivery into plant cells using designed peptide carriers. Biomacromolecules 14, 10–16 (2013).
Lakshmanan, M., Yoshizumi, T., Sudesh, K., Kodama, Y. & Numata, K. Double-stranded DNA introduction into intact plants using peptide–DNA complexes. Plant Biotechnol. 32, 39–45 (2015).
Numata, K., Ohtani, M., Yoshizumi, T., Demura, T. & Kodama, Y. Local gene silencing in plants via synthetic dsRNA and carrier peptide. Plant Biotechnol. J. 12, 1027–1034 (2014).
Chuah, J. A., Yoshizumi, T., Kodama, Y. & Numata, K. Gene introduction into the mitochondria of Arabidopsis thaliana via peptide-based carriers. Sci. Rep. 5, 7751 (2015).
Kenney, J. M., Knight, D., Wise, M. J. & Vollrath, F. Amyloidogenic nature of spider silk. Eur. J. Biochem. 269, 4159–4163 (2002).
Slotta, U., Hess, S., Spieß, K., Stromer, T., Serpell, L. & Scheibel, T. Spider silk and amyloid fibrils: a structural comparison. Macromol. Biosci. 7, 183–188 (2007).
Lundmark, K., Westermark, G. T., Olsen, A. & Westermark, P. Protein fibrils in nature can enhance amyloid protein A amyloidosis in mice: Cross-seeding as a disease mechanism. Proc. Natl. Acad. Sci. USA 102, 6098–6102 (2005).
Numata, K., Cebe, P. & Kaplan, D. L. Mechanism of enzymatic degradation of beta-sheet crystals. Biomaterials 31, 2926–2933 (2010).
Numata, K. & Kaplan, D. L. Mechanisms of enzymatic degradation of amyloid beta microfibrils generating nanofilaments and nanospheres related to cytotoxicity. Biochemistry 49, 3254–3260 (2010).
Numata, K. & Kaplan, D. L. Differences in cytotoxicity of beta-sheet peptides originated from silk and amyloid beta. Macromol. Biosci. 11, 60–64 (2011).
Ayub, Z. H., Arai, M. & Hirabayashi, K. Mechanism of the gelation of fibroin solution. Biosci. Biotechnol. Biochem. 57, 1910–1912 (1993).
Ayub, Z. H., Arai, M. & Hirabayashi, K. Quantitative structural-analysis and physical-properties of silk fibroin hydrogels. Polymer 35, 2197–2200 (1994).
Kim, U. J., Park, J., Li, C., Jin, H. J., Valluzzzi, R. & Kaplan, D. L. Structure and properties of silk hydrogels. Biomacromolecules 5, 786–792 (2004).
Matsumoto, A., Chen, J., Collette, A. L., Kim, U.-J., Altman, G. H., Cebe, P. & Kaplan, D. L. Mechanisms of silk fibroin sol-gel transitions. J. Phys. Chem. B 110, 21630–21638 (2006).
Wang, X. Q., Kluge, J. A., Leisk, G. G. & Kaplan, D. L. Sonication-induced gelation of silk fibroin for cell encapsulation. Biomaterials 29, 1054–1064 (2008).
Yucel, T., Cebe, P. & Kaplan, D. L. Vortex-induced injectable silk fibroin hydrogels. Biophys. J. 97, 2044–2050 (2009).
Kurakazu, M., Katashima, T., Chijiishi, M., Nishi, K., Akagi, Y., Matsunaga, T., Shibayama, M., Chung, U. & Sakai, T. Evaluation of gelation kinetics of tetra-PEG gel. Macromolecules 43, 3935–3940 (2010).
Numata, K., Yamazaki, S., Katashima, T., Chuah, J.-A., Naga, N. & Sakai, T. Silk-pectin hydrogel with superior mechanical properties, biodegradability, and biocompatibility. Macromol. Biosci. 14, 799–806 (2014).
Numata, K., Yamazaki, S. & Naga, N. Biocompatible and biodegradable dual-drug release system based on silk hydrogel containing silk nanoparticles. Biomacromolecules 13, 1383–1389 (2012).
Kikkawa, Y., Hirota, T., Numata, K., Tsuge, T., Abe, H. & Doi, Y. In-situ atomic force microscopy observation of enzymatic degradation in poly(hydroxyalkanoic acid) thin films: Normal and constrained conditions. Macromol. Biosci. 4, 276–285 (2004).
Numata, K., Hirota, T., Kikkawa, Y., Tsuge, T., Iwata, T., Abe, H. & Doi, Y. Enzymatic degradation processes of lamellar crystals in thin films for poly[(R)-3-hydroxybutyric acid] and its copolymers revealed by real-time atomic force microscopy. Biomacromolecules 5, 2186–2194 (2004).
Numata, K., Kikkawa, Y., Tsuge, T., Iwata, T., Doi, Y. & Abe, H. Enzymatic degradation processes of poly[(R)-3-hydroxybutyric acid] and poly[(R)-3-hydroxybutyric acid-co-(R)-3-hydroxyvaleric acid] single crystals revealed by atomic force microscopy: Effects of molecular weight and second-monomer composition on erosion rates. Biomacromolecules 6, 2008–2016 (2005).
Numata, K., Kikkawa, Y., Tsuge, T., Iwata, T., Doi, Y. & Abe, H. Adsorption of biopolyester depolymerase on silicon wafer and poly[(R)-3-hydroxybutyric acid] single crystal revealed by real-time AFM. Macromol. Biosci. 6, 41–50 (2006).
Numata, K., Sato., S., Fujita, M., Tsuge, T., Iwata, T., Doi, Y. & Abe, H. Adsorption effects of poly (hydroxybutyric acid) depolymerase on chain-folding surface of polyester single crystals revealed by mutant enzyme and frictional force microscopy. Polym. Degrad. Stab. 92, 176–183 (2007).
Numata, K., Yamashita, K., Fujita, M., Tsuge, T., Kasuya, K., Iwata, T., Doi, Y. & Abe, H. Adsorption and hydrolysis reactions of poly(hydroxybutyric acid) depolymerases secreted from Ralstonia pickettii T1 and Penicillium funiculosum onto poly[(R)-3-hydroxybutyric acid]. Biomacromolecules 8, 2276–2281 (2007).
Takahashi, Y., Gehoh, M. & Yuzuriha, K. Structure refinement and diffuse streak scattering of silk (Bombyx mori). Int. J. Biol. Macromol. 24, 127–138 (1999).
Acknowledgements
This work was supported by a grant for the RIKEN Biomass Engineering Program, Impulsing Paradigm Change through Disruptive Technologies Program (ImPACT) from the Japan Science and Technology Corporation (JST), Cross-Ministerial Strategic Innovation Promotion Program (SIP) from National Agriculture and Food Research Organization, New Energy and Industrial Technology Development Organization (NEDO), the Sumitomo Foundation, two Grants-in-Aid for Young Scientists (B), a Grant-in-Aid for JSPS Fellows for Research Abroad and a Grant-in-Aid for JSPS Fellows from the Japan Society for the Promotion of Science (JSPS). We expresses special thanks to Professor Emeritus Yoshiharu Doi from Tokyo Institute of Technology, Stern Family Professor of Engineering David L Kaplan from Tufts University, and all the lab members of the Enzyme Research Team in RIKEN Center for Sustainable Resource Science.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no conflict of interest.
Rights and permissions
About this article
Cite this article
Numata, K. Poly(amino acid)s/polypeptides as potential functional and structural materials. Polym J 47, 537–545 (2015). https://doi.org/10.1038/pj.2015.35
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pj.2015.35
This article is cited by
-
Exploring Microbial Contributions to Nutraceutical Production: From Natural to Designed Foods
Molecular Biotechnology (2023)
-
Ameliorative effect of calcium poly(aspartic acid) (PASP-Ca) and calcium poly-γ-glutamic acid (γ-PGA-Ca) on soil acidity in different horizons
Environmental Science and Pollution Research (2023)
-
Structural bases for aspartate recognition and polymerization efficiency of cyanobacterial cyanophycin synthetase
Nature Communications (2022)
-
Preparation and pH/temperature dual drug release behavior of polyamino acid nanomicelles
Polymer Bulletin (2022)
-
Characterization of an l-α,β-diaminopropionic acid polymer with comb-like structure isolated from a poly(ε-l-lysine)-producing Streptomyces sp.
Applied Microbiology and Biotechnology (2021)