Abstract
Background:
Extracellular vesicles (EVs) are a heterogeneous class of lipid bound particles shed by any cell in the body in physiological and pathological conditions. EVs play critical functions in intercellular communication. EVs can actively travel in intercellular matrices and eventually reach the circulation. They can also be released directly in biological fluids where they appear to be stable. Because the molecular content of EVs reflects the composition of the cell of origin, they have recently emerged as a promising source of biomarkers in a number of diseases. EV analysis is particularly attractive in cancer patients that frequently present with increased numbers of circulating EVs.
Methods:
We sought to review the current literature on the molecular profile of prostate cancer-derived EVs in model systems and patient biological fluids in an attempt to draw some practical and universal conclusions on the use of EVs as a tool for liquid biopsy in clinical specimens.
Results:
We discuss advantages and limitations of EV-based liquid biopsy approaches summarizing salient studies on protein, DNA and RNA. Several candidate biomarkers have been identified so far but these results are difficult to apply to the clinic. However, the field is rapidly moving toward the implementation of novel tools to isolate cancer-specific EVs that are free of benign EVs and extra-vesicular contaminants. This can be achieved by identifying markers that are exquisitely present in tumor cell-derived EVs. An important contribution might also derive from a better understanding of EV types that may play specific functions in tumor progression and that may be a source of cancer-specific markers.
Conclusions:
EV analysis holds strong promises for the development of non-invasive biomarkers in patients with prostate cancer. Implementation of modern methods for EV isolation and characterization will enable to interrogate circulating EVs in vivo.
Similar content being viewed by others
Introduction
Prostate cancer (PC) is the most common non-skin cancer in males. Thanks to significant advances in the diagnosis and treatment, the 10-year cancer-specific survival of PC patients exceeds 98%1 enabling many men to live with, rather than die of their disease. In spite of this, more than 180 000 new cases are estimated in 2016 and over 26 000 men are expected to die of PC in the same year.2 Consequently, PC remains the second leading cause of cancer deaths in the United States. Among the key clinical challenges are the inability to accurately distinguish low-risk (indolent) from high-risk (aggressive) disease at the time of diagnosis. In addition, the highly variable clinical course has limited the capacity to predict the biological progression of the disease. Within the existing risk stratification systems for PC, histological grading using the Gleason score system3 remains one of the best independent predictors of clinical outcome. This system was recently modified based on new 2016 World Health Organization PC reporting guidelines.4, 5 The five proposed risk groups (PGG1–PGG5) correspond to genomic groups with increasing number of somatic copy number aberrations.6 Metastatic PC exhibits a distinct signature with specific driving genomic alterations that might provide personalized therapeutic targets.6 Indeed, different genomic signatures might characterize diverse facets of the same lethal disease, and this molecular heterogeneity, along with potentially actionable molecular defects has been identified not only in castration resistance (CR) disease but also in naive PC.7 Identifying alterations that predict high-risk disease, metastasis and/or CR would improve the management of the patients and their long-term outcome.
Although fine-needle biopsies are the standard for PC diagnosis and prognosis, their ability to identify high-risk disease is limited8 Moreover, their invasive nature causes significant morbidity. Analysis of PSA in blood has long been used for early diagnosis and for monitoring of biochemical recurrence, but is flawed with a significant number of false positives.9 These limitations drive ongoing attempts at developing minimally invasive procedures to interrogate cancer-derived macromolecules in circulating tumor cells (CTCs), cell-free DNA (cfDNA) and extracellular vesicles (EVs).
Circulating tumor cells
Despite the intrinsic limitation that CTCs are rare,10 analysis of CTCs has allowed a non-invasive, real-time molecular characterization of cancer in patients with metastatic disease11, 12 and has successfully been used to predict response to Docetaxel and Prednisone in patients with metastatic castration resistant PC (mCRPC).13 Additionally, nuclear expression of the androgen receptor variant V7 (AR-V7) in CTCs seems to report response to taxanes.14 Recently, CTCs have been shown to be heterogeneous and to present characteristics of neuroendocrine differentiation, suggesting that broader criteria for definition and inclusion of CTCs should be used.15
Cell-free DNA
Tumor-derived cfDNA is particularly attractive due to its abundance in plasma. Profiling cfDNA by next-generation sequencing has recently resulted in a strategy for monitoring tumor dynamics and identifying genomic causes of resistance, suggesting a temporal association between cfDNA profiles and clinical progression.16 Similar studies have enabled accurate estimations of absolute androgen receptor (AR) copy number17 and such an analysis of plasma AR in CRPC patients has identified primary resistance to Abiraterone.17 These studies rely on sophisticated computational tools to limit the possibility of missing circulating AR genomic lesions caused by low input cfDNA, which is also contaminated with benign cell-derived DNA.16, 17 Mounting evidence indicates that cfDNA is not the only source of extracellular DNA in plasma, and that a portion of this DNA along with other cancer-derived molecules is carried and protected in EVs.
Extracellular vesicles
EVs are lipid-enclosed particles released by all cellular organisms and containing a repertoire of macromolecules (cargo) that is reflective of the cell of origin. The EV cargo is stable because it is protected from enzymatic degradation by a lipid membrane envelope. EVs are extremely abundant (~1–3 × 1012 exosomes per ml of plasma.18 Direct enumeration of tumor-derived EVs and/or profiling of their molecular cargo in patient body fluids have been shown to provide valuable information about the biology of the tumor.19 While EV profiling offers significant advantages because they contain a complex cargo (proteins, lipids, nucleic acids) that could be interrogated simultaneously, discriminating cancer-derived EVs from the non-cancer ones has been challenging.
EVs are heterogeneous
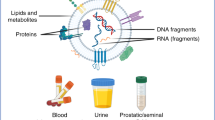
Various populations of EVs have been described.20, 21 While exosomes and ectosomes are the most frequently studied EVs, biofluids contain highly heterogeneous EV populations, of variable size and originating by diverse mechanisms (Figure 1). Importantly, different EV types exhibit distinct molecular and functional characteristics.22, 23 Also, benign cells produce various types of EVs.24 Thus cancer-derived EVs float in a sea of physiologically normal EVs and the use of EVs in liquid biopsies relies on improving the methodologies to selectively purify single EV populations based on their subcellular origin, size, cargo, and donor cell type and status.
The cartoon represents two canonical mechanisms of EV formation from multivesicular bodies (Exo) or direct budding from the plasma membrane, as it is the case for ectosomes (MV and LO). Notably, MV and LO can reach significantly larger dimensions than Exo. EV, extracellular vesicles; LO, large oncosomes; MV, microvesicles.
Size–nomenclature–biogenesis–origin
Exosomes
Exosomes are nanosized vesicles (50 to >100 nm in diameter)22 that originate from the fusion of multivesicular bodies with the plasma membrane (PM),25 or bud directly off the PM26 as recently reviewed.20, 21 Any type of cell from any living organism can shed exosomes.27 Prostate-specific EVs termed ‘prostasomes’ were originally described as exosome-like structures that are released by normal prostate epithelial cells in the seminal fluid.28 Functionally, it is known that prostasomes mediate intercellular communication between epithelial secretory cells and sperm cells. More specifically, secretory cells seem to use this mechanism to nurture sperm cells, thus increasing their motility.29 The term prostasomes has often been used interchangeably with exosomes due to their high cholesterol/phospholipids ratio and to a protein pattern that suggests the origin of prostasomes from multivesicular bodies. Tavoosidana et al.30 reported that detection of prostasomes in peripheral blood is useful for early diagnosis and prognosis in organ-confined PC. However, recent studies have clarified that, unless these EVs are identified in the seminal fluid, the term exosomes is more appropriate to describe them.31
Ectosomes
The term ectosomes indicates exquisitely EVs originating by direct budding and pinching off of the PM.32 These vesicles are generally larger than exosomes (250–1000 nm) and result from PM budding as a consequence of activation of GTPases.32, 33 Ectosomes are also frequently referred to as ‘microvesicles’ (MV). MV were originally described as platelet-derived EVs with a functional role in coagulation34 but several new types of bioactive vesicles derived from a variety of cells, including tumor cells, have been described.
Large oncosomes
Large oncosomes (LO) are atypically large EVs (1–10 μm diameter) that are released by tumor cells transitioning from a mesenchymal to a more rapid, amoeboid motility mode, which results in highly migratory, invasive, and metastatic phenotypes.35, 36, 37, 38 This process occurs in cells with active Akt1 and EGFR signaling,35 and can be induced by silencing of the actin nucleator diaphanous-related formin 3 (DIAPH3).23 They do not appear to be produced by benign cells.39 More recently, the identification of similarly large EVs has been reported21, 40, 41 but it is unclear whether they share similar functional or molecular characteristics to the LO.
EVs as a source of biomarkers in PC
While a discrete portion of molecules is enriched or depleted in EVs in comparison with the cancer cell of origin, suggesting a finely regulated mechanism of export,42, 43 the EV cargo largely reflects the donor cell cargo. Because these molecules are protected from degradation and can be measured in biological fluids, they represent a valuable, non-invasive source of tumor-derived markers, as demonstrated in a large number of neoplasms (Table 1).44, 45, 46, 47, 48, 49
Mass spectrometry
Protein profiling in plasma, serum and other biological fluids are limited by an intrinsic high dynamic range, and by the fact that most of the FDA-approved serum biomarkers are at least five orders of magnitude lower than albumin or other abundant serum proteins. For this reason, most of the preliminary studies on the characterization of the protein cargo of PC-derived EVs have been performed on EVs obtained from PC cell lines. This approach has created an invaluable resource of potential biomarkers and has circumvented the limitations deriving from a direct interrogation of body fluids.
One of the first deep mass spectrometry (MS) analyses of EVs was performed by the Llorente group50 in exosomes derived from PC cells metastatic to the bone.51 In addition to several proteins with extracellular functions, cytoskeleton formation and EV biogenesis, the authors identified CD151 and CUB domain-containing protein 1 (CDCP1) in cancer-EVs but not in benign EVs. CD151 is a tetraspanin family member involved in tumor progression,52, 53 while CDCP1 is a transmembrane glycoprotein identified as a metastasis-associated protein,54 and expressed at high levels in human tumors.55, 56 CDCP1 was also detected in EVs from invasive PC cells with characteristics of epithelial to mesenchymal transition57 but not from isogenic, less aggressive cells.58 Additionally, identification of CDCP1 in urine samples by an antibody specific to the ectodomain of the protein discriminated patients with high-risk versus low-risk PC.58 These results demonstrate that cell-surface proteins can be used to identify tumor-derived EVs and support the use of CDCP1-positive EVs as circulating markers of PC. CDCP1 might also itself prove to be a biomarker.
In order to improve reproducibility and sensitivity of protein detection, Webber et al.59 applied a novel MS platform to exosomes and donor DU145 PC cells. The resulting SOMAscan platform uses a class of protein-binding reagents called SOMAmers (slow off-rate modified aptamers) that allows the detection of a significant number of proteins and can handle a large number of samples. Milk fat globule-EGF factor 8 (MFG-E8), Notch3, disintegrin and metalloproteinase domain-containing protein (ADAM9) were enriched in exosomes in comparison to the tumor cells. Not only did this study contribute to the identification of novel putative extracellular markers of PC, but it also led to the discovery that proteins that are typically considered to be secreted in a free form, are actually exported in EVs. The authors further demonstrated by sucrose density gradient that the soluble form of transforming growth factor β1 (TGF-β1) does not co-isolate with exosome-bound TGF-β1 and they concluded that this is the case also for granulocyte-colony stimulating factor, vascular endothelial growth factor, and interleukin 8, confirming the importance to analyze soluble molecules separately from EV-bound molecules.
A MS approach was also employed by Hosseini-Beheshti et al.60 to profile exosome proteins from AR positive and negative PC cell lines. A number of candidate biomarkers, including calsyntenin-1 (CLSTN1), growth differentiation factor 15 (GDF15), fatty acid synthase (FASN) and clusterin (CLU) were identified in EVs from these tumor cells but not from non-tumorigenic prostate epithelial cells. Finally, Duijvesz et al.61 identified higher levels of programmed cell death 6-interacting protein (PDCD6IP/Alix), FASN, CD9 and enolase 1 (ENO1) in exosomes derived from malignant prostate cell versus benign cell lines.
The Jenster group performed MS on sera of mice with androgen-naive or androgen resistant PC thus providing one of the first evidences for in vivo significance of EVs as biomarkers in this disease.19 They compared the sera profiles to the profiles of exosomes and exosome-deprived supernatant of PC cells to test whether the proteins released in the serum are likely to be packaged in EVs. Strikingly, most of the proteins identified in mouse sera and in PC cell-derived exosomes, including glyceraldehyde 3-phosphate dehydrogenase (GAPDH), ENO1 and lactate dehydrogenase B (LDHB), were found only in the exosome fraction, and not in the exosome-depleted supernatant, suggesting they are exported in EVs. Identification of TM256 by Øverbeye et al.62 in urine exosomes discriminated PC patients (n=16) from controls (n=15) with 94% sensitivity. The sensitivity reached 100% when TM256 was analyzed in combination with the autophagy inhibitor late endosomal/lysosomal adaptor, MAPK and MTOR activator 1 (LAMTOR1). These proteins are candidate EV urinary markers, and the validity of this discovery is supported by the identification, in the same study, of well-known PC markers such as PSA, prostate-specific membrane antigen (FOLH1/PMSA), transglutaminase (TGM4) and transmembrane protease, serine 2 (TMPRSS).
Exosome proteins have also been interrogated for identification of biomarkers of treatment response and particularly for early detection of drug resistance. Kharaziha et al.63 provided evidence that exosomes report acquisition of resistance to docetaxel. In comparison with parental DU145 PC cells, which are sensitive to docetaxel, DU145 cells that have acquired resistance to the compound (Tax-Res) shed more abundant exosomes as measured by NTA. Additionally, exosomes derived from Tax-Res cells carry a different set of proteins while expressing similar levels of exosome markers. Examples are the poly(A)-binding protein (PABP4), Endophilin-A2 and the ATP-dependent drug efflux pumps MDR-1 (or p-glycoprotein) and MDR-3, which were enriched in the Tax-Res exosomes. These observations were further confirmed in a small cohort of patients with CRPC.
Regarding the identification of canonical PC biomarkers in exosomes, an analysis of the literature demonstrates that they contain the mRNA product of the TMPRSS2-ERG fusion as well as PSA.62 However, some studies report detection of PSA in exosome-deprived supernatant rather than as true exosome cargo.19, 61 It would be interesting to investigate whether EV-associated PSA has different function and provides different information than soluble PSA. Additional potential markers are δ-catenin, which was identified in EVs from PC3 cell media and in urine samples of patients with PC,64 and survivin, which was expressed at higher levels in circulating EVs from cancer patients in comparison with healthy controls or patients with BPH.65
As mentioned before, EVs other than exosomes and MV exist. One example is represented by LO, which are particularly large EVs that have not been identified in normal prostate tissue, are present at low levels in tissue of patients with low Gleason Score, organ-confined cancer and at high levels in mCRPC.39 Caveolin-1 (Cav-1), which has been functionally implicated in PC progression66, 67, 68 and has been suggested as a circulating marker of metastatic PC by the Thompson group,69 emerged as a promising LO-associated protein detectable in the circulation of patients with metastatic disease but not in patients with organ-confined PC.70 Importantly, Cav-1 expression was also identified in exosomes, confirming the data from other groups.71 However, flow cytometry experiments indicated that Cav-1-positive LO, but not exosomes, are significantly increased in patients with metastasis.70 One putative marker of LO is cytokeratin 18 (CK18), a protein that is almost completely excluded from exosomes, as demonstrated by gradient centrifugation.23 The presence of CK18-positive EVs in human plasma was associated with cancer in a small cohort of PC patients.23 Additional LO markers are HSPA5, HSPD1, GAPDH and other functionally interesting proteins, including membrane proteins and metabolic enzymes that seem to be implicated in the biological functions of LO.23 Interrogating larger cohorts of patients to validate clinical significance of these markers is essential. Because of the tumor-specificity, differentiating LO from other EV types may increase the granularity of cancer-specific signal. Given their large size, LO could also be easily captured and analyzed by NGS, thus informing the genetic composition of the tumor, risk of recurrence, and disease-free or overall survival. Interestingly, the identification of very large objects (sometimes as large as CTCs) positive for epithelial cell adhesion molecule (EpCAM) and CK18 in the circulation of PC patients was reported in 2010 by Coumans et al.72 and was correlated with reduced survival. Unfortunately, the system currently approved by the FDA uses particularly strict parameters to define a CTC, therefore these abnormally large vesicular structures, which could be LO, are discarded and not incorporated in the analysis. Developing or repurposing microfluidic devices to capture and analyze, specifically, tumor-derived LO, might clarify the function and clinical significance of these vesicles.
In summary, several studies on protein cargo have identified potential EV biomarkers. For most of these, it is unclear whether they are specific to a given EV population (Figure 2). Confirming these data in larger cohorts of patients, and determining whether one or more of these markers are associated with specific EV populations will provide significant advantages to the development of EV-based non-invasive tools for personalized medicine.
Extracellular nucleic acids
A few reports on DNA, and several reports on RNA in EVs have been recently published.73, 74 The evidence that nucleic acids travel protected in EVs offers generous opportunities to nominate tumor type or disease-stage-specific variants or other genomic alterations, as well as aberrant transcripts or noncoding RNAs in EVs by massive parallel sequencing.
DNA
In 2011, Balaj et al.75 were the first to demonstrate that EVs contain DNA. A few years later, the presence of double-stranded DNA (dsDNA), reflecting the genomic DNA (gDNA) of the originating cell, was reported and low-coverage whole-genome sequencing demonstrated that the DNA contained in EVs spans all chromosomes.73, 74 However, single primer PCR was necessary to detect specific DNA aberrations in plasma EVs. Thus, KRAS and P53 mutations, which typically occur in more than 90% of patients with pancreatic cancer,76, 77 were identified in exosomes from pancreatic cancer cells,73 and BRAF and EGFR mutations were identified in exosomes from melanoma and lung cancer cells.74 These seminal findings suggest that more sensitive methods for discovery will improve detection of EVs in which the tumor-derived DNA is highly concentrated. An interesting report from Lázaro-Ibáñez et al. recently tested whether different populations of EVs (exosomes, MV and apoptotic bodies) contain extracellular gDNA with different profiles, and PC-specific mutations.78 Additional studies are needed to understand the functional distribution of extracellular DNA in different EV populations that might accommodate different amounts ad types of DNA.
In order to render this effort clinically applicable, it will be essential to test the presence of frequent mutations in specific clinical groups, standardizing the methods and preserving the EV cargo. For example, because genomic alterations of the phosphoinositide 3-kinase (PI3K) axis, which includes PTEN mutations and deletions, have been identified in tumor tissues from 50% of patients with CRPC by whole-exome and RNA seq,79 this group of patients could be an ideal target to measure PTEN copy number as a EV-based circulating DNA assays. Collaborations with other fields will also help standardization of EV profiling. For example, it is still unclear what the best way to preserve EVs is and perhaps some insights can be derived from the CTC community. Similarly, the cfDNA field might inspire future development of EV assays, since they have established the computational pipelines to analyze highly fragmented plasma DNA. It is important to note that the size of DNA fragments in LO is significantly larger than in other types of EVs (unpublished observations), supporting further investigations in those EVs. We speculate that specific selection of these vesicles might improve designing the approach to detect tumor signal in complex biofluids, thus allowing to monitor the tumor genomic make up by liquid biopsy.
RNA
Donovan et al. recently developed an assay that reports a patient score (EXO106) based on mRNA levels of PC antigen 3 (PCA-3), ETS-related gene (ERG) and SAM pointed domain-containing Ets transcription factor (SPDEF) in urine-derived exosomes.80 In receiver operating characteristic (ROC) analysis, the EXO106 score predicted both PC and high-grade disease (Gleason score 7 or higher) with an area under the curve (AUC) of 0.764. Importantly, EXO106 showed its usefulness in the diagnosis of high-grade PC in patients in ‘grey zone’ serum PSA levels (>2 and <10 ng ml−1). In a follow-up study, the gene expression assay score of PCA-3, ERG and SPDEF assayed in urine exosomes (ExoDx Prostate IntelliScore) outperformed standard of care values. This combination resulted to be more predictive in discriminating PC patients with ⩾GS7 from GS6 or with negative biopsy results (AUC 0.73) in multiple cohorts obtained from different sites. The addition of the ExoDx Prostate IntelliScore seems also to prove useful in identifying patients who are eligible to biopsy among the ones with elevated PSA levels.81 Nilsson et al. had previously examined PSA, PCA-3 and TMPRSS:ERG mRNA levels in urine-derived exosomes and demonstrated a correlation between these transcripts and high Gleason score as well as poor response to androgen deprivation therapy.82 Even if on a small sample size, the results of this study are remarkable and support investigations of the prognostic role of exosome-enclosed mRNA in larger cohorts of patients. Additional studies from Dijkstra et al. and Hendriks et al. have tested the suitability of using urine-derived exosomes for PC diagnosis. However, while the first study suggests the clinical usefulness of urine-derived exosomes, the second proposes whole urine as substrate for biomarkers analysis.83, 84 Of note, both studies highlight the positive effect of digital rectal examination prior urine sampling since it seems to increase material recovering.
In addition to identifying known PC biomarkers, recent studies have attempted to perform extensive, unbiased screenings through array-based approaches to identify novel PC RNA biomarkers (Figure 2). These studies focus particularly on miRNAs, which are stable in human biological fluids because resistant to ribonuclease degradation.85 Korzeniewski et al. reported three miRNAs (miR-483-5p, miR-1275, miR-1290) as the most abundant miRNAs released from PC cells.86 When tested in urine samples, miR-483-5p alone as well as in combination with the other two miRNAs, was able to significantly differentiate PC patients with biopsy-proven tumor mass from patients with microscopic tumor. However, PSA was a better disease predictor, and overall, these miRNAs failed to show advantages over conventional methods. Corcoran et al.87 examined the potential use of exosomal miR-34a as a predictive biomarker of response to docetaxel. Among the miRNAs whose levels correlated with disease progression (miR-598, miR-34a, miR-146a, miR-148a), miR-34a showed the highest clinical relevance based on in silico analysis of publicly available datasets. α-synuclein and solute carrier family 7 member 5 (SLC7A5), which are two common targets of these four miRNAs, also correlated with prognosis in CRPC patients. These results collectively suggest that analyses of EV-bound miRNA might be more useful than analyses of whole miRNA fraction in biofluids (obtained without purifying EVs). However, comparative studies are necessary to make this conclusion.
A large-scale screening was performed by Huang et al.,88 who reported that miR-1290 and miR-375 can be used as prognostic markers of CRPC. Rather than using array-based approach, RNA sequencing with short RNA libraries was conducted in 23 patients with CRPC to identify survival-related miRNAs in plasma exosomes. We believe this is the first RNA sequencing screening performed on plasma EVs in CRPC. Candidate miRNAs were further validated for their prognostic values using PCR in 100 patients. Plasma exosomal miR-1290 and miR-375 were chosen as the two most promising prognostic biomarkers. Interestingly, while miR-375 upregulation in tumor tissues correlated with PC progression,89 miR-1290 was not significantly upregulated in PC tissues.90 This is an interesting result suggesting that EV analyses could allow identification of cancer-derived molecules in biological fluids even if their levels are not increased in tumor tissue. Last but not least, little is known about the potential for use of exosomal long noncoding RNAs (lncRNA) as circulating diagnostic markers for PC. Isin et al. tested the feasibility of using lncRNA-p21 for the detection of PC. Urine lncRNA-p21 was expressed at significantly higher levels in PC (n=30) than in BPH (n=49) and the specificity of the test was increased to 94% when lncRNA-p21 was examined together with PSA.91
Can EVs produce high yield of biomarkers in vivo?
Modern methods for EV isolation and characterization
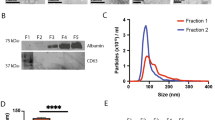
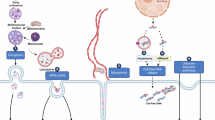
A major challenge in profiling circulating EVs is that differential centrifugation is still the gold standard and one of the most frequently used methods for EV isolation.92 While effective, the methodology is cumbersome, it requires large amount of starting material, and it is not suitable for high-throughput assays nor does it provide individual particle analysis. Luckily the field is advancing rapidly. Multiple alternative approaches have been recently developed for both purification and analysis of EVs (Figure 3). Unfortunately, most of these methods have not been standardized, which makes cross-assay comparisons difficult. It has also prevented the literature from defining specifically what EV population is studied by different investigators. Several kits are commercially available for EV isolation (Table 2). However, these kits tend to precipitate a mixture of EV populations and result in EVs of low purity (EVs are precipitated along with extra-vesicular material). Moreover, most of these methods employ buffers that digest the EV membranes, thus impairing the recovery of intact particles for qualitative and quantitative analyses. While these considerations would support the use of differential centrifugation, a few reports indicate even that methodology is imperfect, as it may favor the formation of particle clumps, interfering with amenable downstream single vesicle characterization.
For the above reasons, several recent studies have focused on developing novel approaches to purify EVs in a more rapid and sensitive manner (Figure 3). One example is the micro-nuclear magnetic resonance (μNMR) system, in which EVs are bound to antibody-coated miniaturized nanoparticles and become super-paramagnetic.93 Antibody-coated photosensitizer-beads that allow amplification of signal by a luminescent proximity homogeneous assay (ExoScreen) have also been employed.94 Recently, a tool that combines magneto and enzymatic detection of signals has been developed (iMEX). In this case EVs are first captured using magnetic beads then combined with a HRP enzyme that allows electrochemical detection.95 As novel detection tools, a time-resolved fluorescence immunoassay (TR-FIA) has recently detected, with high sensitivity, CD9 and CD63-positive exosomes in urine samples from patients with PC.96 Welton et al. recently proposed the application of ready-made chromatography columns for the study of plasma. CD9-positive fractions were thus efficiently separated from albumin.97 This system relies on antibodies and on specificity of EV markers such as EGFRvIII, CD147 and CD9, or EpCAM and CD24. Label-free approaches, to overcome this problem, include nano-plasmonic sensors (nPLEX). This high throughput system can monitor the nPLEX local refractive index, which shifts when EVs bind to a specific sensor.98 It allows elution and recovery of intact EVs, permitting downstream profiling of their cargo. A chip-based approach was also recently employed to analyze EVs in a small cohort of patients with PC. This assay based on electrochemical characterization of EVs detected increased levels of EpCAM and PSMA in patients compared to controls starting from a 25 μl of plasma.99 An aqueous two-phase system was recently proposed for EV isolation from plasma of patients with PC (n=82). Also, in this study, PSMA-positive EVs were associated with high-risk PC and higher risk of biochemical failure.100 Flow cytometric analysis of EVs has made significant advances. In particular, the use of microflow technologies (Apogeeflow.com) has enabled the direct analysis of EVs of various sizes.101, 102, 103 The recent inclusion of standardization, particles has been particularly important for achieving consistent detection and characterization of each EV population. The inclusion of fluorescent detection with small and wide-angle scatter enables microflow technologies to interrogate each particle with fluorescent antibodies, thereby bringing the advantages of traditional flow cytometry to the analysis of EVs.
Conclusion
The EV field is developing rapidly to try to better understand the biology and significance of the cargo with raising opportunities to use EV as biomarkers. Keys to success will be the miniaturization of the assays as well as standardization of EV purification and analysis systems. These recent developments hold high potentials for applicability of EV profile to clinical settings if validated in larger cohorts of patients. Indeed, preliminary investigations in different tumors like glioblastoma, colorectal cancer and ovarian cancer demonstrate that these techniques can discriminate patients from controls, and also identify patients that respond to therapy from the ones who do not.
References
Hamdy FC, Donovan JL, Lane JA, Mason M, Metcalfe C, Holding P et al. 10-Year outcomes after monitoring, surgery, or radiotherapy for localized prostatecancer. N Engl J Med 2016; 375: 1415–1424.
Siegel RL, Miller KD, Jemal A . Cancer statistics, 2016. CA Cancer J Clin 2016; 66: 7–30.
Gleason DF . Histologic grading of prostate cancer: a perspective. Hum Pathol 1992; 23: 273–279.
Epstein JI, Egevad L, Amin MB, Delahunt B, Srigley JR, Humphrey PA et al. The 2014 International Society of Urological Pathology (ISUP) Consensus Conference on Gleason Grading of Prostatic Carcinoma: Definition of Grading Patterns and Proposal for a New Grading System. Am J Surg Pathol 2016; 40: 244–252.
Gordetsky J, Epstein J . Grading of prostatic adenocarcinoma: current state and prognostic implications. Diagn Pathol 2016; 11: 25.
Rubin MA, Girelli G, Demichelis F . Genomic correlates to the newly proposed grading prognostic groups for prostate cancer. Eur Urol 2016; 69: 557–560.
Cancer Genome Atlas Research Network. The molecular taxonomy of primary prostate cancer. Cell 2015; 163: 1011–1025.
D'Elia C, Cerruto MA, Cioffi A, Novella G, Cavalleri S, Artibani W . Upgrading and upstaging in prostate cancer: From prostate biopsy to radical prostatectomy. Mol Clin Oncol 2014; 2: 1145–1149.
Andriole GL, Crawford ED, Grubb RL 3rd, Buys SS, Chia D, Church TR et al. Mortality results from a randomized prostate-cancer screening trial. N Engl J Med 2009; 360: 1310–1319.
Miller MC, Doyle GV, Terstappen LW . Significance of circulating tumor cells detected by the cellsearch system in patients with metastatic breast colorectal and prostate cancer. J Oncol 2010; 2010: 617421.
Gupta GP, Massague J . Cancer metastasis: building a framework. Cell 2006; 127: 679–695.
Gorges TM, Riethdorf S, von Ahsen O, Nastal YP, Rock K, Boede M et al. Heterogeneous PSMA expression on circulating tumor cells: a potential basis for stratification and monitoring of PSMA-directed therapies in prostate cancer. Oncotarget 2016; 7: 34930–34941.
Vogelzang NJ, Fizazi K, Burke JM, De Wit R, Bellmunt J, Hutson TE et al. Circulating tumor cells in a phase 3 study of docetaxel and prednisone with or without lenalidomide in metastatic castration-resistant prostate cancer. Eur Urol 2016; 71: 168–171.
Scher HI, Lu D, Schreiber NA, Louw J, Graf RP, Vargas HA et al. Association of AR-V7 on circulating tumor cells as a treatment-specific biomarker with outcomes and survival in castration-resistant prostate cancer. JAMA Oncol 2016; 2: 1441–1449.
Beltran H, Jendrisak A, Landers M, Mosquera JM, Kossai M, Louw J et al. The initial detection and partial characterization of circulating tumor cells in neuroendocrine prostate cancer. Clin Cancer Res 2016; 22: 1510–1519.
Carreira S, Romanel A, Goodall J, Grist E, Ferraldeschi R, Miranda S et al. Tumor clone dynamics in lethal prostate cancer. Sci Transl Med 2014; 6: 254ra125.
Romanel A, Gasi Tandefelt D, Conteduca V, Jayaram A, Casiraghi N, Wetterskog D et al. Plasma AR and abiraterone-resistant prostate cancer. Sci Transl Med 2015; 7: 312re10.
Li M, Zeringer E, Barta T, Schageman J, Cheng A, Vlassov AV . Analysis of the RNA content of the exosomes derived from blood serum and urine and its potential as biomarkers. Philos Trans R Soc Lond B Biol Sci 2014; 369.
Jansen FH, Krijgsveld J, van Rijswijk A, van den Bemd GJ, van den Berg MS, van Weerden WM et al. Exosomal secretion of cytoplasmic prostate cancer xenograft-derived proteins. Mol Cell Proteomics 2009; 8: 1192–1205.
Minciacchi VR, Freeman MR, Di Vizio D . Extracellular vesicles in cancer: exosomes, microvesicles and the emerging role of large oncosomes. Semin Cell Dev Biol 2015; 40: 41–51.
Ciardiello C, Cavallini L, Spinelli C, Yang J, Reis-Sobreiro M, de Candia P et al. Focus on extracellular vesicles: new frontiers of cell-to-cell communication in cancer. Int J Mol Sci 2016; 17: 175.
Kowal J, Arras G, Colombo M, Jouve M, Morath JP, Primdal-Bengtson B et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc Natl Acad Sci USA 2016; 113: E968–E977.
Minciacchi VR, You S, Spinelli C, Morley S, Zandian M, Aspuria PJ et al. Large oncosomes contain distinct protein cargo and represent a separate functional class of tumor-derived extracellular vesicles. Oncotarget 2015; 6: 11327–11341.
Thery C, Ostrowski M, Segura E . Membrane vesicles as conveyors of immune responses. Nat Rev Immunol 2009; 9: 581–593.
Kowal J, Tkach M, Thery C . Biogenesis and secretion of exosomes. Curr Opin Cell Biol 2014; 29: 116–125.
Booth AM, Fang Y, Fallon JK, Yang JM, Hildreth JE, Gould SJ . Exosomes and HIV Gag bud from endosome-like domains of the T cell plasma membrane. J Cell Biol 2006; 172: 923–935.
Mathivanan S, Ji H, Simpson RJ . Exosomes: extracellular organelles important in intercellular communication. J Proteomics 2010; 73: 1907–1920.
Dubois L, Stridsberg M, Kharaziha P, Chioureas D, Meersman N, Panaretakis T et al. Malignant cell-derived extracellular vesicles express different chromogranin epitopes compared to prostasomes. Prostate 2015; 75: 1063–1073.
Ronquist G . Prostasomes are mediators of intercellular communication: from basic research to clinical implications. J Intern Med 2012; 271: 400–413.
Tavoosidana G, Ronquist G, Darmanis S, Yan J, Carlsson L, Wu D et al. Multiple recognition assay reveals prostasomes as promising plasma biomarkers for prostate cancer. Proc Natl Acad Sci USA 2011; 108: 8809–8814.
Ronquist KG, Sanchez C, Dubois L, Chioureas D, Fonseca P, Larsson A et al. Energy-requiring uptake of prostasomes and PC3 cell-derived exosomes into non-malignant and malignant cells. J Extracell Vesicles 2016; 5: 29877.
Muralidharan-Chari V, Clancy J, Plou C, Romao M, Chavrier P, Raposo G et al. ARF6-regulated shedding of tumor cell-derived plasma membrane microvesicles. Curr Biol 2009; 19: 1875–1885.
Li B, Antonyak MA, Zhang J, Cerione RA . RhoA triggers a specific signaling pathway that generates transforming microvesicles in cancer cells. Oncogene 2012; 31: 4740–4749.
Muller I, Klocke A, Alex M, Kotzsch M, Luther T, Morgenstern E et al. Intravascular tissue factor initiates coagulation via circulating microvesicles and platelets. FASEB J 2003; 17: 476–478.
Di Vizio D, Kim J, Hager MH, Morello M, Yang W, Lafargue CJ et al. Oncosome formation in prostate cancer: association with a region of frequent chromosomal deletion in metastatic disease. Cancer Res 2009; 69: 5601–5609.
Hager MH, Morley S, Bielenberg DR, Gao S, Morello M, Holcomb IN et al. DIAPH3 governs the cellular transition to the amoeboid tumour phenotype. EMBO Mol Med 2012; 4: 743–760.
Wolf K, Mazo I, Leung H, Engelke K, von Andrian UH, Deryugina EI et al. Compensation mechanism in tumor cell migration: mesenchymal-amoeboid transition after blocking of pericellular proteolysis. J Cell Biol 2003; 160: 267–277.
Sanz-Moreno V, Gadea G, Ahn J, Paterson H, Marra P, Pinner S et al. Rac activation and inactivation control plasticity of tumor cell movement. Cell 2008; 135: 510–523.
Di Vizio D, Morello M, Dudley AC, Schow PW, Adam RM, Morley S et al. Large oncosomes in human prostate cancer tissues and in the circulation of mice with metastatic disease. Am J Pathol 2012; 181: 1573–1584.
Wright PK, Jones SB, Ardern N, Ward R, Clarke RB, Sotgia F et al. 17beta-estradiol regulates giant vesicle formation via estrogen receptor-alpha in human breast cancer cells. Oncotarget 2014; 5: 3055–3065.
Ma L, Li Y, Peng J, Wu D, Zhao X, Cui Y et al. Discovery of the migrasome, an organelle mediating release of cytoplasmic contents during cell migration. Cell Res 2015; 25: 24–38.
Clancy JW, Sedgwick A, Rosse C, Muralidharan-Chari V, Raposo G, Method M et al. Regulated delivery of molecular cargo to invasive tumour-derived microvesicles. Nat Commun 2015; 6: 6919.
Inder KL, Ruelcke JE, Petelin L, Moon H, Choi E, Rae J et al. Cavin-1/PTRF alters prostate cancer cell-derived extracellular vesicle content and internalization to attenuate extracellular vesicle-mediated osteoclastogenesis and osteoblast proliferation. J Extracell Vesicles 2014; 3.
Melo SA, Luecke LB, Kahlert C, Fernandez AF, Gammon ST, Kaye J et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 2015; 523: 177–182.
Etayash H, McGee AR, Kaur K, Thundat T . Nanomechanical sandwich assay for multiple cancer biomarkers in breast cancer cell-derived exosomes. Nanoscale 2016; 8: 15137–15141.
Peinado H, Aleckovic M, Lavotshkin S, Matei I, Costa-Silva B, Moreno-Bueno G et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat Med 2012; 18: 883–891.
Akers JC, Ramakrishnan V, Kim R, Skog J, Nakano I, Pingle S et al. MiR-21 in the extracellular vesicles (EVs) of cerebrospinal fluid (CSF): a platform for glioblastoma biomarker development. PLoS ONE 2013; 8: e78115.
Jakobsen KR, Paulsen BS, Baek R, Varming K, Sorensen BS, Jorgensen MM . Exosomal proteins as potential diagnostic markers in advanced non-small cell lung carcinoma. J Extracell Vesicles 2015; 4: 26659.
Skog J, Wurdinger T, van Rijn S, Meijer DH, Gainche L, Sena-Esteves M et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol 2008; 10: 1470–1476.
Hansen AG, Arnold SA, Jiang M, Palmer TD, Ketova T, Merkel A et al. ALCAM/CD166 is a TGF-beta-responsive marker and functional regulator of prostate cancer metastasis to bone. Cancer Res 2014; 74: 1404–1415.
Sandvig K, Llorente A . Proteomic analysis of microvesicles released by the human prostate cancer cell line PC-3. Mol Cell Proteomics 2012; 11: M111 012914.
Rajasekhar VK, Studer L, Gerald W, Socci ND, Scher HI . Tumour-initiating stem-like cells in human prostate cancer exhibit increased NF-kappaB signalling. Nat Commun 2011; 2: 162.
Zijlstra A, Lewis J, Degryse B, Stuhlmann H, Quigley JP . The inhibition of tumor cell intravasation and subsequent metastasis via regulation of in vivo tumor cell motility by the tetraspanin CD151. Cancer Cell 2008; 13: 221–234.
Hooper JD, Zijlstra A, Aimes RT, Liang H, Claassen GF, Tarin D et al. Subtractive immunization using highly metastatic human tumor cells identifies SIMA135/CDCP1, a 135 kDa cell surface phosphorylated glycoprotein antigen. Oncogene 2003; 22: 1783–1794.
Alajati A, Guccini I, Pinton S, Garcia-Escudero R, Bernasocchi T, Sarti M et al. Interaction of CDCP1 with HER2 enhances HER2-driven tumorigenesis and promotes trastuzumab resistance in breast cancer. Cell Rep 2015; 11: 564–576.
He Y, Wu AC, Harrington BS, Davies CM, Wallace SJ, Adams MN et al. Elevated CDCP1 predicts poor patient outcome and mediates ovarian clear cell carcinoma by promoting tumor spheroid formation, cell migration and chemoresistance. Oncogene 2016; 35: 468–478.
Josson S, Gururajan M, Hu P, Shao C, Chu GY, Zhau HE et al. miR-409-3p/-5p promotes tumorigenesis, epithelial-to-mesenchymal transition, and bone metastasis of human prostate cancer. Clin Cancer Res 2014; 20: 4636–4646.
Yang L, Dutta SM, Troyer DA, Lin JB, Lance RA, Nyalwidhe JO et al. Dysregulated expression of cell surface glycoprotein CDCP1 in prostate cancer. Oncotarget 2015; 6: 43743–43758.
Webber J, Stone TC, Katilius E, Smith BC, Gordon B, Mason MD et al. Proteomics analysis of cancer exosomes using a novel modified aptamer-based array (SOMAscan) platform. Mol Cell Proteomics 2014; 13: 1050–1064.
Hosseini-Beheshti E, Pham S, Adomat H, Li N, Tomlinson Guns ES . Exosomes as biomarker enriched microvesicles: characterization of exosomal proteins derived from a panel of prostate cell lines with distinct AR phenotypes. Mol Cell Proteomics 2012; 11: 863–885.
Duijvesz D, Burnum-Johnson KE, Gritsenko MA, Hoogland AM, Vredenbregt-van den Berg MS, Willemsen R et al. Proteomic profiling of exosomes leads to the identification of novel biomarkers for prostate cancer. PLoS ONE 2013; 8: e82589.
Overbye A, Skotland T, Koehler CJ, Thiede B, Seierstad T, Berge V et al. Identification of prostate cancer biomarkers in urinary exosomes. Oncotarget 2015; 6: 30357–30376.
Kharaziha P, Chioureas D, Rutishauser D, Baltatzis G, Lennartsson L, Fonseca P et al. Molecular profiling of prostate cancer derived exosomes may reveal a predictive signature for response to docetaxel. Oncotarget 2015; 6: 21740–21754.
Lu Q, Zhang J, Allison R, Gay H, Yang WX, Bhowmick NA et al. Identification of extracellular delta-catenin accumulation for prostate cancer detection. Prostate 2009; 69: 411–418.
Khan S, Jutzy JM, Valenzuela MM, Turay D, Aspe JR, Ashok A et al. Plasma-derived exosomal survivin, a plausible biomarker for early detection of prostate cancer. PLoS ONE 2012; 7: e46737.
Freeman MR, Yang W, Di Vizio D . Caveolin-1 and prostate cancer progression. Adv Exp Med Biol 2012; 729: 95–110.
Di Vizio D, Sotgia F, Williams TM, Hassan GS, Capozza F, Frank PG et al. Caveolin-1 is required for the upregulation of fatty acid synthase (FASN), a tumor promoter, during prostate cancer progression. Cancer Biol Ther 2007; 6: 1263–1268.
Williams TM, Hassan GS, Li J, Cohen AW, Medina F, Frank PG et al. Caveolin-1 promotes tumor progression in an autochthonous mouse model of prostate cancer: genetic ablation of Cav-1 delays advanced prostate tumor development in tramp mice. J Biol Chem 2005; 280: 25134–25145.
Tahir SA, Ren C, Timme TL, Gdor Y, Hoogeveen R, Morrisett JD et al. Development of an immunoassay for serum caveolin-1: a novel biomarker for prostatecancer. Clin Cancer Res 2003; 9 (10 Pt 1): 3653–3659.
Morello M, Minciacchi VR, de Candia P, Yang J, Posadas E, Kim H et al. Large oncosomes mediate intercellular transfer of functional microRNA. Cell Cycle 2013; 12: 3526–3536.
Logozzi M, De Milito A, Lugini L, Borghi M, Calabro L, Spada M et al. High levels of exosomes expressing CD63 and caveolin-1 in plasma of melanoma patients. PLoS ONE 2009; 4: e5219.
Coumans FA, Doggen CJ, Attard G, de Bono JS, Terstappen LW . All circulating EpCAM+CK+CD45- objects predict overall survival in castration-resistant prostate cancer. Ann Oncol 2010; 21: 1851–1857.
Kahlert C, Melo SA, Protopopov A, Tang J, Seth S, Koch M et al. Identification of double-stranded genomic DNA spanning all chromosomes with mutated KRAS and p53 DNA in the serum exosomes of patients with pancreatic cancer. J Biol Chem 2014; 289: 3869–3875.
Thakur BK, Zhang H, Becker A, Matei I, Huang Y, Costa-Silva B et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell Res 2014; 24: 766–769.
Balaj L, Lessard R, Dai L, Cho YJ, Pomeroy SL, Breakefield XO et al. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat Commun 2011; 2: 180.
Morris JPt, Wang SC, Hebrok M . KRAS, Hedgehog, Wnt and the twisted developmental biology of pancreatic ductal adenocarcinoma. Nat Rev Cancer 2010; 10: 683–695.
Kanda M, Matthaei H, Wu J, Hong SM, Yu J, Borges M et al. Presence of somatic mutations in most early-stage pancreatic intraepithelial neoplasia. Gastroenterology 2012; 142: 730–3 e9.
Lazaro-Ibanez E, Sanz-Garcia A, Visakorpi T, Escobedo-Lucea C, Siljander P, Ayuso-Sacido A et al. Different gDNA content in the subpopulations of prostate cancer extracellular vesicles: apoptotic bodies, microvesicles, and exosomes. Prostate 2014; 74: 1379–1390.
Robinson D, Van Allen EM, Wu YM, Schultz N, Lonigro RJ, Mosquera JM et al. Integrative clinical genomics of advanced prostate cancer. Cell 2015; 161: 1215–1228.
Donovan MJ, Noerholm M, Bentink S, Belzer S, Skog J, O'Neill V et al. A molecular signature of PCA3 and ERG exosomal RNA from non-DRE urine is predictive of initial prostate biopsy result. Prostate Cancer Prostatic Dis 2015; 18: 370–375.
McKiernan J, Donovan MJ, O'Neill V, Bentink S, Noerholm M, Belzer S et al. A novel urine exosome gene expression assay to predict high-grade prostate cancer at initial biopsy. JAMA Oncol 2016; 2: 882–889.
Nilsson J, Skog J, Nordstrand A, Baranov V, Mincheva-Nilsson L, Breakefield XO et al. Prostate cancer-derived urine exosomes: a novel approach to biomarkers for prostate cancer. Br J Cancer. 2009; 100: 1603–1607.
Dijkstra S, Birker IL, Smit FP, Leyten GH, de Reijke TM, van Oort IM et al. Prostate cancer biomarker profiles in urinary sediments and exosomes. J Urol 2014; 191: 1132–1138.
Hendriks RJ, Dijkstra S, Jannink SA, Steffens MG, van Oort IM, Mulders PF et al. Comparative analysis of prostate cancer specific biomarkers PCA3 and ERG in whole urine, urinary sediments and exosomes. Clin Chem Lab Med 2016; 54: 483–492.
Turchinovich A, Weiz L, Langheinz A, Burwinkel B . Characterization of extracellular circulating microRNA. Nucleic Acids Res 2011; 39: 7223–7233.
Korzeniewski N, Tosev G, Pahernik S, Hadaschik B, Hohenfellner M, Duensing S . Identification of cell-free microRNAs in the urine of patients with prostatecancer. Urol Oncol 2015; 33: 16 e7–16 e22.
Corcoran C, Rani S, O'Driscoll L . miR-34a is an intracellular and exosomal predictive biomarker for response to docetaxel with clinical relevance to prostate cancer progression. Prostate 2014; 74: 1320–1334.
Huang X, Yuan T, Liang M, Du M, Xia S, Dittmar R et al. Exosomal miR-1290 and miR-375 as prognostic markers in castration-resistant prostate cancer. Eur Urol 2015; 67: 33–41.
Costa-Pinheiro P, Ramalho-Carvalho J, Vieira FQ, Torres-Ferreira J, Oliveira J, Goncalves CS et al. MicroRNA-375 plays a dual role in prostate carcinogenesis. Clin Epigenetics 2015; 7: 42.
Scaravilli M, Porkka KP, Brofeldt A, Annala M, Tammela TL, Jenster GW et al. MiR-1247-5p is overexpressed in castration resistant prostate cancer and targets MYCBP2. Prostate 2015; 75: 798–805.
Isin M, Uysaler E, Ozgur E, Koseoglu H, Sanli O, Yucel OB et al. Exosomal lncRNA-p21 levels may help to distinguish prostate cancer from benign disease. Front Genet 2015; 6: 168.
Gardiner C, Di Vizio D, Sahoo S, Thery C, Witwer KW, Wauben M et al. Techniques used for the isolation and characterization of extracellular vesicles: results of a worldwide survey. J Extracell Vesicles 2016; 5: 32945.
Shao H, Chung J, Balaj L, Charest A, Bigner DD, Carter BS et al. Protein typing of circulating microvesicles allows real-time monitoring of glioblastoma therapy. Nat Med 2012; 18: 1835–1840.
Yoshioka Y, Kosaka N, Konishi Y, Ohta H, Okamoto H, Sonoda H et al. Ultra-sensitive liquid biopsy of circulating extracellular vesicles using ExoScreen. Nat Commun 2014; 5: 3591.
Jeong S, Park J, Pathania D, Castro CM, Weissleder R, Lee H . Integrated magneto-electrochemical sensor for exosome analysis. ACS Nano 2016; 10: 1802–1809.
Duijvesz D, Versluis CY, van der Fels CA, Vredenbregt-van den Berg MS, Leivo J, Peltola MT et al. Immuno-based detection of extracellular vesicles in urine as diagnostic marker for prostate cancer. Int J Cancer 2015; 137: 2869–2878.
Welton JL, Webber JP, Botos LA, Jones M, Clayton A . Ready-made chromatography columns for extracellular vesicle isolation from plasma. J Extracell Vesicles 2015; 4: 27269.
Im H, Shao H, Park YI, Peterson VM, Castro CM, Weissleder R et al. Label-free detection and molecular profiling of exosomes with a nano-plasmonic sensor. Nat Biotechnol 2014; 32: 490–495.
Zhou YG, Mohamadi RM, Poudineh M, Kermanshah L, Ahmed S, Safaei TS et al. Interrogating circulating microsomes and exosomes using metal nanoparticles. Small 2016; 12: 727–732.
Park YH, Shin HW, Jung AR, Kwon OS, Choi YJ, Park J et al. Prostate-specific extracellular vesicles as a novel biomarker in human prostate cancer. Sci Rep 2016; 6: 30386.
Chandler WL . Measurement of microvesicle levels in human blood using flow cytometry. Cytometry B Clin Cytom 2016; 90: 326–336.
Rose JA, Wanner N, Cheong HI, Queisser K, Barrett P, Park M et al. Flow cytometric quantification of peripheral blood cell beta-adrenergic receptor density and urinary endothelial cell-derived microparticles in pulmonary arterial hypertension. PLoS ONE 2016; 11: e0156940.
Biggs CN, Siddiqui KM, Al-Zahrani AA, Pardhan S, Brett SI, Guo QQ et al. Prostate extracellular vesicles in patient plasma as a liquid biopsy platform for prostate cancer using nanoscale flow cytometry. Oncotarget 2016; 7: 8839–8849.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Minciacchi, V., Zijlstra, A., Rubin, M. et al. Extracellular vesicles for liquid biopsy in prostate cancer: where are we and where are we headed?. Prostate Cancer Prostatic Dis 20, 251–258 (2017). https://doi.org/10.1038/pcan.2017.7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pcan.2017.7
This article is cited by
-
Extracellular vesicles from biological fluids as potential markers in castration resistant prostate cancer
Journal of Cancer Research and Clinical Oncology (2023)
-
Surface protein profiling of prostate-derived extracellular vesicles by mass spectrometry and proximity assays
Communications Biology (2022)
-
Extracellular vesicles as a source of prostate cancer biomarkers in liquid biopsies: a decade of research
British Journal of Cancer (2022)
-
Association of extracellular vesicle inflammatory proteins and mortality
Scientific Reports (2022)
-
Potential and challenges of specifically isolating extracellular vesicles from heterogeneous populations
Scientific Reports (2021)