Abstract
Background:
Although prostate cancer (PCa) is hypothesized to differ in nature between younger versus older patients, the underlying molecular distinctions are poorly understood. We hypothesized that high-throughput transcriptomic analysis would elucidate biological differences in PCas arising in younger versus older men, and would nominate potential age-specific biomarkers and therapeutic targets.
Methods:
The high-density Affymetrix GeneChip platform, encompassing >1 million genomic loci, was utilized to assess gene expression in 1090 radical prostatectomy samples from patients with long-term follow-up. We identified genes associated with metastatic progression by 10 years post-treatment in younger (age<65) versus older (age⩾65) patients, and ranked these genes by their prognostic value. We performed Gene Set Enrichment Analysis (GSEA) to nominate biological concepts that demonstrated age-specific effects, and validated a target by treating with a clinically available drug in three PCa cell lines derived from younger men.
Results:
Over 80% of the top 1000 prognostic genes in younger and older men were specific to that age group. GSEA nominated the proteasome pathway as the most differentially prognostic in younger versus older patients. High expression of proteasomal genes conferred worse prognosis in younger but not older men on univariate and multivariate analysis. Bortezomib, a Food and Drug Administration approved proteasome inhibitor, decreased proliferation in three PCa cell lines derived from younger patients.
Conclusions:
Our data show significant global differences in prognostic genes between older versus younger men. We nominate proteasomeal gene expression as an age-specific biomarker and potential therapeutic target specifically in younger men. Limitations of our study include clinical differences between cohorts, and increased comorbidities and lower survival in older patients. These intriguing findings suggest that current models of PCa biology do not adequately represent genetic heterogeneity of PCa related to age, and future clinical trials would benefit from stratification based on age.
Similar content being viewed by others
Introduction
Close to 1 million men worldwide are diagnosed each year with prostate cancer (PCa).1 The preponderance of men are diagnosed later in life, with a median age at diagnosis of 66 years in the United States.2 Although PCa mainly afflicts men in their seventh decade of life and beyond, there are still a significant number of men who are diagnosed at a younger age.3 Historically, it has been postulated that younger men who are diagnosed with PCa harbor biologically more aggressive disease than their older counterparts, resulting in poorer long-term prognosis for men diagnosed at a young age.4, 5 However, clinical findings to support this notion have to date been mixed.6, 7, 8, 9, 10, 11, 12, 13, 14
Independent of the prognosis of early versus late-life onset PCa, it is possible that the biological pathways that drive this disease differ by age. However, to date, there have been no studies examining the similarities and differences in the prognostic drivers of PCa in different age groups. Identifying these potential age-related biomarkers could improve tailoring of treatment by patient age.
In this study, we sought to define the landscape of gene expression in localized PCas from patients diagnosed at a younger versus older age in the largest high-throughput gene expression profiling experiment in PCa to date. We identified genes prognostic for metastatic progression in younger patients versus older patients, and nominate biological pathways enriched in these prognostic gene sets. To further pursue the top nominated targetable pathway, we investigated the potential of the proteasome pathway as an age-specific biomarker and therapeutic target in PCas from younger patients.
Materials and methods
Study design and tissue samples
Formalin-fixed paraffin-embedded tumor samples were obtained from four prostatectomy patient cohorts enrolled at the Mayo Clinic (MC I and II), Cleveland Clinic (CC) and Thomas Jefferson University (TJU) under informed consent protocols approved by local Institutional Review Boards. The MCI cohort consisted of a nested case–control study with 545 men in matched triples of metastatic progression, biochemical recurrence after radical prostatectomy (RP), and patients with no evidence of disease.15 The MCII cohort consisted of a case–cohort study that sampled a cohort of 1010 high-risk men that underwent RP to generate a final cohort of 232 samples as described previously.16 The TJU cohort is comprised of 143 patients with pT3 or margin-positive disease who underwent RP and post-RP radiotherapy of whom 130 microarray samples were available.17 Patients from the CC cohort were obtained from a case–control study in which 2317 conservatively treated high-risk RP patients who did not receive adjuvant therapy were sampled to achieve a 3:1 ratio for non-metastatic versus metastatic progression, for a total of 183 samples.18 RNA extraction and microarray hybridization were performed using clinical-grade techniques in a Clinical Laboratory Improvement Amendments—certified laboratory facility (GenomeDx Biosciences, San Diego, CA, USA). The normalization and summarization of the microarray samples were done with the Single Channel Array Normalization algorithm with quality control performed as described previously.16, 17, 18, 19 Gene expression for each gene was calculated using the Affymetrix Core level summaries for annotated genes. Microarray data are available on the NCBI Gene Expression Omnibus as accession numbers GSE46691 (MCI), GSE62116 (MCII) and GSE62667 (CC).
Age and prognosis
To evaluate whether age was associated with metastasic progression after RP, patients were stratified by age into those <65 and ⩾65 at the time of RP, which roughly divided the number of patients evenly (median age was 64 years), and is approximately the median age of PCa diagnosis in the United States (66 years). Patient age was assessed as a predictor of metastatic progression within 10 years of RP in a pooled analysis using a random effects model (REM) with inverse variance for weighting using the R ‘meta’ package.
Nomination of metastasis-associated genes in age groups
To nominate genes associated with metastasis after RP, the differential expression of primary tumor tissue from cases that developed metastasis within 10 years of RP was compared with controls that did not using the R package ‘MetaDE’ with a REM.20 Genes were ranked by P-value and the top 1000 genes prognostic for metastases were selected from each age cohort. This was used instead of a P-value cutoff as the younger age group was slightly larger and thus had increased statistical power. Heat maps of the genes associated with metastasis were generated and clustered using hierarchical clustering. Gene expression as a continuous variable was correlated with age at RP using a pooled REM of Spearman’s correlation using the R ‘meta’ package.
Gene set enrichment analysis
Identification of biological concepts enriched in genes associated with metastasis in younger and older men was performed using Gene Set Enrichment Analysis (GSEA). The C2: curated gene sets, C5: GO gene sets and C6: oncogenic signature gene sets were used. The REM T-statistic for genes was scaled to be between −1 and 1 to account for the differences in statistical power, and the difference in the scaled T-statistic between younger and older men was calculated (the delta-T). This value was input to GSEA as a ranked list of all genes.
Univariate and multivariate analyses
Genes with a delta-T<−0.5 in the most negatively enriched gene set were selected for further analysis. The expression of each gene was split into ‘high’ and ‘low’ based on Partition Around Medoids clustering.21, 22 The performance of a gene was evaluated using a REM comparing the 10-year metastasis rate in high versus low expression also using the R ‘meta’ package. Pooled multivariate logistic regression analysis was performed, with age analyzed per year, Gleason and PSA stratified into high/low (Gleason 8–10/⩽7, PSA>8/⩽8), and stratification by cohort as described previously18 to account for baseline differences, both measured and unmeasured, between cohorts. Kaplan–Meier curves were generated and a P-value was calculated using the Log-rank test. All statistical tests were performed using R and SAS v9.3 (Cary, NC, USA).
Drug sensitivity
Experiments with the proteasome inhibitor bortezomib were carried out on three cell lines, LNCaP, VCAP and PC3. Cell lines were purchased from ATCC and tested for mycoplasma contamination. Cells were seeded at a density of 5000 cells per well plated in 96-well culture plates and treated with concentrations from 10 pM to 10 nM. WST-1 assays (Roche, Basel, Switzerland) were performed after 72 h and readings were recorded at 440 nm as previously described.21, 23, 24 GraphPad Prism software (La Jolla, CA, USA) was used to generate non-linear regression curves and calculate IC50 values.
Results
Demographics
Table 1 displays the overall demographics of each cohort. All cohorts had long-term clinical follow-up ranging from a mean of 80 to 160 months post-surgery. The mean follow-up for patients that did not have a metastatic event was 185 months for MCI, 83 months for MCII, 112 months for CC and 104 months for TJU. The cohorts were also as a whole higher risk than the general population, with significant proportions in all cohorts of Gleason 8–10, pre-operative PSA>20 or T3+ disease. An age cutoff of 65 years was used as it was close to the median age of the cohorts (64 years). Supplementary Table 1 shows the age cohort stratified demographics, and demonstrates that there were some significant differences within cohorts.
Age is not associated with prognosis
Age >65 versus ⩽65 was not significantly associated with the 10-year metastasis rate, though we do see a trend toward worse prognosis with older age in all cohorts (Figure 1a). When we assess each cohort individually and compare the mean age at RP for the patients who did and did not develop metastasis within 10 years, they are similar in every cohort (Figure 1b–e).
(a) Forest plot showing the overall effect of age on 10-year metastasis in four clinical cohorts (MCI, MCII, CC and TJU). Age was not significantly associated with metastasis in any cohort individually nor in a pooled random effects model. Odds ratio is comparing the odds of 10-year metastatic progression in men >65 versus ⩽65. Bar plot showing the mean age (±s.e.m.) in patients who had a metastasis by 10 years versus those who did not in CC (b), MCI (c), MCII (d) and TJU (e). The mean ages were not significantly different in any of the groups between patients who metastasized by 10 years and those that did not.
Prognostic genes are different in older versus younger men
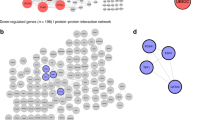
Although age itself was not prognostic in our cohorts, we hypothesized that biological signatures would differ between younger versus older men, and that the genes that are prognostic in younger men would differ from those in older men. We looked at the top 1000 prognostic genes in patients age <65 and ⩾65, and found that only 178 were shared, and the vast majority (822) were unique to each age group (Figure 2a). The 178 genes that were prognostic in older and younger men included well-known PCa biomarkers such as TOP2A and MKI67.25, 26 Hierarchical clustering of the 1000 genes prognostic in those age <65 and the 1000 genes prognostic in those age ⩾65 show that they are able to differentiate patients who will go on to develop metastatic disease, and those who will not, in their respective age groups. These results demonstrate a clear difference in the genes, which are associated with aggressive disease in younger versus older men. Interestingly, we found that the difference in prognostic ability was not owing to differential expression of the genes by age. The most positively correlated gene only had a correlation coefficient of 0.12, and the most negatively correlated gene had a correlation coefficient of −0.13, indicating a weak correlation of gene expression and age (Figure 2b).
(a) Venn diagram in the center of the panel displays the overlap in the top 1000 genes that were associated with metastatic progression across all four cohorts. 343 genes were downregulated (blue) and 479 genes were upregulated (yellow) only in metastatic patients age <65. 321 genes were downregulated (red) and 501 genes were upregulated (green) only in metastatic patients age ⩾65. 128 genes were upregulated (yellow–green) and 50 genes were downregulated (purple) in metastatic patients independent of age. The heat map on the left panel displays the expression of all 1000 genes prognostic in younger patients. The genes represented by the yellow bar are upregulated and the genes represented by the blue bar are downregulated in metastatic patients age <65. The heat map on the right panel displays the expression of all 1000 genes prognostic in older patients. The genes represented by the green bar are upregulated and the genes represented by the red bar are downregulated in metastatic patients age ⩾65. Hierarchical clustering in both age groups was able to stratify metastasis (the top bar above the heat maps). (b) Bar plot that compares the pooled Spearman’s correlation coefficient of the expression of all genes versus age. The correlation coefficients ranged from −0.13 to 0.12.
Age-specific predictors of metastasis
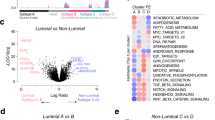
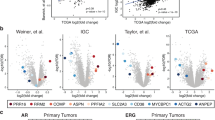
We then sought to characterize the biological differences leading to metastasis in younger versus older men. All genes on the microarray were ranked based on the difference between how prognostic they were in younger versus older men. A positive delta-T signified that higher expression of a gene confers worse prognosis in older men and/or better prognosis in younger men. A negative delta-T signifies the opposite, that higher expression confers better prognosis in older men and/or worse prognosis in younger men. GSEA was then run on this ranked list, and the top 10 out of >5000 gene sets are shown in Figure 3. Four out of the top five most negatively enriched gene sets (gene sets preferentially associated with worse prognosis in younger men compared with older men) consisted of genes associated with proteasomes/protein degradation (Figures 3a and c). Another group of gene sets associated with translation initiation was also prominently enriched, with an extracellular matrix pathway finishing out the top 10. Other notably negatively enriched gene sets were the vascular endothelial growth factor pathway and MTOR pathway ranked 22nd and 23rd respectively. Interestingly, eight out of the top 10 positively enriched gene sets (gene sets preferentially associated with worse prognosis in older men compared with younger men) involved ion channels (Figures 3b and d).
(a) GSEA-enrichment plot of the most negatively enriched gene set: Biocarta proteasome pathway. (b) GSEA-enrichment plot of the most positively enriched gene set: ion channel activity. (c) Bar plot depicting the normalized enrichment scores (NES) of the top 10 most negatively enriched gene sets, which contain several gene sets related to proteasomes (green) and translation initiation (red). (d) Bar plot depicting the normalized enrichment scores (NES) of the top 10 most positively enriched gene sets, which contain several gene sets related to ion channels (blue).
We focused on the top negatively enriched gene set that contains proteasomal genes curated from Biocarta, given the existence of Food and Drug Administration approved proteasome inhibitors (bortezomib). We found that high expression of five (PSMB4, PSMB7, PSMD14, PSMB2 and PSMD11) out of the top six genes (Figure 4a) were all associated with significantly worse 10-year metastatic rate in younger men, and the final gene PSMD6 showed the same trend with borderline significance (Figures 4c–h). None of these six genes were prognostic in older men (data not shown). In addition, we found that high expression of three or more of any of these six proteasome genes could be combined into a classifier, which was also significantly prognostic in younger but not older men (Figure 4b). On multivariable analysis (Table 2), high expression in three or more of the proteasome genes was significantly and strongly prognostic for 10-year metastasis rate even after taking into account clinical and pathologic variables such as Gleason and PSA as well as inter-cohort differences (odds ratio=2.81, P=0.00048). On Kaplan–Meier analysis in the pooled cohort, the proteasome classifier predicts metastasis-free survival in younger patients (Figure 4i; hazard ratio=1.8, P=0.00036) but not in older patients (Figure 4j; hazard ratio=1.2, P=0.22).
(a) Table showing the delta-T values of the top proteasomal genes (delta-T<0.5) from the most negatively enriched gene set (Biocarta proteasome pathway). (b) Forest plot showing that high expression of three or more of any of these proteasomal genes conferred worse prognosis only in younger men. When examining these genes individually, high expression of PSMB4 (c), PSMB7 (d), PSMD14 (f), PSMB2 (g) and PSMD11 (h) all conferred significantly worse prognosis only in younger men, with PSMD6 showing the same trend with borderline significance (e). Kaplan–Meier curves show high expression of three or more of any of these proteasomal genes confers worse metastasis-free survival in younger (i) but not older (j) men.
Proteasome inhibitors inhibit growth in cell lines derived from younger men
The nomination of proteasomal genes as prognostic in younger but not older men was intriguing because bortezomib has undergone clinical trials in PCa. We subsequently characterized the in vitro response to bortezomib in three widely used PCa cell lines, PC3, LNCaP and VCAP (Figure 5). We found that proliferation of all three cell lines were inhibited by low concentrations of bortezomib, with similar IC50s of 4.26 nM for PC3, 7.59 nM for LNCaP and 2.41 nM for VCAP, which are consistent with previously reported results.27, 28 All of these cell lines were derived from patients age <65 (PC3: 62 years old,29 LNCaP: 50 years old30 and VCAP: 59 years old31). To our knowledge, there are no PCa cell lines available from older patients.
Discussion
Current understanding of the age-specific differences in PCa tumor biology is limited. Therefore, we performed the largest high-throughput gene expression profiling experiment in PCa to date on over a thousand clinical samples. We then used unbiased approaches to identify the most age-specific prognostic genes and characterized their biologic function.
Our findings suggest that there was no significantly higher risk for developing metastatic disease in older or younger men. Men <65 years of age had the same incidence of metastases 10 years post-treatment as men who were ⩾65 years old at the time of treatment. Although age was not prognostic for metastatic progression, the genes associated with metastatic disease differed drastically between younger and older patients. Of the top 1000 genes associated with metastases in young and older patients, only 178 of the 2000 identified genes overlapped between these two age groups, suggesting that a stark contrast may exist between the genomic predictors of metastasis in men <65 and ⩾65 at time of treatment. Well-known PCa biomarkers such as TOP2A and MKI67 were unsurprisingly prognostic in all ages. Even though there was great distinction between the prognostic genes between the two age cohorts, overall, there was very little correlation with any individual gene and patient age. When we examined the individual genes that displayed the most age-specific prognostic potential, biological clusters of ion channels, translation initiation and proteasomes were identified as the top positively and negatively enriched gene sets out of >5000 analyzed.
The proteasome is a multi-subunit complex responsible for cellular protein degradation, and given that there is a Food and Drug Administration approved proteasomal inhibitor (bortezomib), we focused our remaining analysis on the proteasomal genes. Individual proteasomal genes were combined into a simple classifier that could predict metastatic progression only in younger men, even after accounting for PSA, Gleason and inter-cohort differences in a cohort-stratified pooled multivariate analysis. We show that bortezomib inhibits growth in three commonly used cell line models derived in patients under the age of 65. Although the exact mechanism for an age-specific role of proteasomes is unclear, there is a preponderance of evidence that the aging process reduces proteasomal activity in a wide range of tissues.32, 33, 34, 35, 36, 37 It is possible that PCa in younger men, who have more proteasomal activity, remains dependent on proteasomes for essential cellular functions and so can be successfully targeted. In older men, cancer cells may have adapted to lower proteasomal activity, and thus are less affected by proteasomal inhibition. Our findings are of significant clinical relevance as several early-phase clinical trials have assessed the use of bortezomib in the treatment of PCa. In advanced hormone resistant PCa, bortezomib has been underwhelming to date.38, 39, 40, 41 However, in our data, proteasomes were prognostic from the time of initial prostatectomy for development of future metastases in men <65 years old, not at a metastatic stage. Bortezomib has only been assessed in PCa patients in an earlier disease state in very small clinical trials without evaluation of metastasis or survival as an end point but these trials suggest that treatment can change the PSA trajectory.42, 43 These results with our current findings suggest that the use of botezomib for localized PCa in young men is an area worthy of further investigation.
Although our study was very large with over 1000 patients, it is based on retrospective data and therefore does not control for any unmeasured confounding factors. There are also potential batch effects from the multi-institutional and multi-cohort nature of our data, though we attempt to correct for this using the same clinical grade microarray platform for all samples, and by statistically correcting for this using REMs and stratification. Examining different age groups can also be confounded by increased comorbidities and lower survival in older patients, which we attempt to somewhat mitigate by focusing on metastatic progression. Finally, although cell lines present an easily studied in vitro model system for studying PCa, their biology may have diverged from their original in vivo phenotype during the immortalization process. As described above, there is also an absence of cell lines representing men who were diagnosed with PCa at an older age. This could represent a contributing factor to why therapies that look promising in cell line models fail to validate in clinical trials, which usually do not stratify by age.
In summary, our data suggest that age alone is not prognostic for future development of metastatic disease in men with PCa treated with RP. However, there is a striking difference between the genomic correlates of metastatic progression in men <65 and ⩾65 years of age. Notably, we identify proteasomes as a potential therapeutic target in localized PCa especially for men <65 years old. We believe our results support continued study of proteasome inhibitors in localized PCa in younger patients, and if these results are independently validated, we propose that future clinical trials should consider age stratification or selection.
References
Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM . Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer 2010; 127: 2893–2917.
American Cancer Society. What are the key statistics about prostate cancer? 2015.
Walsh PC . Cancer surveillance series: interpreting trends in prostate cancer—part I: evidence of the effects of screening in recent prostate cancer incidence, mortality, and survival rates. J Urol 2000; 163: 364–365.
Sandhu DP, Munson KW, Benghiat A, Hopper IP . Natural history and prognosis of prostate carcinoma in adolescents and men under 35 years of age. Br J Urol 1992; 69: 525–529.
Kerr LA, Zincke H . Radical retropubic prostatectomy for prostate cancer in the elderly and the young: complications and prognosis. Eur Urol 1994; 25: 305–311 discussion 311-302.
Hamstra DA, Bae K, Pilepich MV, Hanks GE, Grignon DJ, McGowan DG et al. Older age predicts decreased metastasis and prostate cancer-specific death for men treated with radiation therapy: meta-analysis of radiation therapy oncology group trials. Int J Radiat Oncol Biol Phys 2011; 81: 1293–1301.
Smith CV, Bauer JJ, Connelly RR, Seay T, Kane C, Foley J et al. Prostate cancer in men age 50 years or younger: a review of the Department of Defense Center for Prostate Disease Research multicenter prostate cancer database. J Urol 2000; 164: 1964–1967.
Briganti A, Spahn M, Joniau S, Gontero P, Bianchi M, Kneitz B et al. Impact of age and comorbidities on long-term survival of patients with high-risk prostate cancer treated with radical prostatectomy: a multi-institutional competing-risks analysis. Eur Urol 2013; 63: 693–701.
Siddiqui SA, Sengupta S, Slezak JM, Bergstralh EJ, Leibovich BC, Myers RP et al. Impact of patient age at treatment on outcome following radical retropubic prostatectomy for prostate cancer. J Urol 2006; 175 (3 Pt 1): 952–957.
Parker CC, Gospodarowicz M, Warde P . Does age influence the behaviour of localized prostate cancer? BJU Int 2001; 87: 629–637.
Parker PM, Rice KR, Sterbis JR, Chen Y, Cullen J, McLeod DG et al. Prostate cancer in men less than the age of 50: a comparison of race and outcomes. Urology 2011; 78: 110–115.
Bechis SK, Carroll PR, Cooperberg MR . Impact of age at diagnosis on prostate cancer treatment and survival. J Clin Oncol 2011; 29: 235–241.
Tward JD, Lee CM, Pappas LM, Szabo A, Gaffney DK, Shrieve DC . Survival of men with clinically localized prostate cancer treated with prostatectomy, brachytherapy, or no definitive treatment: impact of age at diagnosis. Cancer 2006; 107: 2392–2400.
Salinas CA, Tsodikov A, Ishak-Howard M, Cooney KA . Prostate cancer in young men: an important clinical entity. Nat Rev Urol 2014; 11: 317–323.
Nakagawa T, Kollmeyer TM, Morlan BW, Anderson SK, Bergstralh EJ, Davis BJ et al. A tissue biomarker panel predicting systemic progression after PSA recurrence post-definitive prostate cancer therapy. PloS One 2008; 3: e2318.
Karnes RJ, Bergstralh EJ, Davicioni E, Ghadessi M, Buerki C, Mitra AP et al. Validation of a genomic classifier that predicts metastasis following radical prostatectomy in an at risk patient population. J Urol 2013; 190: 2047–2053.
Den RB, Feng FY, Showalter TN, Mishra MV, Trabulsi EJ, Lallas CD et al. Genomic prostate cancer classifier predicts biochemical failure and metastases in patients after postoperative radiation therapy. Int J Radiat Oncol Biol Phys 2014; 89: 1038–1046.
Prensner JR, Zhao S, Erho N, Schipper M, Iyer MK, Dhanasekaran SM et al. RNA biomarkers associated with metastatic progression in prostate cancer: a multi-institutional high-throughput analysis of SChLAP1. Lancet Oncol 2014; 15: 1469–1480.
Erho N, Crisan A, Vergara IA, Mitra AP, Ghadessi M, Buerki C et al. Discovery and validation of a prostate cancer genomic classifier that predicts early metastasis following radical prostatectomy. PloS One 2013; 8: e66855.
Wang X, Kang DD, Shen K, Song C, Lu S, Chang LC et al. An R package suite for microarray meta-analysis in quality control, differentially expressed gene analysis and pathway enrichment detection. Bioinformatics 2012; 28: 2534–2536.
Prensner JR, Chen W, Han S, Iyer MK, Cao Q, Kothari V et al. The long non-coding RNA PCAT-1 promotes prostate cancer cell proliferation through cMyc. Neoplasia 2014; 16: 900–908.
Reynolds GR AP, De La Iglesia B, Rayward-Smith VJ . Clustering Rules: A Comparison of Partitioning and Hierarchial Clustering Algorithms. J Math Model Algorithms 2006; 5: 475–504.
Prensner JR, Chen W, Iyer MK, Cao Q, Ma T, Han S et al. PCAT-1, a long noncoding RNA, regulates BRCA2 and controls homologous recombination in cancer. Cancer Res 2014; 74: 1651–1660.
Han S, Brenner JC, Sabolch A, Jackson W, Speers C, Wilder-Romans K et al. Targeted radiosensitization of ETS fusion-positive prostate cancer through PARP1 inhibition. Neoplasia 2013; 15: 1207–1217.
Kosari F, Munz JM, Savci-Heijink CD, Spiro C, Klee EW, Kube DM et al. Identification of prognostic biomarkers for prostate cancer. Clin Cancer Res 2008; 14: 1734–1743.
Tollefson MK, Karnes RJ, Kwon ED, Lohse CM, Rangel LJ, Mynderse LA et al. Prostate cancer Ki-67 (MIB-1) expression, perineural invasion, and gleason score as biopsy-based predictors of prostate cancer mortality: the Mayo model. Mayo Clin Proc 2014; 89: 308–318.
Hu W, Zheng RR, Cui HX, Yue D, Wang Y, Jiang YH . Effects of bortezomib in sensitizing human prostate cancer cell lines to NK-mediated cytotoxicity. Asian J Androl 2012; 14: 695–702.
Kiliccioglu I, Konac E, Varol N, Gurocak S, Yucel Bilen C . Apoptotic effects of proteasome and histone deacetylase inhibitors in prostate cancer cell lines. Genet Mol Res 2014; 13: 3721–3731.
Kaighn ME, Narayan KS, Ohnuki Y, Lechner JF, Jones LW . Establishment and characterization of a human prostatic carcinoma cell line (PC-3). Invest Urol 1979; 17: 16–23.
Horoszewicz JS, Leong SS, Kawinski E, Karr JP, Rosenthal H, Chu TM et al. LNCaP model of human prostatic carcinoma. Cancer Res 1983; 43: 1809–1818.
Korenchuk S, Lehr JE, MClean L, Lee YG, Whitney S, Vessella R et al. VCaP, a cell-based model system of human prostate cancer. In Vivo 2001; 15: 163–168.
Carrard G, Bulteau AL, Petropoulos I, Friguet B . Impairment of proteasome structure and function in aging. Int J Biochem Cell Biol 2002; 34: 1461–1474.
Bulteau AL, Petropoulos I, Friguet B . Age-related alterations of proteasome structure and function in aging epidermis. Exp Gerontol 2000; 35: 767–777.
Carrard G, Dieu M, Raes M, Toussaint O, Friguet B . Impact of ageing on proteasome structure and function in human lymphocytes. Int J Biochem Cell Biol 2003; 35: 728–739.
Bulteau AL, Szweda LI, Friguet B . Age-dependent declines in proteasome activity in the heart. Arch Biochem Biophys 2002; 397: 298–304.
Ethen CM, Hussong SA, Reilly C, Feng X, Olsen TW, Ferrington DA . Transformation of the proteasome with age-related macular degeneration. FEBS Lett 2007; 581: 885–890.
Viteri G, Carrard G, Birlouez-Aragon I, Silva E, Friguet B . Age-dependent protein modifications and declining proteasome activity in the human lens. Arch Biochem Biophys 2004; 427: 197–203.
Dreicer R, Petrylak D, Agus D, Webb I, Roth B . Phase I/II study of bortezomib plus docetaxel in patients with advanced androgen-independent prostate cancer. Clin Cancer Res 2007; 13: 1208–1215.
Hainsworth JD, Meluch AA, Spigel DR, Barton J Jr., Simons L, Meng C et al. Weekly docetaxel and bortezomib as first-line treatment for patients with hormone-refractory prostate cancer: a Minnie Pearl Cancer Research Network phase II trial. Clin Genitourin Cancer 2007; 5: 278–283.
Morris MJ, Kelly WK, Slovin S, Ryan C, Eicher C, Heller G et al. A phase II trial of bortezomib and prednisone for castration resistant metastatic prostate cancer. J Urol 2007; 178: 2378–2383.
Price N, Dreicer R . Phase I/II trial of bortezomib plus docetaxel in patients with advanced androgen-independent prostate cancer. Clin Prostate Cancer 2004; 3: 141–143.
Kraft AS, Garrett-Mayer E, Wahlquist AE, Golshayan A, Chen CS, Butler W et al. Combination therapy of recurrent prostate cancer with the proteasome inhibitor bortezomib plus hormone blockade. Cancer Biol Ther 2011; 12: 119–124.
Ayala G, Yan J, Li R, Ding Y, Thompson TC, Mims MP et al. Bortezomib-mediated inhibition of steroid receptor coactivator-3 degradation leads to activated Akt. Clin Cancer Res 2008; 14: 7511–7518.
Acknowledgements
We acknowledge the assistance of Steven Kronenberg with graphic design. FYF, EMS, AER, EAK and RBD are supported by the Prostate Cancer Foundation. GenomeDx Biosciences helped collect and manage the microarray data used in this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
SGZ and FYF report travel grants from GenomeDx Biosciences. NE and ED are employees of GenomeDx Biosciences. EMS, AER, PLN and EAK are consultants for GenomeDx Biosciences. RBD reports grants from GenomeDx Biosciences. RBJ reports a patent ‘Cancer Diagnostics Using Biomarkers’ licensed to GenomeDx Biosciences. FYF has previously served on an advisory board for GenomeDx Biosciences. The remaining authors declare no conflicts of interest.
Additional information
Supplementary Information accompanies the paper on the Prostate Cancer and Prostatic Diseases website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Zhao, S., Jackson, W., Kothari, V. et al. High-throughput transcriptomic analysis nominates proteasomal genes as age-specific biomarkers and therapeutic targets in prostate cancer. Prostate Cancer Prostatic Dis 18, 229–236 (2015). https://doi.org/10.1038/pcan.2015.22
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/pcan.2015.22