Abstract
Applications of natural viruses to nanotechnology have attracted much attention due to their fascinating nanostructures. However, chemical strategy to de novo design virus-like nanostructures is still in its infancy. In this review article, we describe novel C3-symmetric molecular design of peptides and their characteristic self-assembly into virus-like nanostructures. We have developed trigonal conjugates of β-sheet-forming peptides, trigonal conjugates of glutathione and a viral β-annulus peptide fragment.
Similar content being viewed by others
Introduction
In biological systems, bottom-up construction of nano-sized functional materials has been carried out by molecular self-assembly. Many kinds of supramolecular nanostructures, as exemplified by photosynthetic reaction centers, cell membranes, chromatins, ion channels, flagella, actin filaments, microtubules, viruses and clathrin, are spontaneously formed by the self-assembly of biomolecules such as proteins, lipids and nucleic acids. Development of artificial nanomaterials self-organized from biomolecular building blocks are of interest because of their unique morphology, fascinating functionality, diversity of molecular design and high biocompatibility.1, 2, 3 Since Seeman pioneered the new scientific field of ‘DNA nanotechnology’ in 1991, various DNA nano-architectures such as polyhedrons, spheres and sheets, which are self-assembled from elaborately programmed DNA building units bearing junction structures, have been developed.4, 5, 6, 7, 8, 9‘DNA origami’ structures have been constructed by folding natural long single-strands of DNA with programmed short DNA building units.10, 11 These DNA superstructures have been used as nano-actuators and templates for the arrangement of functional molecules and nanoparticles.
Viruses are also one of the natural supramolecular assemblies with discrete size and specific aggregation number, which possess rod-like or spherical morphology.12 Recently, the application of bacteriophages, such as M13 phage,13 and plant viruses,14, 15 such as cowpea mosaic virus and cowpea chlorotic mottle virus, to nanotechnology have attracted much attention due to their fascinating nanostructures. For example, Douglas and Young16 reported mineralization of polyoxometalates (paratungstate and decavanadate) and single crystal γ-FeOOH17 in the nano-space inside a cowpea chlorotic mottle virus capsid. Furthermore, Cornelissen and co-workers18 succeeded in inclusion of horse radish peroxidase and green fluorescence protein19 into the modified cowpea chlorotic mottle virus capsid. In addition, Finn and co-workers20 have developed functional CPMV capsids modified with oligonucleotides, gold nanoparticles21 and oligosaccharides22 onto the surface. The virus nanotechnologies depend on the structure of ‘ready-made’ capsids; however, chemical strategy to de novo design ‘tailor-made’ virus-like nanocapsule is still in its infancy. Development of designed capsid molecules for reconstructing viral architectures would enhance the potential of virus-like nanocapsules and notably contribute to advances in nano-bioscience.
In this focus review article, we describe our approach to development of virus-inspired peptide nanoassemblies. Moreover, novel C3-symmetric molecular design of peptides and their characteristic self-assembling behavior are described.
Self-assembly of spherical viruses and virus-inspired artificial molecules
In the past decades, three-dimensional structures of many simple spherical viruses have been revealed by X-ray crystallography.12 Most capsids in spherical viruses are formed by the assembly of multiples of 60 proteins, having an icosahedral symmetry (including C5-, C3-, and C2-symmetric axes) as a result. The number and arrangement of subunits are related to the triangulation number (T number) that is derived from quasi-equivalence theory, meaning the number of equilateral triangles composed of protein trimers on a face of icosahedron. For example, satellite tobacco necrosis virus (T=1, diameter: 18 nm), the smallest virus discovered, is self-assembled from 60 equivalent protein subunits. Tomato bushy stunt virus (T=3, diameter: 33 nm) is generated from 180 quasi-equivalent protein subunits, in which the dodecahedral internal skeleton is composed of a C3-symmetric β-annulus motif.23 Such spherical protein assemblies from C3-symmetric subunits are also found in non-viral systems. Clathrin lattice, which participates in receptor-mediated endocytosis, is aggregated from the C3-symmetric peptide triskerions in the presence of a magnesium ion.24, 25 Ferritin, which is an intracellulariron storage protein, has C3-symmetric pores.26
Chemists have developed many organic nanocapsules inspired by spherical viruses and clathrin. Rebek and colleague27 developed molecular-sized nanocapsules from the self-assembly of two bent molecules via hydrogen bonding. Fujita and co-workers28, 29 devised self-assembled coordination cages with a size of 3–5 nm. However, the size of these supramolecular nanocapsules are evidently smaller than that of natural viruses, and consequently their applications would be limited by the inclusion of small guest molecules.
Trigonal conjugates of β-sheet-forming peptides
To date, artificial supramolecular assemblies consisting of amphiphilic α-helix peptides,30, 31, 32 coiled-coil α-helix peptides33, 34, 35 and β-sheet-forming peptides36, 37 have been reported. As mentioned above, many biological supramolecules often have highly symmetric units to achieve effective assembly. The symmetric pre-organization of peptide chains might reduce the entropy loss in the self-assembling process and afford unique assemblies in water. Yeates and co-workers38 have prepared a fusion protein of bromoperoxidase (trimer-forming unit) together with M1 matrix protein of influenza virus (dimer-forming unit), which self-assembled into tetrahedral cages with a size of 15 nm. We have applied the C3-symmetric molecular design inspired by spherical viruses and clathrin to synthetic peptide assemblies.
In 2005, we reported the first example of artificial C3-symmetric peptide conjugates (Trigonal-(FKFE)2) that self-assembled into viral-sized peptide nanospheres in water (Figure 1).39 Trigonal-(FKFE)2 conjugate was prepared by coupling thiol groups of β-sheet-forming peptides (CFKFEFKFE) with iodoacethylated trimesoyl core molecule. Circular dichroism and Fourier transform infrared spectra of acidic solutions (pH 3.3) of Trigonal-(FKFE)2 revealed the formation of anti-parallel β-sheet structure. Scanning electron microscopy of Trigonal-(FKFE)2 showed the presence of spherical assemblies with a size of 22–34 nm. The average diameter of the nanospheres, as determined by dynamic light scattering, was 19.1±4.0 nm, which is comparable with the estimated diameter of a dodecahedron structure (about 16 nm). It is noteworthy that nanospheres are selectively generated without the formation of fibrous assemblies.
In contrast, a trigonal peptide conjugate consisting of three β-sheet-forming peptides (FKFECKFE) with wheel-like arrangement (Wheel-FKFE) self-assembled into nanofibers with uniform width (Figure 2).40 It seems that the peptide units of Wheel-FKFEs in nanofibers form anti-parallel β-sheets with attractive intermolecular ionic complementarity. If Trigonal-(FKFE)2s form nanofibers, they would have aligned to form parallel β-sheets against the repulsive ionic complementarity and led to the formation of nanofibers of Trigonal-(FKFE)2.
Self-assembly of trigonal peptide conjugate consisting of three β-sheet-forming peptides with wheel-like arrangement (wheel-FKFE) into nanofibers with uniform width in water. Adapted with permission from Matsuura et al.40 Copyright 2008 American Chemical Society. A full color version of this figure is available at Polymer Journal online.
Tryptophane zipper is a secondary structure motif that forms a stable twisted β-hairpin structure due to the interaction between tryptophane residues.41, 42C3-symmetric peptide conjugate (Trigonal-WTW) that bears three tryptophane zipper-forming peptides (CKTWTWTE) showed pH-responding self-assembly in water, namely, nanospheres (about 30 nm) via formation of tryptophane zipper at pH 7, mixture of nanofibers and nanospheres at pH 11 and irregular aggregates at pH 3 (Figure 3).43 It is believed that Trigonal-WTW formed nanospheres by forming anti-parallel β-sheet-like structures due to the attractive ionic complementarity at pH 7. Recently, we have also reported syntheses of pentagonal conjugates of tryptophane zipper-forming peptides (CKTWTWTE), and nanofibers were obtained at pH 7.44 The difference in morphology between trigonal- and pentagonal-tryptophane zipper conjugates might arise from difference in the peripheral density of peptide chains and the conformation of scaffold.
pH-responding self-assembly of C3-symmetric peptide conjugate bearing tryptophane zipper-forming peptides (Trigonal-WTW) in water. Adapted with permission from Matsuura et al.43 Copyright 2011 Royal Society of Chemistry. A full color version of this figure is available at Polymer Journal online.
Trigonal conjugates of glutathione
Recently, it was reported that simple dipeptide and tripeptide derivatives act as self-assembling units. For example, Gazit and others45, 46 found that aromatic dipeptide (Phe-Phe) formed crystalline nanotube assemblies in water. Atkins et al.47 reported that oxidized glutathione (GSSG) formed organogel in various organic solvents, whereas fluorescence hydrogel was generated from glutathione-pyrene conjugate in water.48
We have developed a trigonal glutathione (TG) conjugate that spontaneously aggregated into nanospheres with a size of 100–250 nm in water (Figure 4a).49 Although the size and morphology of the assemblies were minimally affected by the concentration and pH, the nanospheres underwent structural changes from hollow to filled spheres depending on the concentration. Uranine (a guest molecule) was encapsulated into nanospheres that were formed from TG at lower concentrations (<1 mm), while it became hardly bound at higher TG concentration (>1 mm). By adopting a similar strategy, Gazit and co-workers50 reported that C3-symmetric dipeptide WW conjugate self-assembled into nanocapsules with a size of 200–500 nm in methanol–water mixtures.
Self-assembly of trigonal-glutathione (a) and conformation-regulated trigonal glutathione (b) into nanospheres in water. Adapted with permission from Matsuura et al.49, 52 Copyright 2009 Royal Society of Chemistry and 2010 Chemical Society of Japan. A full color version of this figure is available at Polymer Journal online.
TG nanospheres underwent disulfide recombination upon the addition of dithiothreitol, which was confirmed by reverse-phase high-performance liquid chromatography and matrix assisted laser desorption ionization time-of-flight mass spectrometry analyses. Interestingly, although the spherical morphology was maintained, the guest molecules encapsulated inside the TG nanospheres were gradually released during the disulfide recombination (Figure 5).51 This controlled release system, based on bond recombination of nanoassemblies, would provide a novel guideline for design of a drug-delivery system.
Schematic illustration of disulfide recombination in spherical assembly of Trigonal glutathione (TG). Adapted with permission from Matsuura et al.51 Copyright 2011 Chemical Society of Japan. DTT, dithiothreitol
In order to construct more regular nanoassemblies from glutathione derivatives, 1,3,5-tris(aminomethyl)-2,4,6-triethylbenzene was used as the core of a new TG.52 The glutathione arms on the 1,3,5-positions were assembled on the one side of the scaffold due to the alternative conformation of the core.53 The conformation-regulated trigonal glutathione was combined into hard spherical assemblies with a size of 310±50 nm (Figure 4b). The spherical superstructures self-assembled from conformation-regulated trigonal glutathione showed regular morphology and enhanced rigidity compared with those formed from conformationally non-regulated TG.
Virus-like nanocapsules self-assembled from viral β-annulus peptide fragment
Tomato bushy stunt virus capsid consists of 180 quasi-equivalent protein subunits that comprise 388 amino acids each.23 If virus-like nanocapsules can be constructed from peptide fragments containing a subunit of viral capsid, the peptide fragments would be promising candidates as components of chemically designable artificial nanoassemblies.
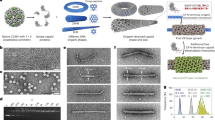
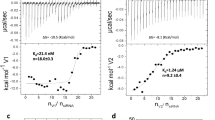
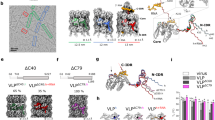
Recently, we found that 24-mer β-annulus peptides (INHVGGTGGAIMAPVAVTRQLVGS) of tomato bushy stunt virus, which participates in the formation of dodecahedral internal skeleton, spontaneously self-assembled into hollow nanocapsules with a size of 30–50 nm in water(Figure 6).54 The nanocapsule possesses a definite critical aggregation concentration (CAC=25 μm), and its size is almost unaffected by peptide concentration above CAC. The hollow structure of the present assembly from β-annulus peptide was clearly revealed by synchrotron small angle X-ray scattering. These findings provide a minimized design of artificial viral capsids that can be modified to extend the molecular design of functional nanocapsules, such as DNA and protein carriers, and the platform for artificial vaccine by proper surface modifications.
Self-assembly of 24-mer β-annulus peptide fragment from tomato bushy stunt virus into virus-like nanocapsule with a size of 30–50 nm in water. The red amino-acid sequences should form β-structure and intermolecular hydrogen bonds. Adapted with permission from Matsuura et al.54 Copyright 2010 Wiley-VCH.
Conclusion
We have provided a novel molecular design for the construction of peptide-based nanoassemblies. The building of artificial viral capsids from designed peptide conjugates and peptide fragments is epoch-making discovery, which will open a field of innovation in virus-based nanotechnology. The peptide-based artificial viral capsids can be dressed-up by proper modification on the surface or in the interior to make functional nanocapsules. More elaborate molecular design might make it possible to construct more complicated biomolecular assemblies such as adenovirus and T4 bacteriophage. Subsequent tasks toward artificial living system include giving special functions, such as actuator, self-replication and information transmission, to the artificial biomolecular assemblies.
References
Niemeyer, C. M. Nanoparticles, Proteins, and Nucleic Acids: Biotechnology Meets Materials Science. Angew. Chem. Int. Ed. 40, 4128–4158 (2001).
Niemeyer, C. M. & Mirkin, C. A. (Eds.) Nanobiotechnology, (Wiley-VCH, Weinheim 2004).
Mirkin, C. A. & Niemeyer, C. M. (Eds.) Nanobiotechnology II, (Wiley-VCH, Weinheim 2007).
Seeman, N. C Nucleic acid nanostructures and topology. Angew. Chem. Int. Ed. 37, 3220–3238 (1998).
Seeman, N. C. DNA in a material world. Nature 421, 427–431 (2003).
Shih, W. M., Quispe, J. D. & Joyce, G. F. A 1.7-kilobase single-stranded DNA that folds into a nanoscale octahedron. Nature 427, 618–621 (2004).
Goodman, R. P., Schaap, I. A. T., Tardin, C. F., Erben, C. M., Berry, R. M., Schmidt, C. F. & Turberfield, A. J. Rapid chiral assembly of rigid DNA building blocks for molecular nanofabrication. Science 310, 1661–1665 (2005).
Matsuura, K., Yamashita, T., Igami, Y. & Kimizuka, N. ‘Nucleo-nanocages’: designed ternary oligodeoxyribonucleotides spontaneously form nanosized DNA cages. Chem. Commun. 376–377 (2003).
Matsuura, K., Masumoto, K., Igami, Y., Fujioka, T. & Kimizuka, N. In situ observation of spherical DNA assembly in water and the controlled release of bound dyes. Biomacromolecules 8, 2726–2732 (2007).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Nangreave, J., Han, D., Liu, Y. & Yan, H. DNA origami: a history and current perspective. Curr. Opin. Chem. Biol. 14, 608–615 (2010).
Branden, C. & Tooze, J. Introduction to Protein Structure 2nd ed. (Garland Publishing, New York 1999).
Mao, C., Solis, D. J., Reiss, B. D., Kottmann, S. T., Sweeney, R. Y., Hayhurst, A., Georgiou, G., Iverson, B. & Belcher, A. M. Virus-based toolkit for the directed synthesis of magnetic and semiconducting nanowires. Science 303, 213–217 (2004).
Douglas, T. & Young, M. Viruses: making friends with old foes. Science 312, 873–875 (2006).
Steinmetz, N. F. & Evans, D. J. Utilisation of plant viruses in bionanotechnology. Org. Biomol. Chem. 5, 2891–2902 (2007).
Douglas, T. & Young, M. Host–guest encapsulation of materials by assembled virus protein cages. Nature 393, 152–155 (1998).
Douglas, T., Strable, E., Willits, D., Aitouchen, A., Libera, M. & Young, M. Protein engineering of a viral cage for constrained nanomaterials synthesis. Adv. Mater. 14, 415–418 (2002).
Comellas-Aragonès, M., Engelkamp, H., Claessen, V. I., Sommerdijik, N. A. J. M., Rowan, A. E., Christianen, P. C. M., Maan, J. C., Verduin, B. J. M., Cornelissen, J. J. L. M. & Nolte, R. J. M. A virus-based single-enzyme nanoreactor. Nat. Nanotech. 2, 635–639 (2007).
Minten, I. J., Hendriks, L. J. A., Nolte, R. J. M. & Cornelissen, J. J. L. M. Controlled encapsulation of multiple proteins in virus capsids. J. Am. Chem. Soc. 131, 17771–17773 (2009).
Strable, E., Johnson, J. E. & Finn, M. G. Natural nanochemical building blocks: icosahedral virus particles organized by attached oligonucleotides. Nano Lett. 4, 1385–1389 (2004).
Wang, Q., Lin, T., Tang, L., Johnson, J. E. & Finn, M. G Icosahedral virus particles as addressable nanoscale building blocks. Angew. Chem. Int. Ed. 41, 459–462 (2002).
Kaltgrad, E., O’Reilly, M. K., Liao, L., Han, S., Paulson, J. C. & Finn, M. G. On-virus construction of polyvalent glycan ligands for cell-surface receptors. J. Am. Chem. Soc. 130, 4578–4579 (2008).
Olson, J., Bricogne, G. & Harrison, S. C. Structure of tomato bushy stunt virus IV. The virus particle at 2.9 Å resolution. J. Mol. Biol. 171, 61–93 (1983).
Harrison, S. C. & Kirchhausen, T. Clathrin, cages, and coated vesicles. Cell 33, 650–652 (1983).
Fotin, A., Cheng, Y., Sliz, P., Grigorieff, N., Harrison, S. C., Kirchhausen, T. & Walz, T. Molecular model for a complete clathrin lattice from electron cyomicroscopy. Nature 432, 573–579 (2004).
Theil, E. C. Ferritin: Structure, gene regulation, and cellular function in animals, plants and microorganisms. Annu. Rev. Biochem. 56, 289–315 (1987).
Conn, M. M. & Rebek, J. Self-assembling capsules. Chem. Rev. 97, 1647–1668 (1997).
Tominaga, M., Suzuki, K., Kawano, M., Kusukawa, T., Ozeki, T., Sakamoto, S., Yamaguchi, K. & Fujita, M. Finite, spherical coordination networks that self-organize from 36 small components. Angew. Chem. Int. Ed. 43, 5621–5625 (2004).
Sun, Q.-F., Iwasa, J., Ogawa, D., Ishido, Y., Sato, S., Ozeki, T., Sei, Y., Yamaguchi, K. & Fujita, M. Self-assembled M24L48 polyhedra and their sharp structural switch upon subtle ligand variation. Science 328, 1144–1147 (2011).
Fujita, K., Kimura, S. & Imanishi, Y. Spherical self-assembly of a synthetic α-helical peptide in water. Langmuir 15, 4377–4379 (1999).
Kimura, S., Kim, D.-H., Sugiyama, J. & Imanishi, Y. Vesicular self-assembly of a helical peptide in water. Langmuir 15, 4461–4463 (1999).
Morikawa, M., Yoshihara, M., Endo, T. & Kimizuka, N. α-Helical polypeptide microcapsules formed by emulsion-templated self-assembly. Chem. Eur. J. 11, 1574–1578 (2005).
Ryadnov, M. G. & Woolfson, D. N. Engineering the morphology of a self-assembling protein fibre. Nat. Mater. 2, 329–332 (2003).
Ryadnov, M. G. & Woolfson, D. N. Introducing branches into a self-assembling peptide fiber. Angew. Chem. Int. Ed. 42, 3021–3023 (2003).
Zhou, M., Bentley, D. & Ghosh, I. Helical supramolecules and fibers utilizing leucine zipper-displaying dendrimers. J. Am. Chem. Soc. 126, 734–735 (2004).
Marini, D. M., Hwang, W., Lauffenburger, D. A., Zhang, S. & Kamm, R. D. Left-handed helical ribbon intermediates in the self-assembly of a β-sheet peptide. Nano Lett. 2, 295–299 (2002).
Yokoi, H., Kinoshita, T. & Zhang, S. Dynamic reassembly of peptide RADA16 nanofiber scaffold. Proc. Natl. Acad. Sci. USA 102, 8414–8419 (2005).
Padilla, J. E., Colovos, C. & Yeates, T. O. Nanohedra: using symmetry to design self-assembling protein cages, layers, crystals, and filaments. Proc. Natl. Acad. Sci. USA 98, 2217–2221 (2001).
Matsuura, K., Murasato, K. & Kimizuka, N. Artificial peptide-nanospheres self-assembled from three-way junctions of β-sheet-forming peptides. J. Am. Chem. Soc. 127, 10148–10149 (2005).
Murasato, K., Matsuura, K. & Kimizuka, N. Self-assembly of nanofiber with uniform width from wheel-type Trigonal-β-Sheet forming peptide. Biomacromolecules 9, 913–918 (2008).
Cochran, A. G., Skelton, N. J. & Starovasnik, M. A. Tryptophan zippers: stable, monomeric β-hairpins. Proc. Natl. Acad. Sci. USA 98, 5578–5583 (2001).
Streicher, W. W. & Makhatadze, G. I. Calorimetric evidence for a two-state unfolding of the β-hairpin peptide Trpzip4. J. Am. Chem. Soc. 128, 30–31 (2006).
Matsuura, K., Hayashi, H., Murasato, K. & Kimizuka, N. Trigonal tryptophane-zipper as a novel building block for pH-responding peptide nano-assemblies. Chem. Commun. 47, 265–267 (2011).
Matsuura, K., Murasato, K. & Kimizuka, N. Syntheses and self-assembling behaviors of pentagonal conjugates of tryptophane zipper-forming peptide. Int. J. Mol. Sci. 12, 5187–5199 (2011).
Reches, M. & Gazit, E. Casting metal nanowires within discrete self-assembled peptide nanotubes. Science 300, 625–627 (2003).
Gazit, E. Self-assembled peptide nanostructures: the design of molecular building blocks and their technological utilization. Chem. Soc. Rev. 36, 1263–1269 (2007).
Lyon, R. P. & Atkins, W. M. Self-assembly and gelation of oxidized glutathione in organic solvents. J. Am. Chem. Soc. 123, 4408–4413 (2001).
Mahajan, S. S., Paranji, R., Mehta, R., Lyon, R. P. & Atkins, W. M. A Glutathione-Based hydrogel and its site-selective interactions with water. Bioconj. Chem. 16, 1019–1026 (2005).
Matsuura, K., Matsuyama, H., Fukuda, T., Teramoto, T., Watanabe, K., Murasato, K. & Kimizuka, N. Spontaneous Self-assembly of nano-spheres from trigonal conjugate of glutathione in water. Soft Matter. 5, 2463–2470 (2009).
Ghosh, S., Reches, M., Gazit, E. & Verma, S. Bioinspired design of nanocages by self-assembling triskelion peptide elements. Angew. Chem. Int. Ed. 46, 2002–2004 (2007).
Matsuura, K., Tochio, K., Watanabe, K. & Kimizuka, N. Controlled release of guest molecules from spherical assembly of trigonal gultathione by disulfide recombination. Chem. Lett. 40, 711–713 (2011).
Matsuura, K., Fujino, K., Teramoto, T., Murasato, K. & Kimizuka, N. Glutathione nanospheres: self-assembly of conformation-regulated trigonal-glutathiones in water. Bull. Chem. Soc. Jpn. 83, 880–886 (2010).
Wiskur, S. L., Ait-Haddou, H., Lavigne, J. J. & Anslyn, E. V Teaching old indicators new tricks. Acc. Chem. Res. 34, 963–972 (2001).
Matsuura, K., Watanabe, K., Sakurai, K., Matsuzaki, T. & Kimizuka, N. Self-assembled synthetic viral capsids from a 24-mer viral peptide fragment. Angew. Chem. Int. Ed. 49, 9662–9665 (2010).
Acknowledgements
This research was partially supported by a Grant-in-Aid for Scientific Research on the Innovative Areas of ‘Fusion Materials’ (no. 2206) from the Ministry of Education, Science, Sports and Culture of Japan (MEXT), and by a Grant-in-Aid for Scientific Research (B) (no. 22350075) from Japan Society for the Promotion of Science. Synchrotron small angle X-ray scattering experiment was performed by Professor Salurai and Mr Matsuzaki (The University of Kitakyushu) on SPring-8 BL40B2 and BL45XU. I am deeply indebted to Professor Nobuo Kimizuka (Kyushu University) for his continuous encouragement and discussion throughout this work. Most of this study was conducted by our students. I am very much indebted to many of my co-workers and students. I thank to Dr Joseph Hui (Kyushu University) for reading this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Matsuura, K. Construction of spherical virus-inspired peptide nanoassemblies. Polym J 44, 469–474 (2012). https://doi.org/10.1038/pj.2012.16
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pj.2012.16
Keywords
This article is cited by
-
Continual reproduction of self-assembling oligotriazole peptide nanomaterials
Nature Communications (2017)
-
Antimicrobial peptide capsids of de novo design
Nature Communications (2017)
-
Self-assembled artificial viral capsid decorated with gold nanoparticles
Polymer Journal (2015)
-
Guest-binding behavior of peptide nanocapsules self-assembled from viral peptide fragments
Polymer Journal (2013)