Abstract
Dishevelled (Dvl) is a key regulator of Wnt signaling both in the canonical and non-canonical pathways. Here we report the identification of a regulatory domain of ubiquitination (RDU) in the C-terminus of Dvl. Mutations in the RDU resulted in accumulation of polyubiquitinated forms of Dvl, which were mainly K63 linked. Small interfering RNA-based screening identified Usp14 as a mediator of Dvl deubiquitination. Genetic and chemical suppression of Usp14 activity caused an increase in Dvl polyubiquitination and significantly impaired downstream Wnt signaling. These data suggest that Usp14 functions as a positive regulator of the Wnt signaling pathway. Consistently, tissue microarray analysis of colon cancer revealed a strong correlation between the levels of Usp14 and β-catenin, which suggests an oncogenic role for Usp14 via enhancement of Wnt/β-catenin signaling.
Similar content being viewed by others
Introduction
The Wnt signaling pathway regulates key developmental processes during embryogenesis as well as various homeostatic mechanisms in adult animals.1, 2 Aberrant activation of Wnt signaling is observed in diverse human disorders, including cancer.3 The canonical Wnt pathway (Wnt/β-catenin pathway) maintains β-catenin at a level insufficient for transcriptional activation, through proteolysis successively mediated via (1) phosphorylation by the destruction complex consisting of Axin, glycogen synthase kinase 3β, adenomatous polyposis coil and casein kinase 1, (2) polyubiquitination by SCFβ-TrCP E3 ligase and (3) proteasomal degradation.4 Wnt proteins bind to Frizzled (Fzd) receptor and low-density lipoprotein-related protein 5/6 co-receptor, followed by translocation of Dishevelled (Dvl) and Axin from cytoplasm to plasma membrane to form the Wnt/Fzd/LRP/Dvl/Axin receptor complex, which functions to suppress cytoplasmic β-catenin levels. The accumulation of β-catenin in the cytosol drives its movement into the nucleus, where it activates the expression of various target genes.4, 5
Vertebrates have three Dvl homologs — Dvl1, Dvl2 and Dvl3 — and they are common downstream mediators of Wnt signaling in both canonical and non-canonical pathways.6 In canonical Wnt signaling, translocation of Dvls to the receptor complex is essential for phosphorylation of the PPPSP motif in lipoprotein-related protein 5/6, which functions as an Axin docking site to promote stabilization of β-catenin.7, 8 In non-canonical Wnt signaling, which requires neither lipoprotein-related protein 5/6 phosphorylation nor β-catenin-mediated gene expression, Fzd-bound Wnt activates Dvls and regulates various downstream effectors, including Rac, Rho, CaMKII and the PKC pathway.9
Dvls contain three conserved domains, including DIX (Dvl and Axin), PDZ (Post Synaptic Density-95, discs-large and Zonula occludens-1) and DEP (Dvl, Egl-10 and Pleckstrin) domains. The Dvl and Axin domain is important for canonical Wnt signaling, the DEP domain for non-canonical and the PDZ domain for both Wnt signaling pathways.6 Therefore, it is thought that Dvls can differentiate the canonical or non-canonical Wnt signaling pathways in response to various combinations of Wnt/receptor activation, although detailed molecular mechanisms remain elusive. Upon binding of Wnt to receptors, extensive phosphorylation, which is essential for the activation of both canonical and non-canonical signaling, occurs in the central regions of Dvl containing the PDZ and DEP domains. Kinases and other factors involved in this hyperphosphorylation event include casein kinase 1, casein kinase 2, PAR1, β-arrestin and PP2A.10 The activity of Dvl is also negatively regulated through polyubiquitination by factors such as KLHL12, NEDL1 and inversin, with modified Dvl being degraded by the proteasome.10, 11 In addition to the ubiquitin-proteasome system, lysosomal degradation pathways may also regulate the stability of Dvl, according to some reports.12, 13 In particular, ubiquitination of Dvl by the von Hippel–Lindau E3 ligase is thought to promote its degradation by the autophagy-lysosome system.14 Most studies on the regulation of Dvl ubiquitination have focused on the identification of E3 ligases. However, it has been recently shown that CYLD, a deubiquitinating enzyme (DUB), controls Wnt/β-catenin signaling by disassembling K63-linked polyubiquitin chains of Dvl,15 thereby providing another mechanism for the regulation of Wnt signal transduction.
When we compared the sequences of Dvl proteins across species, we found that an ∼40-amino-acid span in the C-terminus of Dvl is highly conserved among all vertebrates but not present in Drosophila Dishevelled (Dsh). The function of this region is not well understood. In this report, we provide evidence that the C-terminus of Dvl regulates the ubiquitination status of Dvl via direct binding of K63-linked polyubiquitin chains. Mutation of key hydrophobic residues in the C-terminus of Dvl resulted in increased K63-linked ubiquitination of Dvl. These results led us to hypothesize that the C-terminal domain of Dvl may cooperate with unknown DUB(s) to control the ubiquitination status of Dvl. Based on systematic small interfering RNA (siRNA) screening and biochemical analysis, we identified Usp14 as a major DUB of Dvl. Specifically, Usp14 regulates the ubiquitination of Dvl and its subsequent phosphorylation, which is essential for activation of downstream Wnt signaling. We also identified a strong correlation between the levels of Usp14 and β-catenin in human colorectal cancer cells. Our data collectively indicate that Usp14 may have oncogenic potential via enhancement of Wnt/β-catenin signaling in colorectal cancer.
Results
Mutations in the C-terminal domain of Dvl enhance K63-linked polyubiquitination of Dvl
An ∼40-amino-acid segment in the C-terminus of Dvl is highly conserved among vertebrates, while being absent in Drosophila (Supplementary Figures S1a and S1b).6 This region stabilizes the interaction between Dvl and Fzd through direct contact with Fzd.16 Whereas BLAST searches on this conserved sequence revealed no significantly similar sequence, we found that the C-terminal domain of Dvl retained hydrophobic patches as well as several key residues that are similar to those of ubiquitin-associated domains, in which these residues participate in ubiquitin recognition (Supplementary Figure S1c). It has been reported that the sequences of bona fide ubiquitin-associated domains from Rad23, Ede1, Ddi1 and Dsk2 share no significant overall homology, whereas they form highly similar three-helix structures, mainly comprising hydrophobic residues.17
We reasoned that if the C-terminal domain of Dvl behaves as a ubiquitin-associated-like domain, then deletion of this domain may affect ubiquitination of Dvl. To test this possibility, we transiently overexpressed wild-type Dvl (Dvl-WT) and a C-terminal domain deletion mutant (Dvl-ΔRDU; RDU stands for regulatory domain of ubiquitination, see below) in HEK293T cells and compared their stability and ubiquitination. While the steady-state levels of Dvl-WT and Dvl-ΔRDU were comparable, the levels of polyubiquitinated forms of Dvl-ΔRDU were strikingly elevated over that of wild type (Figure 1a). As immunoprecipitation was performed under denaturing conditions (Supplementary Figure S4a; see details in Materials and methods), increased ubiquitination seemed to occur on Dvl rather than proteins associated with it.
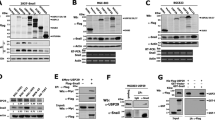
Mutations in C-terminal domain of Dvl enhance level of K63-linked polyubiquitination. (a) Deletion of the RDU of Dvl increases polyubiquitination. Plasmids expressing HA-tagged ubiquitin were co-transfected with myc-tagged Dvl-WT, Dvl-ΔRDU or empty vector into HEK293T cells. Total cell lysates were subjected to immunoprecipitation with anti-myc antibody, followed by immunoblotting with anti-HA antibody. α-tubulin is a loading control. (b) Polyubiquitin chains of Dvl-ΔRDU mainly form through K63 linkages. Experiments were performed as described in (a) except for the usage of flag-tagged Ub-K48R or Ub-K63R mutants. After immunoprecipitation using anti-myc antibody, ubiquitinated fractions of Dvl were detected by immunoblotting with anti-flag antibody. (c) C-terminus of Dvl containing the RDU (Dvl-ct) directly interacts with K63-linked polyubiqutin chains, but not with monoubiquitin or K48-linked polyubiquitin chains. Recombinant GST-Dvl-ct, GST-Dvl-ct-VK2 or GST alone (empty) were incubated with various ubiquitin species, followed by GST-pull-down, SDS–PAGE and immunoblotting for Dvl-bound ubiquitin proteins (right, top). Input is 10% of total ubiquitins used in the pull-down assay (left). Coomassie Brilliant Blue staining of pulled-down GST species is shown in the bottom panels as a control (right, bottom). (d) Point mutations of putative ubiquitin recognition residues (V644 and V667) in the RDU of Dvl promote polyubiquitination. Flag-tagged K48R ubiquitin was coexpressed with wild-type Dvl or its mutant variants into HEK293T cells, followed by immunoprecipitation/immunoblotting for detection of polyubiquitinated forms of Dvl. (e) Polyubiquitin chain of mutant Dvl is preferentially K63-linked. As in (e), but endogenous ubiquitins and anti-K63-linkage-specific Ub antibody (Apu3) were used. (f) Dvl-ΔRDU, Dvl-V667K and Dvl-VK2 had significantly reduced reporter activity compared with wild-type Dvl. Result of the TOPFlash reporter analysis performed in HEK293T cells in triplicates is shown (top). P<0.00002. As transfection and loading control, total lysates subjected to immunoblotting for Dvl using anti-myc and β-actin antibodies are shown (bottom). (g) In contrast to wild-type Dvl, which mainly exists in phosphorylated forms, a significant fraction of Dvl-ΔRDU mutant is not phosphorylated. After transient expression of myc-tagged Dvl-WT and Dvl-ΔRDU mutants, lysates were collected, incubated with λ phosphatase and subjected to immunoblotting with anti-myc antibody. Band for endogenous levels of phosphorylated c-jun (p-c-jun) was abolished after treatment with λ phosphatase (marked with arrowhead). β-actin was used as a loading control.
It has been suggested that the lysine 48-linked ubiquitin chains are involved in proteasomal degradation, whereas lysine 63-linked ubiquitin chains usually have roles in non-degradative signaling pathways.18, 19 Interestingly, increased ubiquitination of Dvl-ΔRDU was observed upon expression of ubiquitin with a point mutation at lysine 48 (Ub-K48R), but not with a point mutation at lysine 63 (Ub-K63R) (Figure 1b). This result suggests that the polyubiquitin chains formed on Dvl-ΔRDU are assembled mainly through K63 linkages. As the C-terminal domain of Dvl regulates Dvl ubiquitination, we dubbed this the ‘RDU’.
To clarify the relationship between the RDU and Dvl ubiquitination, we examined whether the RDU is a ubiquitin binding element. The C-terminus of Dvl (Dvl-CT) was tested together with ubiquitin chains in an in vitro binding assay. Recombinant GST-Dvl-CT was incubated with monoubiquitin, K48-linked polyubiquitin or K63-linked polyubiquitin (Ub2 to Ub7). GST-Dvl-CT significantly bound to K63-linked polyubiquitin (Figure 1c), whereas mono UB and K48-linked polyubiquitin were not pulled down with GST-Dvl-CT. Short K63-linked chains with up to four ubiquitin groups also failed to show Dvl-CT binding in this assay. Overall, these data suggest that the RDU of Dvl binds to K63-linked polyubiquitin chains.
We next attempted to identify residues in the RDU element that are critical for ubiquitin chain binding. Hydrophobic interactions are typically critical for specific ubiquitin recognition.20 We therefore hypothesized that conserved hydrophobic residues Val644 and Val667 of the RDU may be important for the proper conformation of the RDU as well as ubiquitin recognition.21 Indeed, tandem substitution of these residues to lysine (the Dvl-VK2 mutation) strongly impaired recognition of K63 polyubiquitin chains (Figure 1c).
We next introduced the VK2 mutation into full-length Dvl and tested whether it affects modification of the protein by ubiquitin. As with the RDU deletion, we observed a dramatic stimulation of Dvl ubiquitination (Figure 1d). A single substitution, V667K, was sufficient to produce this effect (Figure 1d). The chains observed are striking in length. These results suggest that increased ubiquitination of Dvl may be the outcome of a compromised RDU-ubiquitin interaction. Application of chain-linkage specific ubiquitin antibodies further confirmed that the polyubiquitin chains of these Dvl mutants contained K63 linkages (Figure 1e).
To determine whether deletion of the RDU affects Dvl activity in Wnt signaling, a TOPFlash reporter assay was performed (see Materials and methods). Reporter activity was significantly reduced when cells transiently expressed Dvl-ΔRDU as compared with Dvl-WT (Figure 1f), consistent with the findings of Tauriello et al.16 In addition, Dvl-V667K and Dvl-VK2 showed similar reporter activity as Dvl-ΔRDU (Figure 1f). Phosphorylation of Dvl is necessary for the activation of Wnt signaling.22, 23 We reasoned that the impaired activation of Wnt signaling by Dvl-ΔRDU expression could involve abnormal phosphorylation. Incubation with λ phosphatase resulted in faster migration of Dvl-WT but not Dvl-ΔRDU, suggesting that Dvl-ΔRDU is hypophosphorylated (Figure 1g). Disappearance of phospho-c-jun after incubation with λ phosphatase was used as a positive control for dephosphorylation (Figure 1g). In summary, these data indicate that the RDU of Dvl is essential for the regulation of ubiquitination and phosphorylation of Dvl, both of which promote the activation of downstream Wnt signaling.
Usp14 deubiquitinates K63-linked polyubiquitin chains of Dvl
As the RDU suppresses ubiquitination of Dvl, we hypothesized that the RDU may function in cooperation with an unknown DUB. To identify a DUB that targets the polyubiquitin chains of Dvl, we selected nine candidate DUBs that included potential interaction partners of Dvl from the NCBI database and other DUBs showing impaired Wnt responsiveness upon shRNA-mediated knockdown.15 When Usp14 (designated as DUB6 in Figure 2a) was knocked down by siRNA, strikingly increased Dvl polyubiquitination was observed on endogenous Dvl2 (Figure 2a). A similar result was obtained when cells were treated IU1 (Figure 2b), a small molecule inhibitor of Usp14.24 These results suggest that Usp14 is a key mediator of Dvl deubiquitination. Consistent with these results, overexpression of a catalytically inactive Usp14-C114A mutant (Usp14-CA), which can act as a dominant negative,24 resulted in significant accumulation of polyubiquitinated Dvl (Figure 2c). In summary, the steady-state levels of Dvl ubiquitination are established through a rapid cycle of ubiquitination and deubiquitination in which Usp14 has a dominant role.
Inhibition of Usp14 increases level of polyubiquitinated Dvl. (a) Knockdown of DUB6 (Usp14) increases polyubiquitination of endogenous Dvl. siRNAs for different DUBs or control siGFP were co-transfected with plasmids expressing flag-tagged Ub-K48R into HEK293T cells. Total cell lysates were subjected to immunoprecipitation with anti-Dvl2 antibody or control immunoglobulin G, followed by immunoblotting with anti-flag antibody to detect the ubiquitinated forms of Dvl2. siRNA targeting Usp14 yielded ∼90% knockdown efficiency. Note that the data for DUB9/Usp36 knockdown are excluded in the figure due to severe cell death as previously reported.34 (b) As in a, except that cells were also treated with IU1, an inhibitor of Usp14, at 40 μm for 2 days. Effect of IU1 on Dvl2 polyubiquitination was comparable to that of Usp14 knockdown by siRNA. Dvl2 and Usp14 levels were measured, and endogenous β-actin was used as a loading control. (c) Expression of catalytically inactive Usp14-CA promotes Dvl polyubiquitination. Wild-type and mutant variants of Usp14 were co-transfected with flag-tagged Ub-K48R into HEK293T cells. Protein samples were analyzed by immunoprecipitation/immunoblotting as indicated. (d) As in c, except that Usp14 siRNA was simultaneously introduced along with transient co-expression of Usp14 variants and flag-Ub-K48R. Ectopic expression of wild-type Usp14 and Usp14-ΔUBL was sufficient to reverse Dvl2 ubiquitination induced by Usp14 knockdown. (e) Usp14 is essential for transduction of Wnt signaling. TOPFlash assay was performed in HEK293T cells introduced with either siGFP or siUsp14. At 2 days post transfection of siRNA and reporter plasmids, Wnt3a CM were treated for 0, 4, 8 and 12 h. Data represent average values of relative TOPFlash over FOPFlash ratios from one representative experiment performed in triplicate. Error bars indicate standard deviations in triplicate (*P<0.002). (f) The Usp14-CA mutant inhibits reporter activity induced by Dvl, but not β-catenin. Catalytically inactive Usp14-CA mutant was coexpressed with β-catenin or Dvl, and reporter assays were performed as described in (e) (*P<0.0003). (g) Ubiquitin chain trimming assay. HEK293T cells under Usp14 siRNA conditions were transfected with Ub-K48R only (left) or Ub-K48R and Dvl-ΔRDU (right). At 2 days post transfection, cells were treated with Wnt3a CM for 1 h and whole-cell lysates were collected. Polyubiquitinated species of endogenous and transfected Dvl were enriched by immunoprecipitation using anti-Dvl2 (left) and anti-myc antibodies (right), respectively. Immunoprecipitates were incubated with BSA (marked with ‘-’), recombinant GST-Usp14-WT, or GST-Usp14-CA protein for an additional 1 h. Levels of Dvl ubiquitination were analyzed by SDS–PAGE and immunoblotting with anti-Flag antibody. Levels of GST-Usp14-WT and GST-Usp14-CA were determined by Coomassie Brilliant Blue staining.
Usp14 is a proteasome-associated DUB known to be activated by proteasomes.24 However, the detailed biochemical mechanism of activation is still undetermined. Through both catalytic and non-catalytic mechanisms, Usp14 functions as an endogenous inhibitor of mammalian proteasomes.24, 25 Specifically, the ubiquitin-like (UBL) domain of Usp14 is thought to interact directly with subunit Rpn1 in the proteasome regulatory particle (also called 19S and PA70026, 27). Interestingly, ectopically expressed Usp14-ΔUBL behaved as full-length Usp14 in reversing the effect of Usp14 siRNA on Dvl polyubiquitination, suggesting that the disassembly of ubiquitin chains on Dvl may be a proteasome-independent activity of Usp14 (Figure 2d).
As inhibition of Usp14 activity increased Dvl polyubiquitination, we examined whether knockdown of Usp14 has any effect on Wnt signaling. Significantly induced Wnt reporter activity was observed in control HEK293T cells 8 h after treatment with Wnt3a-conditioned media (CM), whereas Usp14 knockdown cells displayed only slightly increased Wnt reporter activity (Figure 2e). Accordingly, transient overexpression of Usp14-CA reduced Wnt reporter activity (Supplementary Figure S2a). In contrast, overexpression of Usp14-WT or Usp14-ΔUBL had little effect on Wnt signal output (Supplementary Figure S2a), suggesting that Usp14 is not a limiting component in the Wnt pathway in HEK293T cells.
Increased Wnt reporter activity upon ectopic expression of Dvl, but not β-catenin, was reduced by expression of Usp14-CA (Figure 2f). This suggests that regulation of Wnt signaling by Usp14 is controlled at the level of Dvl, rather than downstream components such as β-catenin. This finding is consistent with the effect of Usp14 on Dvl ubiquitination (Figure 2a). To test direct deubiquitination of Dvl by Usp14, we performed a ubiquitin chain trimming assay using immunopurified Dvl-ubiquitin conjugates and recombinant Usp14 in a proteasome-free condition (Figure 2g and Supplementary Figure S2b). HEK293T cells were treated with Usp14 siRNA to knock down endogenous Usp14, followed by transient expression of Ub-K48R. After incubation with Wnt3a CM, whole-cell extracts were collected and endogenous Dvl proteins enriched by immunoprecipitation. Incubation of recombinant Usp14-WT with immunoprecipitates significantly reduced the high-molecular weight species of polyubiquitinated Dvl, whereas catalytically inactive Usp14-CA had little or no effect (Figure 2g). In a similar assay, polyubiquitin chains of Dvl-ΔRDU mutants were hardly susceptible to deubiquitination by Usp14 (Figure 2g), further suggesting that the RDU domain is essential for Usp14 interaction and subsequent deubiquitination.
RDU of Dvl is important for interaction with Usp14
To test for an interaction between Usp14 and Dvl, co-immunoprecipitation assays were performed. HEK293T cells were transfected with Usp14 constructs, and cell extracts were subjected to immunoprecipitation using anti-Dvl antibody. Neither endogenous Usp14, transiently overexpressed Usp14-WT, nor the Usp14-ΔUBL mutant co-immunoprecipitated with endogenous Dvl, regardless of the presence of Wnt3a CM (Figure 3a and Supplementary Figure S3a). This result could potentially be attributed to the rapid dissociation of Usp14 from Dvl when deubiquitination events were completed. However, even catalytically inactive Usp14-CA mutant did not co-immunoprecipitate with Dvl in normal conditions (Figure 3a). Only when cells were incubated with Wnt3a CM did the Usp14-CA mutant show strong interaction with endogenous Dvl (Figure 3a and Supplementary Figure S3a), suggesting that the interaction is stabilized upon activation of Dvl by Wnt. In a time course experiment, significant interaction between Usp14-CA and Dvl was observed from 30 min to 2 h following incubation with Wnt3a CM (Figure 3b). To confirm the colocalization of endogenous Dvl and Usp14 in the presence of Wnt3a CM, indirect immunofluorescent analysis using wild-type murine embryonic fibroblasts (MEFs) was performed. Although there was no obvious membrane localization of Dvl upon Wnt3a CM treatment, colocalized fluorescence signals of endogenous Dvl and Usp14 clearly peaked at 30–45 min upon Wnt3a CM treatment (Supplementary Figure S3b). Therefore, deubiquitination of Dvl by Usp14 may occur via their transient interaction during Wnt signal transduction.
Dvl transiently interacts with Usp14 in the presence of Wnt3a. (a) Usp14-CA was transiently overexpressed in HEK293T cells in parallel with an empty vector control. At 2 days post transfection, cells were cultured in Wnt3a CM or control media for 2 h. Total cell lysates were collected, subjected to immunoprecipitation with anti-Dvl2 antibody or control immunoglobulin G, and analyzed by immunoblotting for Usp14. Dvl2 and Usp14 in total lysates are shown in the bottom panel. (b) Interaction between Dvl and Usp14 occurs at an early time point after treatment with Wnt3a CM. HEK293T cells were transfected with Usp14-CA. At 2 days post transfection, cells were cultured in Wnt3a CM for various times as indicated in the figure followed by immunoprecipitation/immunoblotting. (c) The RDU is necessary for interaction between Dvl and Usp14. HEK293T cells were transfected with plasmids expressing myc epitope-tagged WT-, ΔRDU-, or VK2-Dvl, then cultured in Wnt3a CM for 30 min. Interaction was analyzed by immunoprecipitation using anti-myc antibody.
To determine whether the RDU and its putative ubiquitin binding sites are necessary for the interaction between Dvl and Usp14, a binding assay similar to that in Figures 3a and b was performed using transiently overexpressed rather than endogenous Dvl. Whereas Dvl-WT strongly interacted with Usp14-CA in the presence of Wnt3a CM, Dvl-ΔRDU or Dvl-VK2 displayed no interaction under the same conditions (Figure 3c). Overall, these data suggest that the binding of polyubiquitin chains by the C-terminal domain of Dvl creates a specific conformation necessary for interaction with Usp14 (see below).
Dvl-ΔRDU displays increased ubiquitination on K444 and K451
To map the sites of enhanced ubiquitin modification on Dvl-ΔRDU, we performed electron spray ionization-mass spectrometry. HEK293T cells were co-transfected with myc-Dvl-ΔRDU and Ub-K48R plasmids, followed by a two-step immunoprecipitation as shown in Supplementary Figure S4a. Immunoprecipitated proteins were resolved by SDS–PAGE and extracted from gel slices above the myc-Dvl-ΔRDU band (Figure 4a). We observed ubiquitinated Dvl when region I, but not region II, (denoted in Figure 4a) was used in electron spray ionization-mass spectrometry analysis, which is consistent with the previous finding that K63-linked ubiquitinated forms of Dvl are found in the lower molecular weight region of the gel, just above unmodified Dvl.15 Electron spray ionization-mass spectrometry analysis identified five candidate ubiquitination sites (K50, K285, K387, K444 and K451) (Figure 4b and Supplementary Figure S4b). K50 was previously identified as a principal K63-linked ubiquitination site in CYLD-deficient cells, and K285 has been assigned as a K48-linked ubiquitination site.15 In subsequent experiments, we focused on K444 and K451, which are highly conserved residues within the DEP domain. When these and two other highly conserved lysine residues (413 and 461) in the DEP domain were changed into arginine, the level of ubiquitinated forms of the Dvl-ΔRDU mutant (Dvl-ΔRDU-DEP4KR) was strongly reduced compared with that of Dvl-ΔRDU (Figure 4c). As Dvl-ΔRDU-DEP4KR was only weakly ubiquitinated, we reasoned that knockdown of Usp14 might have little effect on the reporter activity induced by Dvl-DEP4KR. Indeed, the reporter activity induced by Dvl-WT, but not Dvl-DEP4KR, was inhibited by knockdown of Usp14 (Figure 4d). Together, these results strongly suggest that the increased ubiquitination of Dvl-ΔRDU upon knockdown of Usp14 occurs within the DEP domain, particularly on K444 and K451. We then tested whether the reduced reporter activity in response to Wnt3a CM in the presence of siUsp14 (Figure 2e) might be due to blocking of the interaction between Fzd and ubiquitinated Dvl. Interestingly, Dvl-VK2 did not interact with Fzd5 whereas Dvl-DEP4KR was comparable with Dvl-WT in its interaction with Fzd5 (Figure 4e). These data suggest that deubiquitination of Dvl by Usp14 is necessary for the interaction between Fzd and Dvl.
Ubiquitination of K444 and K451 is regulated by Usp14. (a) Isolation of ubiquitinated forms of myc-Dvl-ΔRDU. Plasmids expressing myc-Dvl-ΔRDU and Flag-Ub (K48R) were co-transfected into HEK293T cells. Dvl was enriched by two-step immunoprecipitation, followed by SDS–PAGE/Coomassie Brilliant Blue staining. Gel slices marked (I) and (II) were excised for ESI-MS analysis as described under ‘Materials and methods’. (b) K444 and K451 are ubiquitinated in myc-Dvl-ΔRDU by ESI-MS analysis. Schematic diagram for the candidate ubiquitination sites identified in this analysis (top). ESI-MS spectra for ubiquitinated Dvl on K444 and K451 are presented. (c) Ubiquitination of Dvl-ΔRDU is reduced by the DEP4KR mutations. Dvls (WT, ΔRDU and ΔRDU-DEP4KR) were co-transfected with Flag-Ub-K48R into HEK293T cells. Protein samples were analyzed by immunoprecipitation/immunoblotting as indicated. (d) Usp14 knockdown inhibits reporter activity induced by WT-Dvl but not that induced by Dvl-DEP4KR. (e) Dvl-DEP4KR, but not Dvl-VK2, interacted with Fzd5 as wild-type Dvl. Dvls (WT, VK2 and DEP4KR) were co-transfected with or without HA-Fzd5 into HEK293T cells. Protein samples were analyzed by immunoprecipitation/immunoblotting as indicated.
Attenuation of Wnt signal transduction by Usp14 inhibition
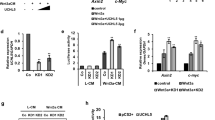
To further test whether Usp14 is necessary for Wnt signaling, we employed MEFs from Usp14−/− mice.28 Compared with wild type, Usp14−/− MEFs displayed weak phosphorylation of endogenous Dvl2 and LRP6 proteins upon Wnt3a CM treatment (Figure 5a). In wild-type MEFs, both Dvl2 and LRP6 became prominently phosphorylated at ∼1 to 2 h following Wnt3a CM treatment, overlapping with the peak period of Dvl-Usp14 interaction (Figure 3b). Similar to the effect of Usp14 knockout, wild-type MEFs treated with IU1 showed significantly reduced LRP6 phosphorylation (Figure 5b). Consistent with these findings, the Wnt3a CM-induced expression of Wnt target genes, such as Nkd1 and Lef1, was significantly impaired in Usp14−/− MEFs or wild-type MEFs treated with IU1 (Figure 5c). To further confirm that Usp14 is important for Wnt pathway function, an axis duplication assay using Xenopus embryos was performed. Consistent with the data obtained using cell culture systems, ectopic expression of WT Usp14 enhanced, whereas Usp14-CA inhibited, Dvl2-induced axis duplication (Figure 5d). Overall, these results suggest that Usp14 has a positive role in the regulation of Wnt/β-catenin signaling. In addition, the delayed phosphorylation of Dvl2 in Usp14−/− MEFs upon Wnt5a CM treatment may indicate that regulation of Dvl polyubiquitination is implicated in non-canonical Wnt signaling pathways as well (Supplementary Figure S5a).
Inhibition of Usp14 delays Wnt signal transduction. (a) Phosphorylation of Dvl2 and LRP6 in response to Wnt is delayed in Usp14−/− MEFs. Wild-type and Usp14−/− MEFs were treated with Wnt3a CM for various times as indicated. Protein samples were collected and analyzed by SDS–PAGE/immunoblotting. Anti-pLRP6 antibodies recognize phospho-Ser1490 of LRP6. (b) Inhibition of Usp14 by IU1 delays Wnt3a-mediated phosphorylation of LRP6. HEK293T cells were treated with IU1 for 4 h and subsequently incubated in Wnt3a CM containing IU1 for various times as indicated. Protein samples were collected and analyzed by SDS–PAGE/immunoblotting. (c) A null mutation in Usp14 attenuates expression of Wnt target genes, Nkd1 (left) and Lef1 (right). Wild-type and Usp14−/− MEFs, and IU1-treated or -untreated wild-type MEF cells were incubated with Wnt3a CM for 6 h before collection of mRNA. Relative mRNA levels of Lef1 and Nkd1 in cells were measured by standard qPCR methods (* indicates P<0.0006). (d) Axis duplication was assessed in Xenopus tadpole embryo. mRNAs for GFP-xDvl2 (500 pg), Usp14 (250 pg), Usp14-CA (250 pg), and Usp14ΔUBL (250 pg) were injected as indicated. Each number of analyzed embryos (n) is indicated. (e) Treatment with Wnt3a induced the level of polyubiquitinated Dvl2 in HEK293T cells when deubiquitination was blocked by ectopic expression of Usp14-CA. Flag-tagged K48R ubiquitin was co-transfected with empty or Usp14-CA constructs in HEK293T cells, followed by treatment with Wnt3a-conditioned media for indicated time periods and immunoprecipitation/immunoblotting to detect polyubiquitinated Dvl2.
As a lack of Dvl deubiquitination by Usp14 appears to attenuate Wnt signaling, we proposed that an elevated forward rate of ubiquitination, which will be counteracted by Usp14 activity, might be induced by Wnt3a CM treatment. We could observe only modestly increased polyubiquitinated Dvl after 2 h Wnt3a CM treatment, perhaps due to active ubiquitination and deubiquitination during this time period (Figure 5e). In contrast, transient expression of inactive Usp14-CA dramatically increased the levels of polyubiquitinated Dvl2 (Supplementary Figure S5b). The enhanced ubiquitination in the presence of Usp14-CA was detected within 30 min after treatment of Wnt3a CM and steadily increased with time (Figure 5e). In summary, our data strongly suggest that Usp14-mediated deubiquitination is necessary for productive signaling upon Wnt3a CM treatment.
Usp14 levels are strongly correlated with β-catenin in human colon cancer
Abnormal activation of the Wnt signaling pathway is known to be linked to the initiation of most colorectal cancers.29 As Usp14 appeared to be a positive regulator of the Wnt signaling pathway, we hypothesized that an elevated level of Usp14 may potentiate β-catenin-mediated colon cancer development. To test this hypothesis, we first examined Usp14 levels in well-characterized colorectal carcinoma cell lines and compared them with those of other immortalized cell lines (Figure 6a). SW480 and LOVO colon cancer cells exhibited significantly increased levels of Usp14 in comparison with wild-type MEFs, HEK293 cells and other cancer cell lines derived from prostate (DU145), ovarian (HeLa) and breast cancer (MCF-7), whereas α7, a subunit of the 26S proteasome, was not increased. DLD1 and Caco2 showed modestly increased or at least similar levels of Usp14 as control cells, whereas HCT116 cells displayed low Usp14 expression. We next examined the level of Usp14 in human colon cancer tissues. Three colon cancer tissues were initially obtained from three individual colon cancer patients, and each colon tissue was divided into normal (N) or tumorous (T) tissues based on histological analysis. All pairs (N/T) from same individuals showed significantly increased Usp14 levels in cancerous colon tissues compared with normal counterparts (Figure 6b). We also observed upregulated Usp14 mRNA levels in the colon cancer tissues (Supplementary Figure S6).
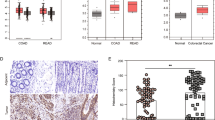
Level of Usp14 is strongly correlated with expression of β-catenin in human colorectal cancer. (a) Endogenous Usp14 levels in colorectal carcinoma and other immortalized cell lines. Each lane was loaded with 20 μg of whole-cell extract. Wild-type and Usp14−/− MEFs were compared as positive and negative controls of Usp14 signaling, respectively. β-actin and α7 were used as loading controls. (b) Comparison of Usp14 protein levels in normal (N) and tumorous (T) human colon tissues. Each pair was derived from an individual colon cancer patient. (c) Immunohistological analysis of Usp14 in colorectal cancer tissue microarray. (a) Immunoreactivity for Usp14 was weak in the cytoplasm of normal colonic mucosal epithelial cells. (b) Colon cancer cells exhibited moderate and strong Usp14 signals in moderately differentiated cancer. (c) Negative Usp14 staining in colon cancer. See Supplementary Table S1 for tissue microarray results from 122 colorectal cancer tissues.
To further test the correlation between Usp14 and β-catenin in colorectal cancer, we performed a tissue microarray using 122 colorectal carcinoma and control tissues. In immunohistochemical analysis, weak Usp14 immunoreactivity was observed in the cytoplasm of corresponding normal colorectal mucosal cells (Figure 6c), and surrounding stromal cells such as fibroblasts were negative for Usp14 immunostaining. Significantly overexpressed Usp14 was detected in 71 (58.2%) of 122 colorectal carcinomas (Figure 6c and Supplementary Table S1). Tumor tissues, even different portions of the same tumor, showed similar expression levels of Usp14, and therefore, no evidence of intratumoral heterogeneity (data not shown). Based on Duke’s stage classification, Usp14 overexpression was categorized into different stages: 8 of 12 cases corresponding to stage A (66.7%), 26 of 47 to stage B (55.3%), 32 of 55 to stage C (58.2%), and 5 of 8 to stage D (62.5%) (Supplementary Table S1). In addition, Usp14 protein was overexpressed in 37 of 61 cases with lymph node metastasis (60.7%). Statistically, there was no relationship between altered Usp14 expression and clinicopathological parameters, including clinical stage (Bartholomew’s test, P>0.05) and lymph node metastasis (χ2 test, P>0.05). However, altered Usp14 levels were closely associated with β-catenin expression in colorectal cancer tissues (P<0.0001) (Supplementary Table S1). This correlation was much stronger than that between Pin1, colon cancer marker and β-catenin.30 Overall, these data suggest that increased expression of Usp14 may enhance β-catenin-mediated transformation of normal colon cells, but not metastasis of malignant colon cancer cells.
Discussion
We report here a novel signaling mechanism in the Wnt pathway, involving K63-linked polyubiquitination of Dvl. These ubiquitin chains are induced by Wnt conditioned medium and rapidly disassembled by a specific DUB, Usp14. Dvl was found to interact with Usp14 in a Wnt-dependent manner to promote rapid disassembly of Dvl-bound, K63-linked chains. Contrary to its established mechanistic role, Usp14 mediates deubiquitination of Dvl without a requirement for its UBL domain. Because the UBL domain of Usp14 is critical for its association with the proteasome, it is likely that free Usp14 is competent to disassemble K63 chains bound to Dvl. Lack of deubiquitination by Usp14 prevented activation of Wnt/β-catenin signaling, based on Dvl and LRP6 phosphorylation, reporter activities, as well as a Xenopus embryonic axis duplication assay, suggesting that Usp14 has a positive role in Wnt/β-catenin signaling. Consistently, the level of Usp14 was strongly correlated with the level of β-catenin in human colorectal cancer.
Based on these observations, we propose a model wherein Usp14-mediated deubiquitination of Dvl regulates the progression of downstream Wnt signaling upon external stimulation by Wnt (Figure 7). Active K63-linked ubiquitination of Dvl occurred with Wnt treatment and did not result in proteasomal degradation (Figure 5e and Supplementary Figure S5b). Therefore, we reasoned that the interaction between the RDU and K63-linked polyubiquitin of Dvl may produce a conformation necessary for interaction with Usp14 (Figure 7). Usp14 deubiquitinates K63-linked polyubiquitin and dissociates from Dvl, which is subsequently phosphorylated in response to Wnt. Properly phosphorylated Dvl then functions as an activator of Wnt signaling. However, deletion of the RDU or mutation of residues that participate in the interaction with polyubiquitin may prevent the conformational change necessary for interaction with Usp14. This could lead to increased Dvl ubiquitination levels as well as reduced phosphorylation. Similarly, knockdown of Usp14 results in increased ubiquitination of Dvl and reduced phosphorylation of Dvl, thereby preventing the activation of downstream Wnt/β-catenin signaling. Therefore, Usp14 functions as a novel regulator of the Wnt signaling pathway.
Model of a novel regulatory mechanism of Wnt signaling via Usp14-mediated Dvl deubiquitination. In the presence of Wnt, K63-linked ubiquitination of Dvl mediated by unknown E3 ligase increases, and the RDU of Dvl binds to the K63-linked ubiquitin chain, which in turn causes conformational changes and allows interaction with Usp14. Usp14 preferentially deubiquitinates K63-linked polyubiquitin chains of Dvl allowing multiple phosphorylation events in Dvl. Deubiquitinated and phosphorylated Dvl interacts with Fzd, thereby activating downstream Wnt signaling.
Consistent with our model, the level of Usp14 was elevated in human colorectal cancer tissues displaying high levels of β-catenin (Supplementary Table S1). The mechanistic cause of this elevation is yet to be determined and it needs to be determined whether increased Usp14 mRNA expression promotes or simply accompanies the progression of colon cancer. However, it is clear that increased Usp14 reinforces β-catenin-mediated Wnt signaling, which may enhance the onset of cancer formation.
Usp14rrk114/rrk114 (considered as Usp14−/− in our manuscript) mice display a male-specific sterility defect as well as a neurologic phenotype culminating in widespread paralysis and premature death (Crimmins, 2009) #228,. As Usp14rrk114/rrk114 is not Usp14-null mice and the knockout phenotypes of different Wnts are so diverse (See Wnt home page, http://www.stanford.edu/group/nusselab/cgi-bin/wnt/), it is difficult to identify obvious similarity between Usp14−/− and Wnt knockout mice phenotypes. In addition, the fact that Usp14−/− MEFs showed delayed Wnt signal transduction (Figures 5a and b), rather than complete blocking, suggests that there are certain functional redundancies shared by other deubiquitinase(s).
The ubiquitin ligases that induce K48-linked ubiquitination of Dvl are known, whereas the E3 ligase that controls K63-linked ubiquitination has not been identified. Gao et al.14 showed that von Hippel–Lindau induces polyubiquitination of Dvl, which in turn leads to autophagosomal degradation.14 As they show that p62, which is known to prefer K63-linked polyubiquitinated substrates,31, 32 recognizes ubiquitinated Dvl for autophagic degradation, von Hippel–Lindau may catalyze K63-linked polyubiquitination of Dvl. Identification of the E3 ligase for K63-linked polyubiquitination of Dvl in Wnt signaling will be of interest, as it may be a useful therapeutic target for diseases caused by misregulation of Wnt signaling.
Here, we reveal a novel role and regulatory mechanism for Usp14. It has been shown that Usp14 is involved in the processing of polyubiquitinated substrates bearing either pure K63 chains or mixed linkage types in a manner that depends on proteasome association.24 However, our data suggest that Usp14 deubiquitinates K63-linked polyubiquitin chains in vivo and in vitro without association with proteasome (Figure 2 and Supplementary Figure S2b). We propose that an unknown modification of Usp14 in the context of Wnt signaling and specific conformation of Dvl caused by K63-linked ubiquitination, permit Usp14 to interact with Dvl and have deubiquitinase activity without association of proteasome. This hypothesis may explain why no Usp14-interacting partners other than the proteasome have been identified. Examination of the nature of Usp14 modification in the presence of Wnt and identification of enzymes responsible for the modification will provide a framework for the study of inducible regulation of Usp14.
High expression levels of Usp14 in colorectal cancer (18 of 99 (18.9%)) have been shown to be associated with liver and lymph node metastasis.33 Our data from tissue microarray indicated that a much higher proportion of colorectal carcinomas have high levels of Usp14 (71 of 122 (58.2%)) (Figure 6c and Supplementary Table S1). More significantly, most colorectal carcinomas that had a high level of Usp14 also showed a high level of β-catenin (63 of 71 (88.7%)). We suggest that Usp14 enhances the responsiveness to external Wnt signal, thus increasing the β-catenin level, which then serves as an initial hit for the oncogenic transformation of colorectal cells. Therefore, it will be interesting to examine whether the level of Usp14 can be used as a prognostic marker and therapeutic target for other Wnt-dependent types of cancer.
Materials and methods
All material and methods are described in Supplementary Information.
References
Logan CY, Nusse R . The Wnt signaling pathway in development and disease. Annu Rev Cell Dev Biol 2004; 20: 781–810.
Clevers H . Wnt/beta-catenin signaling in development and disease. Cell 2006; 127: 469–480.
Reya T, Clevers H . Wnt signalling in stem cells and cancer. Nature 2005; 434: 843–850.
MacDonald BT, Tamai K, He X . Wnt/beta-catenin signaling: components, mechanisms, and diseases. Dev Cell 2009; 17: 9–26.
Clevers H, Nusse R . Wnt/beta-catenin signaling and disease. Cell 2012; 149: 1192–1205.
Wallingford JB, Habas R . The developmental biology of Dishevelled: an enigmatic protein governing cell fate and cell polarity. Development 2005; 132: 4421–4436.
Zeng X, Tamai K, Doble B, Li S, Huang H, Habas R et al. A dual-kinase mechanism for Wnt co-receptor phosphorylation and activation. Nature 2005; 438: 873–877.
Pan W, Choi SC, Wang H, Qin Y, Volpicelli-Daley L, Swan L et al. Wnt3a-mediated formation of phosphatidylinositol 4,5-bisphosphate regulates LRP6 phosphorylation. Science 2008; 321: 1350–1353.
Veeman MT, Axelrod JD, Moon RT . A second canon. Functions and mechanisms of beta-catenin-independent Wnt signaling. Dev Cell 2003; 5: 367–377.
Gao C, Chen YG . Dishevelled: The hub of Wnt signaling. Cell Signal 2010; 22: 717–727.
Tauriello DV, Maurice MM . The various roles of ubiquitin in Wnt pathway regulation. Cell Cycle 2010; 9: 3700–3709.
Jung H, Kim HJ, Lee SK, Kim R, Kopachik W, Han JK et al. Negative feedback regulation of Wnt signaling by Gbetagamma-mediated reduction of Dishevelled. Exp Mol Med 2009; 41: 695–706.
Zhang L, Gao X, Wen J, Ning Y, Chen YG . Dapper 1 antagonizes Wnt signaling by promoting dishevelled degradation. J Biol Chem 2006; 281: 8607–8612.
Gao C, Cao W, Bao L, Zuo W, Xie G, Cai T et al. Autophagy negatively regulates Wnt signalling by promoting Dishevelled degradation. Nat Cell Biol 2010; 12: 781–790.
Tauriello DV, Haegebarth A, Kuper I, Edelmann MJ, Henraat M, Canninga-van Dijk MR et al. Loss of the tumor suppressor CYLD enhances Wnt/beta-catenin signaling through K63-linked ubiquitination of Dvl. Mol Cell 2010; 37: 607–619.
Tauriello DV, Jordens I, Kirchner K, Slootstra JW, Kruitwagen T, Bouwman BA et al. Wnt/beta-catenin signaling requires interaction of the Dishevelled DEP domain and C terminus with a discontinuous motif in Frizzled. Proc Natl Acad Sci USA 2012; 109: E812–E820.
Heessen S, Masucci MG, Dantuma NP . The UBA2 domain functions as an intrinsic stabilization signal that protects Rad23 from proteasomal degradation. Mol Cell 2005; 18: 225–235.
Pickart CM . Ubiquitin enters the new millennium. Mol Cell 2001; 8: 499–504.
Komander D, Rape M . The ubiquitin code. Annu Rev Biochem 2012; 81: 203–229.
Beal RE, Toscano-Cantaffa D, Young P, Rechsteiner M, Pickart CM . The hydrophobic effect contributes to polyubiquitin chain recognition. Biochemistry 1998; 37: 2925–2934.
Kozlov G, Nguyen L, Lin T, De Crescenzo G, Park M, Gehring K . Structural basis of ubiquitin recognition by the ubiquitin-associated (UBA) domain of the ubiquitin ligase EDD. J Biol Chem 2007; 282: 35787–35795.
Bryja V, Schulte G, Rawal N, Grahn A, Arenas E . Wnt-5a induces Dishevelled phosphorylation and dopaminergic differentiation via a CK1-dependent mechanism. J Cell Sci 2007; 120 (Pt 4): 586–595.
Price MA . CKI, there's more than one: casein kinase I family members in Wnt and Hedgehog signaling. Genes Dev 2006; 20: 399–410.
Lee BH, Lee MJ, Park S, Oh DC, Elsasser S, Chen PC et al. Enhancement of proteasome activity by a small-molecule inhibitor of USP14. Nature 2010; 467: 179–184.
Lee MJ, Lee BH, Hanna J, King RW, Finley D . Trimming of ubiquitin chains by proteasome-associated deubiquitinating enzymes. Mol Cell Proteomics 2011; 10: R110 003871.
Leggett DS, Hanna J, Borodovsky A, Crosas B, Schmidt M, Baker RT et al. Multiple associated proteins regulate proteasome structure and function. Mol Cell 2002; 10: 495–507.
Rosenzweig R, Bronner V, Zhang D, Fushman D, Glickman MH . Rpn1 and Rpn2 coordinate ubiquitin processing factors at proteasome. J Biol Chem 2012; 287: 14659–14671.
Crimmins S, Sutovsky M, Chen PC, Huffman A, Wheeler C, Swing DA et al. Transgenic rescue of ataxia mice reveals a male-specific sterility defect. Dev Biol 2009; 325: 33–42.
Medema JP, Vermeulen L . Microenvironmental regulation of stem cells in intestinal homeostasis and cancer. Nature 2011; 474: 318–326.
Kim CJ, Cho YG, Park YG, Nam SW, Kim SY, Lee SH et al. Pin1 overexpression in colorectal cancer and its correlation with aberrant beta-catenin expression. World J Gastroenterol 2005; 11: 5006–5009.
Long J, Gallagher TR, Cavey JR, Sheppard PW, Ralston SH, Layfield R et al. Ubiquitin recognition by the ubiquitin-associated domain of p62 involves a novel conformational switch. J Biol Chem 2008; 283: 5427–5440.
Wooten MW, Geetha T, Babu JR, Seibenhener ML, Peng J, Cox N et al. Essential role of sequestosome 1/p62 in regulating accumulation of Lys63-ubiquitinated proteins. J Biol Chem 2008; 283: 6783–6789.
Shinji S, Naito Z, Ishiwata S, Ishiwata T, Tanaka N, Furukawa K et al. Ubiquitin-specific protease 14 expression in colorectal cancer is associated with liver and lymph node metastases. Oncol Rep 2006; 15: 539–543.
Endo A, Matsumoto M, Inada T, Yamamoto A, Nakayama KI, Kitamura N et al. Nucleolar structure and function are regulated by the deubiquitylating enzyme USP36. J Cell Sci 2009; 122 (Pt 5): 678–686.
Acknowledgements
We thank Kay Hofmann (University of Cologne) for his assistance with bioinformatics. This work was supported by funding from the National Research Foundation of Korea (2010-0021106 and 2012R1A2A2A01012472 to E-h. J. and 2012-0003217 to M.J.L.) and by the National Institutes of Health grant GM095526 (DF). HJ was supported by the Brain Korea 21 program.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogenesis website .
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Jung, H., Kim, BG., Han, W. et al. Deubiquitination of Dishevelled by Usp14 is required for Wnt signaling. Oncogenesis 2, e64 (2013). https://doi.org/10.1038/oncsis.2013.28
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/oncsis.2013.28