Abstract
Melanoma is a heterogeneous tumor with different subpopulations showing different proliferation rates. Slow-cycling cells were previously identified in melanoma, but not fully biologically characterized. Using the label-retention method, we identified a subpopulation of slow-cycling cells, defined as label-retaining cells (LRC), with strong invasive properties. We demonstrate through live imaging that LRC are leaving the primary tumor mass at a very early stage and disseminate to peripheral organs. Through global proteome analyses, we identified the secreted protein SerpinE2/protease nexin-1 as causative for the highly invasive potential of LRC in melanomas.
Similar content being viewed by others
Introduction
Melanoma accounts for ~4% of all skin cancer cases but for >70% of all skin cancer deaths. Although tumor development and progression, from benign proliferative lesions to dysplasia to overt malignancy, typically follow a stepwise pattern, metastatic dissemination remains difficult to predict and treat because of the heterogeneity in this disease. Heterogeneity manifests both at the genetic and epigenetic/phenotypic levels. Of all human cancers, melanomas have the highest mutational load with hundreds to thousands of mutations per tumor. While the field has identified ~20 driver mutations in this disease,1, 2, 3, 4, 5, 6, 7, 8, 9 the different combinations of driver mutations that are possible make it challenging to identify genetic subgroups for outcome prediction of continuous growth, invasion and dissemination, or, in advanced metastatic disease, immunotherapy or targeted therapy.
Heterogeneity of melanoma is displayed not only between tumors but also within tumors. Differences manifest in growth rates, pigmentation, differentiation states to other cell types, structural cellular elements, matrix formation, or host cellular infiltrates. While most cancers contain unique cell populations generally described as cancer-initiating cells, melanoma does not follow this pattern. Instead, melanomas contain subpopulations whose presence is dynamically regulated by microenvironmental signals. We previously defined a slowly-cycling subpopulation in melanoma that is important for tumor maintenance and drug resistance. This subpopulation, which proliferates at a much slower rate than the main population, was defined by expression of the H3K4 demethylase JARID1B10 and represents only 0.5 to 5% of all cells. It is regulated by hypoxia, growth factors, and cytokines such as IGF-1 or TNF-α.10, 11, 12 Expression of the transcription factor microphtalmia-associated transcription factor (MITF) has been used as biomarker to distinguish populations of melanoma cells that are highly proliferative and poorly invasive (MITFHigh) or slowly proliferative and highly invasive (MITFLow).13, 14, 15 MITF is a master regulator for melanocytes and melanoma cells and essential for the survival of all pigmented cells whereas JARID1B expression characterizes a subpopulation within slow-cycling melanoma cells, since not all the slow-cycling melanoma cells express JARID1B. We focused in the presented experiments on slow-proliferating melanoma cells that included both JARID1B-posive and -negative melanoma cells.
All cells can take up a membrane-staining dye, but after two to three cell divisions the dye is diluted and becomes undetectable by fluorescence microscopy whereas label-retaining cells (LRC) retain the dye for longer periods of time, which defines melanoma slow-cycling cells as biologically LRC. Similar dye-retaining experiments are well described for stem cells with high turnover of the main population. For example, epithelial stem cells in the skin were defined by retention of bromo-deoxy-uridine.19, 20
Here we characterize the invasive properties of slowly-cycling melanoma cells both in vitro and in vivo. We demonstrate that human melanoma LRC are highly invasive and disseminate more rapidly in immunodeficient mice to lymph nodes, lungs, liver and spleen than non-LRC. After dissemination, significantly larger metastases are observed in LRC-injected mice. We then identified a secreted protein, SerpinE2/Protease nexin-1 (PN-1) or glia-derived nexin, as causative for the highly invasive potential of LRC. SerpinE2 is a member of the Serpin gene superfamily with anti-serine protease activity and acts as major regulator for both tissue-type and urokinase plasminogen activators.
Results
Melanoma LRC are slow-cycling and invasive in vitro
Slow-cycling melanoma cells have been identified by us and others10, 21, 22 but knowledge of their biological properties has been limited to growth and drug resistance. When cultured cells from human metastatic melanomas are labeled with membrane dyes such as CellTrace Violet (Molecular Probes, Eugene, OR, USA) or PKH26, they dilute the dyes with each cell division. Thus, only non-proliferating or slow-dividing cells retain the label for prolonged periods of time, defining this subpopulation as LRC. In vitro, using CellTrace Violet, LRC were found in all metastatic melanoma cell lines analyzed (Figures 1a and b). In each cell line this LRC subpopulation emerged after 4 days in culture and could be detected for up to 3 weeks, at which point the membrane dye could no longer be detected due to cell divisions (Figure 1b; Supplementary Figure 1A). Their presence was not associated with any genetic subgroups of melanomas such as those representing mutant BRAF, NRAS or BRAF/NRAS wild type. Of all cell surface markers previously used to define melanoma-initiating cells, including CD20, CD271 or CD13322, 23, 24 we saw no distinct expression patterns in the LRC populations except that CD20 was upregulated in LRC of 4/8 melanoma cell lines (Supplementary Figure 1B). Most, but not all, LRC express the H3K4 demethylase JARID1B,10 which we confirmed at the single-cell level by mRNA fluorescence in situ hybridization (FISH; Figure 1d). LRC are enriched in G2/M of the cell cycle (Figure 1c; Supplementary Figure 1C) and no differences were observed between LRC and non-LRC in S phase (Supplementary Figure 1D). LRC are highly invasive through basement membrane matrix, Matrigel® (Corning, Tewksbury, MA, USA), when tested in Boyden chamber assays. Differences in invasiveness varied among cell lines with LRC being 2- to 8-fold more invasive than their respective non-LRC populations (Figure 1e; Supplementary Table 6).
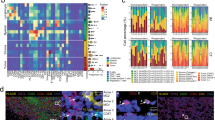
Melanoma cells identified as slow-cycling, LRC are predominantly in the G2/M cell cycle phase and are more invasive in vitro. (a) Six melanoma cell lines were labeled with CellTrace Dye and analyzed at different time points (days 0–4–8–12). Day 0 indicates how the cells appear directly after CellTrace loading. LRC are defined and gated as shown in the dot plots; numbers indicate % of LRC detected on day 12 in culture; standard error of the mean (s.e.m.) in parenthesis. (b) Relative dilution of the CellTrace Dye for LRC (red circles), non-LRC (yellow inverted triangles) and total cells (gray squares). Slopes indicate mean fluorescence intensity ratio between CellTrace-stained and unstained cells. Mean±s.e.m. are reported. (c) Cells were loaded with CellTrace Dye and stained with propidium iodide for cell cycle analyses after 7 days in culture. Percentage of cells in each cell cycle stage is shown in the Y-axes. Data represent mean±s.e.m. Three representative cell lines are shown. (d) LRC (red), non-LRC (orange) and total (gray) were sorted and analyzed by mRNA FISH for JARID1B expression in single cells, with representative images below (total magnification 100 ×). Data indicate that some but not all LRC are JARID1B-positive. Violin plots depict mRNA counts (Y-axes); ns=not significant. (e) Invasive capacity of LRC (red bars) and non-LRC (orange) in Boyden chamber assays after sorting, for three representative cell lines. Data represent mean±s.e.m. of three independent experiments of the fold change of LRC over non-LRC. All reported P-values were determined using a 2-sided t-student test.
Melanoma LRC exit the primary tumor mass and disseminate in mouse tissues
We then investigated in vivo the growth and invasive dynamics of LRC originally isolated from metastatic lesions. Figure 2a provides a schematic for the experiment, in which we labeled melanoma cells in vitro with the membrane dye PKH26 and injected 106 cells subcutaneously into SCID Hairless Outbred (SHO®; Charles River Laboratories, Wilmington, MA, USA) mice. PKH26 was chosen because the fluorescence allows scanning of labeled cells by IVIS® (PerkinElmer, Waltham, MA, USA) in vivo. SHO mice were selected because they are hairless and have thin skin, allowing better scanning of internal organs than other mouse strains with fur. We monitored the mice weekly for fluorescent signals by placing them in dorsal, ventral, and lateral positions to test the hypothesis that the normally proliferating cells at the injection site (primary tumor) would rapidly lose the fluorescent signal, whereas LRC would readily disseminate, retain the signal, and remain visible at a distant site. Indeed, PKH26 was detected in the primary tumors only at the beginning of the experiment and it was diluted when tumors expanded in size (Figures 2b and c). Demonstrating the consistency of the technique, the PKH26 dilution followed the same kinetics, whether tumors grew faster (451Lu) or slower (WM989). The dye was no longer detectable when tumors reached ~500 mm3 in size.
Melanoma LRC disseminate in vivo early from the primary tumor site. (a) Schematic for detecting LRC in vivo. Melanoma cells (WM989 and 451Lu) were labeled with PKH26 and subcutaneously injected into SCID Hairless Outbred mice. Mice were imaged weekly using IVIS (dorsal, ventral and lateral scanning). Between days 0 and 20, the fluorescent signal was detected in the primary tumor at the injection site; after day 20, proliferating tumor cells lost the dye at the primary site. Instead, fluorescent signals indicating the presence of LRC were detected in distant sites. (b) Mice on left: dorsal view of mice scanned 7 days post-tumor injection. For the left two mice of each melanoma, palpable tumors overlapped with fluorescence signals. Graphs on right: tumor growth (black line, left Y-axis) and dilution of PKH26 signal (red line, right Y-axis) at the primary tumor site (n=5). Arrows indicate the cross point of the curves. After reaching ~500 mm3 tumor size, PKH26 signal is rapidly lost. (c) Dorsal (left), ventral (middle), and lateral (right) views of mice scanned when tumors reached ~700 mm3 size. (d) Scatter plots showing quantification of the fluorescent signals for individual mice at days 0–20 (left plots) and 20–60 (right plots) post injection (n=5), for the same two melanoma cell lines as in c. Primary tumors (blue), peritoneal cavities (red), and left flank (black). (e) Bars represent flow cytometry quantification of melanoma LRC (CD146+/PKH26+ positive cells) in primary tumor (gray), mouse lymph nodes (LN, yellow), lungs (blue), liver (green) and spleen (purple); Zebra plot (below) are representative examples from lungs. Upper right quadrant: melanoma LRC double positive for CD146 and PKH26; lower right quadrant: melanoma non-LRC cells (CD146+ positive). Numbers are % of positive cells after subtraction of the percentage detected in the isotype control (left plot). (f) Representative photomicrographs of immunofluorescence performed on frozen tissue from lungs and spleen for a combination of melanoma markers (CD146, β3 integrin, MSCP-1). Cells can be detected as clusters (white square in spleen) or disseminated (white arrows). Bottom panel is the negative antibody control staining.
Over time, as the primary tumor grew, the signal disappeared from it, but appeared at distal locations. Signal was detected in mice injected with WM989 cells and scanned in the ventral (5/5 positive) and lateral (4/5 positive) positions; for 451Lu cells, 2/5 mice were positive when imaged in the ventral position (Figures 2c and d). When ND238 cells were injected into 10 mice, 8/10 showed positive signal in their peritoneal cavities (Supplementary Figure 2A). Similarly, after injection of WM3942 cells, we detected positive PKH26 signal in 4/4 mice left flank and in 4/4 peritoneal cavities (Supplementary Figure 2B). Thus in all 4 models analyzed we could observe the same general dissemination LRC. When the primary tumors reached ~1000 mm3 in size, mice were sacrificed and tumors and organs (lungs, liver, spleen and detectable lymph nodes) were harvested and analyzed by flow cytometry for melanoma LRC. Confirming that LRC migrated from the primary tumor, we found that the percentage of LRC (mouse lineage negative, CD146+, PKH26+) was higher in all distal organs compared with the primary tumor (Figure 2e; Supplementary Figure 2C). A representative example is shown in the Zebra plots of Figure 2e, with LRC represented by the upper right quadrants. Results were confirmed by immunohistochemistry of tissue sections from lungs and spleen (Figure 2f). We were able to monitor LRC in the same location for the same animal for up to 20 (5–20) days after the initial detection of signal. These observations indicate that the majority of LRC do not begin proliferation immediately after entering an organ but can remain dormant for extended time periods (Supplementary Figure 2D).
To directly compare the ability of LRC and non-LRC to disseminate, we sorted LRC from non-LRC and injected s.c. both populations into different recipient mice. Sorted LRC and non-LRC rapidly re-equilibrated their parental redistribution in vitro (Supplementary Figure 3A). This dynamic is well established for JARID1B (Roesch 2010) and we believe it occurs also in vivo. The total metastatic burden (measured as metastatic spots detected by live imaging) was higher for mice injected with LRC than non-LRC (Supplementary Figure 3B). Representative images of mouse lungs showed larger metastatic areas for those from LRC than those non-LRC (Supplementary Figure 3C). The ability to regain proliferative capacity of LRC is underscored by the observation that LRC and non-LRC generate primary tumors in recipient mice with comparable growth rates (Supplementary Figure 4A). Moreover, a majority of cells in LRC-derived xenografts is positive for the Ki67 proliferation marker (Supplementary Figure 4B). Injection of cells labeled with luciferase, allowed us to image organs at long delayed time point, and these experiments confirming the presence of growing metastatic lesions in distant organs (representative image in Supplementary Figure 5A).
Proteome analyses reveal upregulation of SerpinE2, PDGFRL and BMP1 in LRC
To understand the invasive phenotype of slowly-proliferating cells, we performed mass spectrometry based comparative and semi-quantitative proteomics using tryptic digest of LRC and non-LRC from five different melanoma cell lines by mass spectrometry. From this extensive analysis, we selected for further study proteins preferentially expressed in LRC over non-LRC. Most consistently overexpressed in LRC, across all cell lines, were three proteins, SerpinE2, platelet-derived growth factor receptor-like (PDGFRL) and bone morphogenetic protein 1 (BMP1; Figure 3a; Supplementary Tables 1–5). We validated only SerpinE2 and BMP1 because these proteins were more likely to be relevant for cancer cell aggressiveness (and limited tools were available to study PDGFRL). We confirmed that SerpinE2 and BMP1 are expressed at higher levels in LRC than non-LRC by single-cell mRNA FISH and qRTPCR on LRC and non-LRC total cells (Figure 3b; Supplementary Figure 6A) and by immunofluorescence (Figure 3c). To ensure that SerpinE2 and BMP1 expression was not an in vitro artifact, we validated expression in primary tumor sites and distal metastatic organs by immunofluorescence and flow cytometry (Figure 3d; Supplementary Figure 6B). We additionally observed that human SerpinE2 expression is increased in mouse spleen compared with primary tumors (Supplementary Figure 6C). We then investigated whether SerpinE2 expression is associated with the slow-cycling phenotype in melanoma and whether it correlated with expression of JARID1B and MITF. SerpinE2 is enriched ~6-fold in slow-cycling cells identified as double positive for JARID1B and PKH26 compared with fast proliferating cells (JARID1B-/PKH26-; Supplementary Figure 6D). At the single-cell level SerpinE2 expression is positively correlated to JARID1B expression in the total LRC and non-LRC fractions (Supplementary Figure 6E). Conversely, there is no correlation between SerpinE2 and MITF expression, neither in LRC or non-LRC (Supplementary Figure 6F). This suggests that SerpinE2 and JARID1B are partially overlapping markers and enriched in LRC. Their expression is nevertheless not exclusive for LRC. Non-LRC can express JARID1B and SerpinE2, albeit at significant lower levels. We then focused on SerpinE2 as a candidate for highly invasive LRC.
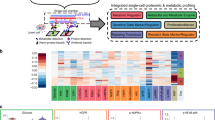
Label-retaining, disseminated melanoma cells express high levels of SerpinE2. (a) Table lists proteins identified after whole-cell proteomic analyses of LRC and non-LRC. The last column lists the cell lines with differential expression of the indicated protein. (b) Violin plots (left panels) illustrate quantification of SerpinE2 (white spot, top) and BMP1 (white spots, bottom) as determined by mRNA FISH. mRNA counts for single cells are shown on the Y-axes. Note differences in Y-axes. Representative photomicrographs are shown on the right (total magnification: 100 ×). (c) Protein expression was analyzed by immunofluorescence and quantified at single-cell levels (scatter plots on left and representative images on right). LRC (red), non-LRC (orange), total cells (gray). (d) Flow cytometry of disseminated WM989 melanoma cells detected in three mouse organs (top panel). The percentage of cells double positive for CD146 and SerpinE2 is shown in the upper right quadrants; the lower right quadrants indicate the percentage of cells singly positive for CD146. Micrographs show examples of disseminated SerpinE2-positive cells (arrows, middle and right panels) in mouse lungs. Left panel: negative control.
SerpinE2 is critical for melanoma invasion and its expression level correlates with tumor progression
To investigate the role of SerpinE2 in melanoma invasion, we sorted melanoma LRC and non-LRC and performed an invasion assay overnight in the presence of human recombinant SerpinE2. In the presence of exogenous SerpinE2, non-LRC gained the ability to invade, whereas the invasive capacity of LRC was not significantly affected (Figure 4a; Supplementary Figure 7A; Supplementary Table 7).25 For validation, we blocked the anti-proteolytic activity of SerpinE2 with a specific monoclonal antibody.26 Neutralization of SerpinE2 reduced melanoma invasion in a dose-dependent manner in all four models tested (Figure 4b; Supplementary Figure 7B). SerpinE2 knockdown with shRNA (Supplementary Figure 8; Supplementary Table 8) also confirmed the inhibition of invasion (Figure 4c). Notably, invasion was affected only when successful knockdown was achieved (Sh_13-16), and it was not impaired in the presence of non-effective Sh_17 (Figure 4c; Supplementary Figure 8). For further analyses, we chose Sh_14 and Sh_16, showing most effective knockdown across all cell lines tested (Supplementary Figure 8). SerpinE2 knockdown specifically affected melanoma invasion, without impairing cell proliferation or viability (Supplementary Figure 9).
SerpinE2 drives melanoma invasiveness. (a) Boyden chamber invasion assay of sorted label-retaining cells (LRC) or non-LRC in presence (blue columns) or absence (gray columns) of human recombinant SerpinE2. Bars show fold change in invasion (mean±s.e.m.) of samples with SerpinE2 over untreated cells. (b) Invasion assay in presence of an anti-human SerpinE2 neutralizing antibody or Isotype control (ctrl). The percentage of invading cells after neutralization is shown. Data represent mean±s.e.m. of three independent experiments. (c) Invasion assay after SerpinE2 knockdown. Data shown are percentage of invading cells found after silencing with 5 different shRNA (Sh_13-17) relative to Sh_ctrl (ctrl). Bars represent mean±s.e.m.
SerpinE2 is nearly absent in supernatants from the five normal melanocyte cultures (MC) tested, but is present in supernatants from all seven melanoma lines tested (WM; Figure 5a). The 2D finding was confirmed in 3D skin reconstructs, with only melanoma cells staining positive for SerpinE2 (Figure 5b). SerpinE2 mRNA expression is absent in normal skin as shown by analysis of a public data set (GEM_1375) and increases with malignant progression (Figure 5c). Immunohistochemistry analyses on patient tissues confirmed the association between SerpinE2 expression and melanoma progression, with the strongest expression in invasive and tumorigenic vertical growth phase (VGP) primary tumors and metastatic lesions (Figures 5d and e). Benign lesions (nevi) showed a dotted expression pattern (Figure 5d, arrows) typical of Golgi-associated localization, which switches to extracellular staining in malignant lesions, supporting the involvement of secreted SerpinE2 in invasive progression. Intriguingly, the variability of SerpinE2 expression in VGP reflects the known cellular heterogeneity of this type of lesion,27, 28 whereas metastatic lesions show more uniform staining, suggesting a possible selection of SerpinE2-high expressing cells during metastasis formation.
SerpinE2 expression is increased in malignant cells and correlates with tumor progression. (a) Western blot analysis of SerpinE2 secretion by melanoma cells (n=21) and human melanocytes (n=5) into culture supernatants. Recombinant SerpinE2 protein (500 ng) was used as a positive control and Ponceau’s staining is shown for loading control. Graph shows quantification (right panel). (b) SerpinE2 staining of 3D skin reconstructs with melanocytes (left) and melanoma cells (right). (c) GEM_1375 data set analysis of SerpinE2 mRNA expression in melanomas, non-malignant nevi and normal skin. (d) Examples of immune histochemistry at two different magnifications of: normal skin, benign nevi, in situ, vertical growth phase (VGP) and lymph node metastatic melanomas (n=15 for each group). SerpinE2 protein expression is indicated by purple staining. Arrows indicate the dotted SerpinE2 expression pattern found in benign nevi only. (e) Intensity score of SerpinE2 expression in normal skin, benign nevi and malignant melanomas (radial growth phase (RGP), VGP and metastatic). Bars represent mean±SEM of tissue sections of 15 specimens from each group; P-value after t-student test are given.
Discussion
We describe a subpopulation of human melanoma cells that proliferates much slower than the main tumor population but is highly invasive, both in vitro and in vivo. This subpopulation does not correlate with any genetic signatures of melanoma cells such as the status of BRAF, NRAS or PTEN mutations. We defined this subpopulation as LRC because of the dye-labeling technique used for their identification. After injection into immunodeficient mice, LRC migrate from the primary tumor to distal sites (spleen, lungs and lymph nodes); this migration could be visualized in vivo for ~20 days.
While proliferation is considered a hallmark of aggressive cancers, we demonstrate here that slow-proliferating cells are more invasive and thus the most dangerous cells (Figures 1 and 2; Supplementary Figure 2). Our findings are in-line with reports from cultured tumor cells that LRC have increased invasive potential, for example in pancreatic adenocarcinoma,29 and that non-proliferative, senescent-like melanoma cells can be invasive in vitro.30, 31 The secreted protein SerpinE2 strongly augments LRC’s invasive potential. The identification and characterization of this slowly-proliferating melanoma subpopulation urges us to begin adjusting our treatment strategies of biologically early disease, because we may not detect individual, disseminated tumor cells with current diagnostic techniques.
Invasion and dissemination are the first steps in the metastatic cascade, with many cells able to survive when circulating in the lymph and blood vascular systems, but only a small percentage is able to initiate growth at the new site. Understanding the biological events at the distal sites is critical for being able to prevent or slow metastases. We observed that some LRC in peripheral organs remained detectable for days without any dye lost, i.e., they remained dormant (Supplementary Figure 2D).
This dynamic resembles clinical patterns, where tumor cells that disseminate early can remain dormant for extended time periods but then (via unknown mechanisms) regain proliferative potential and give rise to clinically detectable tumor masses.
Proliferation at the primary tumor site is associated with metastasis formation in the regional lymph node,32 but melanoma can recur much later (>5 years) in patients with no/low proliferation in their primary tumors.33 Even very late recurrences (>10 years), observed in approximately 7% of melanoma patients, can be associated with negative sentinel lymph node biopsy,34 suggesting that dissemination has occurred through the vascular system and that the disseminated cells have remained dormant for extended times. The mechanisms driving the switch between dormant state and metastasis development are not clear, but likely result from cues in the microenvironment. The clinical implication of the LRC phenotype is that if some cells are able to disseminate in the first phases of tumor growth, they will escape surgical excision, which is the predominant treatment for primary melanomas. Therefore, separate strategies are needed to target these cells for achieving a complete eradication of the tumor.
Using the unbiased approach of directly comparing proteomes from total extracts of matched LRC and non-LRC, we identified SerpinE2, BMP1 and PDGFRL as potential markers for melanoma LRC (Figure 3; Supplementary Figure 6; Supplementary Tables 1–4). As PDGFRL has been poorly studied and no antibodies were available, this protein was not investigated further. BMP1 is a member of the bone morphogenic protein family that is normally secreted during cartilage development and has a major role in proto-collagen and proto-laminin 5 cleavage.35–37 In cancer development and progression, BMP1 is associated with metastasis in non-small cell lung cancer38 and structurally analogous members are associated with colon cancer progression,39 likely via Src activation,40 but in general the role of BMP1 in cancer is not well defined.
SerpinE2 is secreted by nerve and vascular cells in physiological conditions and by platelets upon activation at tissue injury sites.41 It has anti-serine protease activity to many proteases but its main function is to regulate the plasminogen-plasmin axis, functioning as a plasminogen activator inhibitor.42, 43, 44 SerpinE2 inhibits its targets by the standard serpin inhibitory mechanism, involving protease binding and cleavage of the reactive center loop, triggering a large conformational change to the serpin molecule trapping the targeted protease in a covalent inactive complex.45, 46 In mammalian cells, serpin-protease complexes bind to members of the low-density lipoprotein receptor family such as low-density lipoprotein receptor-related protein 1 (alpha2-macroglobulin receptor or CD91), triggering a variety of signaling events.47
SerpinE2 has a pro-invasive role in cancer.48 Earlier reports found that SerpinE2 promotes invasion of pancreatic, testicular, mammary, prostate, and lung cancers and gliomas.49, 50, 51, 52, 53, 54, 55 SerpinE2 is upregulated in epithelial cells after oncogenic activation56 and during the progression from pre-neoplastic lesions to medulloblastoma.57 SerpinE2 has not been studied in detail in melanoma, except a recent report associates SerpinE2 to melanoma invasiveness.58 It is not clear how serpins and SerpinE2 contribute to metastasis formation; two recent reports suggest two different mechanisms of action. Serpins could shield metastatic cells from Fas-mediated killing activated by plasminogen-plasmin, providing a pro-metastatic advantage,59 or serpins could act as anticoagulants at the vascular/extravascular interface in the tumor, increasing vascular leakiness favoring extravasation.60 One possible scenario is that LRC, through active secretion of SerpinE2, could direct modulate non-LRC properties, making them more invasive similarly to report in a Zebrafish melanoma model.61 Our data confirm the findings in the fish models that non-LRC express SerpinE2 receptors (Supplementary Figure 5B) and that recombinant SerpinE2 in culture media can directly stimulate non-LRC invasion (Figure 4a; Supplementary Figure 7A). We can also postulate that SerpinE2 contributes to both invasion and survival of LRC. For example, several SerpinE2-associated processes may contribute to LRC extravasation, invasion, and survival in the newly-seeded tissue: cell cycle exit promotion and integrin inactivation achieved in an urokinase Plasminogen activator receptor-dependent manner,62, 63 induction of metalloproteinase release,53, 55 or apoptosis prevention.64
We directly linked SerpinE2 to melanoma invasiveness by first showing that SerpinE2-positive cells have disseminated to mouse lungs, liver and lymph nodes (Figure 3d; Supplementary Figures 6B, C), and then showing that exogenous SerpinE2 stimulates invasion of non-LRC and its neutralization prevents invasion by LRC (Figure 4; Supplementary Figure 7). Moreover, our data confirm that SerpinE2 is exclusively linked to melanoma cells’ invasive capacities. SerpinE2 does not control melanoma cell proliferation, since its abrogation does not change proliferation dynamics or cell viability (Supplementary Figure 9). Furthermore, our data show that there is no direct correlation with MITF expression (Supplementary Figure 6F) or Ki67 (Supplementary Figure 4C). In particular in skin reconstruct, SerpinE2-positive cells were either positive or negative for Ki67. It appears that scattered SerpinE2+ cells are generally negative for Ki67 (Supplementary Figure 4C, top lane) while SerpinE2+ cells found in clusters stain positive for Ki67 (Supplementary Figure 4C, bottom lane). In some areas it is also possible the co-existence of SerpinE2-positive cells that are either positive or negative for Ki67 (Supplementary Figure 4C, middle panel). Unfortunately, there are technical limitations that do not allow live imaging of single-cell migration nor can we detect co-expression of SerpinE2 and PKH26 in vivo.
Our in vitro observations (Figures 4 and 5) were confirmed in patients’ tissues (Figures 5c–e), where SerpinE2 expression correlated with highly malignant lesions. Normal tissue surrounding melanoma lesions is completely negative for SerpinE2. In VGP primary melanoma, two populations of cells co-exist, those that are high- and low-expressing the protein. Vertical growth is the first invasive step of melanomas, in which both invasive and not invasive cells are present within the tumor mass. Increased expression of SerpinE2 could favor the selection of invasive cells to develop a metastatic mass, that is homogenously highly positive for SerpinE2 (Figure 5f). Once disseminated, LRC regain proliferation capacity as demonstrated in vitro and in vivo (Supplementary Figures 3A, 4B and C) and can regenerate the tumor mass. Our data are in line with clinical reports that describe increased SerpinE2 in aggressive stages of gastric,65 colon66 and oral squamous cell carcinomas.67 To complete our study on SerpinE2 role in melanoma, we analyzed SerpinE2 expression in tissue of patients using publicly available data sets. When the TCGA data set was analyzed (469 melanoma samples), we found SerpinE2 mRNA expression upregulated in 6% of patients. These patients show better survival, compared to patients with unvariated expression of SerpinE2. The differences between relative expression levels of SerpinE2 and survival may depend on the type of treatment patients received subsequently to tissue removal. Expression may also depend on size of lesion, location and presence of host cells. In a smaller set of 38 patients with mostly stage 3 melanoma (GSE19234), SerpinE2 expression is associated with poor survival, when compared to patients with relatively low expression of SerpinE2. These descrepancies suggest the need for further correlative studies on SerpinE2 expression with survival, particularly in VGP primary melanomas and biologically early (i.e., regional lymph node) metastases.
In summary, we have shown that melanoma slow-cycling cells (LRC) exist in vitro and in vivo. They disseminate early during tumor growth and are able to efficiently give rise to metastatic lesions at distant sites. We identify a new role for SerpinE2 as responsible for melanoma LRC invasiveness, by what appears to directly involve its anti-proteolytic activity; therefore, a potential new target to prevent or limit melanomas’ early dissemination. We summarized our hypothesis in Supplementary Figure 10. We believe that melanoma slow-cycling cells are primed to invade due to their transcriptional program. Among LRC, SerpinE2high cells are favored in the dissemination process and are selected during organ colonization. Once disseminated, LRC-SerpinE2high cells proliferate and generate non-LRC without losing SerpinE2 expression. Thus, metastases express almost homogenously high SerpinE2. Since SerpinE2 is expressed exclusively by melanoma cells, it is potentially a useful tool to detect isolated malignant cells disseminated in normal organ parenchyma.
Materials and methods
LRC detection
Melanoma cells were stained with CellTrace Violet cell proliferation kit or with PHK26 Red Fluorescent Cell Linker mini-kit (Sigma-Aldrich, St. Louis, MO, USA) according to the manufacturer’s protocol. Fluorescence dilution was measured at indicated time points by flow cytometry (LSRII, Becton Dickinson, Franklin Lakes, NJ, USA) and analyzed using FlowJo Software v10.0 (FlowJo, LLC, Ashland, OR, USA) after normalization on unstained cells. Labeled cells were grown in culture in melanoma media (MCDB154+L-15+2%FBS) for 10–14 days, harvested and stained with DAPI before sorting with MoFlo® Astrios™ (Beckman Coulter, Brea, CA, USA).
In vivo studies
Melanoma cells were stained with PHK26 and 1 × 106 cells/mouse were injected subcutaneously with Matrigel in 1:1 ratio in SHO mice. Tumor growth was measured once to twice a week with manual caliper and mice were imaged by IVIS200 (PerkinElmer) once a week. Dorsal, ventral and lateral scanning was performed. Fluorescent signal was quantified through Living Image 4.3.1 (PerkinElmer), after subtraction of the background. Because IVIS allows in vivo fluorescence detection below 1 cm depth, we also harvested organs at the end of the experiment for additional LRC detection through imaging and flow cytometry.
Proteomic analyses
Proteins at 50 μg/sample were used for proteomic analyses. Proteins were obtained as described.68, 69 Filter-aided sample preparation was performed as described.70, 71 Eluates were vacuum-concentrated and reconstituted in 2 mm ammonia formate buffer and subjected to Tandem Mass Tag™ system (Thermo Fisher Scientific, Waltham, MA, USA) labeling according to manufacturer’s instructions. Two-dimensional liquid chromatography was performed by reversed-phase chromatography at high and low pH as described.72 Mass spectrometry was performed on a hybrid linear trap quadruple Orbitrap Velos mass spectrometer (Thermo Fisher Scientific) using Xcalibur software 2.1.0 (Thermo Fisher Scientific) coupled to an Agilent 1200 HPLC nanoflow system (Proxeon, Odense, Denmark). MS data files were processed with MSConvert (ProteoWizard Library v2.1.2708) and converted into Mascot (MatrixScience, London, UK) files. Peak lists were searched against the human SwissProt database version v2013.01_2013011073 with the search engines Mascot (v2.3.02) and Phenyx (v2.5.14, GeneBio, Geneva, Switzerland).74 Proteins with ⩾2 unique peptides above a score T1, or with a single peptide above a score T2, were selected as unambiguous identifications. Additional peptides for these validated proteins with score >T3 were also accepted. For Mascot and Phenyx, T1, T2 and T3 were equal to 16, 40, 10 and 5.5, 9.5, 3.5, respectively (P-value <10–3). Following the selection criteria, proteins were grouped based on shared peptides, and only the group reporters are considered in the final output of identified proteins. Peptide-spectrum match conflicts between Mascot and Phenyx were discarded. The entire procedure was repeated against a reversed database to assess the protein group false discovery rate (FDR). Peptide and protein group identifications were <0.1 and <1% FDR, respectively. The R software package Isobar75 was used to calculate protein ratios and assess their significance. Protein ratios were calculated based on unique peptides. Proteins were denoted as significantly regulated when their ratio P-value was <0.05, and the ratio itself was >2.5 times the median absolute deviation from the median.
Immunohistochemistry
Fifteen radial growth phase melanomas, 15 metastatic melanomas, and 10 nevi were analyzed. Formalin-fixed, paraffin-embedded tissue sections (1–2 μm) were stained with mouse anti-human SerpinE2 antibody (Origene) followed by polymer staining and Fast-Red for 7 min at RT. Nuclei were counterstained with Hematoxylin for 5 min at RT. Antigen retrieval was achieved at 99 °C for 20 min with ER2 solution (Leica Microsystems, Inc., Buffalo Grove, IL, USA).
Western blotting
For culture supernatant analyses, 2E6 cells were plated in a 6 cm-diameter culture dish for 48 h with 8 ml of culture media. Culture media were harvested, filtered through Millex-GP Syringe-driven Filter Unit (EMD Millipore, part of Merck KGaA, Darmstadt, Germany) and 40 μl of supernatant/sample were used. Samples were boiled 10 min in NuPAGE LDS sample buffer (Thermo Fisher Scientific), loaded on 10% gels, transferred onto nitrocellulose filters and immunoblotted with mouse anti-human SerpinE2 antibody (R&D), diluted 1:500. Anti-mouse secondary antibody was from IRDye (LI-COR Biosciences, Lincoln, NE, USA) and was used at 1:10 000 dilution and membranes were analyzed by Odyssey (LI-COR Biosciences). Densitometry analysis was performed using Odyssey and data were normalized on the total protein detected after Ponceau’s staining.
References
Hodis E, Watson IR, Kryukov GV, Arold ST, Imielinski M, Theurillat JP et al. A landscape of driver mutations in melanoma. Cell 2012; 150: 251–263.
Krauthammer M, Kong Y, Bacchiocchi A, Evans P, Pornputtapong N, Wu C et al. Exome sequencing identifies recurrent mutations in NF1 and RASopathy genes in sun-exposed melanomas. Nat Genet 2015; 47: 996–1002.
Wagle N, Van Allen EM, Treacy DJ, Frederick DT, Cooper ZA, Taylor-Weiner A et al. MAP kinase pathway alterations in BRAF-mutant melanoma patients with acquired resistance to combined RAF/MEK inhibition. Cancer Discov 2014; 4: 61–68.
Van Allen EM, Miao D, Schilling B, Shukla SA, Blank C, Zimmer L et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 2015; 350: 207–211.
Johannessen CM, Johnson LA, Piccioni F, Townes A, Frederick DT, Donahue MK et al. A melanocyte lineage program confers resistance to MAP kinase pathway inhibition. Nature 2013; 504: 138–142.
Van Allen EM, Wagle N, Sucker A, Treacy DJ, Johannessen CM, Goetz EM et al. The genetic landscape of clinical resistance to RAF inhibition in metastatic melanoma. Cancer Discov 2014; 4: 94–109.
Konieczkowski DJ, Johannessen CM, Abudayyeh O, Kim JW, Cooper ZA, Piris A et al. A melanoma cell state distinction influences sensitivity to MAPK pathway inhibitors. Cancer Discov 2014; 4: 816–827.
Cancer Genome Atlas Network. Genomic classification of cutaneous melanoma. Cell 2015; 161: 1681–1696.
Shain AH, Yeh I, Kovalyshyn I, Sriharan A, Talevich E, Gagnon A et al. The genetic evolution of melanoma from precursor lesions. N Engl J Med 2015; 373: 1926–1936.
Roesch A, Fukunaga-Kalabis M, Schmidt EC, Zabierowski SE, Brafford PA, Vultur A et al. A temporarily distinct subpopulation of slow-cycling melanoma cells is required for continuous tumor growth. Cell 2010; 141: 583–594.
Roesch A, Vultur A, Bogeski I, Wang H, Zimmermann KM, Speicher D et al. Overcoming intrinsic multidrug resistance in melanoma by blocking the mitochondrial respiratory chain of slow-cycling JARID1B(high) cells. Cancer Cell 2013; 23: 811–825.
Ostyn P, El Machhour R, Begard S, Kotecki N, Vandomme J, Flamenco P et al. Transient TNF regulates the self-renewing capacity of stem-like label-retaining cells in sphere and skin equivalent models of melanoma. Cell Commun Signal 2014; 12: 52.
Goodall J, Carreira S, Denat L, Kobi D, Davidson I, Nuciforo P et al. Brn-2 represses microphthalmia-associated transcription factor expression and marks a distinct subpopulation of microphthalmia-associated transcription factor-negative melanoma cells. Cancer Res 2008; 1: 68.
Cheli Y, Giuliano S, Botton T, Rocchi S, Hofman V, Hofman P et al. Mitf is the key molecular switch between mouse or human melanoma initiating cells and their differentiated progeny. Oncogene 2001; 19: 30.
Cheli Y, Giuliano S, Fenouille N, Allegra M, Hofman V, Hofman P et al. Hypoxia and MITF control metastatic behaviour in mouse and human melanoma cells. Oncogene 2012; 10: 31.
Bianchi-Smiraglia A, Bagatti A, Fink EE, Moparthy S, Wawrzyniak JA et al. Microphtalmia-associated transcription factor suppress invasion by reducing intracellular GTP pools. Oncogene 2017; 5: 36.
Falletta P, Sanchez-Del-Campo L, Chauhan J, Efffern M, Kenyon A et al. Translational reprogramming is an evolutionary conserved driver of pnhenotypic plasticity and therapeutic resistance in melanoma. Genes Dev 2017; 1: 31.
Fane ME, Chhabra Y, Hollingsworth DE, Simmons JL, Spoerri L et al. NFIB mediates BRN2 driven melanoma cell migration and invasion through regulation of EXH2 and MITF. EBio Med 2017; 16: 63.
Ito M, Liu Y, Yang Z, Nguyen J, Liang F, Morris RJ et al. Stem cells in the hair follicle bulge contribute to wound repair but not to homeostasis of the epidermis. Nat Med 2005; 11: 1351–1354.
Zhou M, Leiberman J, Xu J, Lavker RM . A hierarchy of proliferative cells exists in mouse lens epithelium: implications for lens maintenance. Invest Ophthalmol Vis Sci 2006; 47: 2997–3003.
Wilson BJ, Saab KR, Ma J, Schatton T, Putz P, Zhan Q et al. ABCB5 maintains melanoma-initiating cells through a proinflammatory cytokine signaling circuit. Cancer Res 2014; 74: 4196–4207.
Fang D, Nguyen TK, Leishear K, Finko R, Kulp AN, Hotz S et al. A tumorigenic subpopulation with stem cell properties in melanomas. Cancer Res 2005; 65: 9328–9337.
Boiko AD, Razorenova OV, van de Rijn M, Swetter SM, Johnson DL, Ly DP et al. Human melanoma-initiating cells express neural crest nerve growth factor receptor CD271. Nature 2010; 466: 133–137.
Monzani E, Facchetti F, Galmozzi E, Corsini E, Benetti A, Cavazzin C et al. Melanoma contains CD133 and ABCG2 positive cells with enhanced tumourigenic potential. Eur J Cancer 2007; 43: 935–946.
Raj A, van den Bogaard P, Rifkin SA, van Oudenaarden A, Tyagi S . Imaging individual mRNA molecules using multiple singly labeled probes. Nat Methods 2008; 5: 877–879.
Kousted TM, Skjoedt K, Petersen SV, Koch C, Vitved L, Sochalska M et al. Three monoclonal antibodies against the serpin protease nexin-1 prevent protease translocation. Thromb Haemost 2014; 111: 29–40.
Croteau W, Jenkins MH, Ye S, Mullins DW, Brinckerhoff CE . Differential mechanisms of tumor progression in clones from a single heterogeneous human melanoma. J Cell Physiol 2013; 228: 773–780.
Lee JJ, Cook M, Mihm MC, Xu S, Zhan Q, Wang TJ et al. Loss of the epigenetic mark, 5-Hydroxymethylcytosine, correlates with small cell/nevoid subpopulations and assists in microstaging of human melanoma. Oncotarget 2015; 6: 37995–38004.
Dembinski JL, Krauss S . A distinct slow-cycling cancer stem-like subpopulation of pancreatic adenocarcinoma cells is maintained in vivo. Cancers 2010; 2: 2011–2025.
Hoek KS, Eichhoff OM, Schlegel NC, Dobbeling U, Kobert N, Schaerer L et al. In vivo switching of human melanoma cells between proliferative and invasive states. Cancer Res 2008; 68: 650–656.
Webster MR, Xu M, Kinzler KA, Kaur A, Appleton J, O'Connell MP et al. Wnt5A promotes an adaptive, senescent-like stress response, while continuing to drive invasion in melanoma cells. Pigment Cell Melanoma Res 2015; 28: 184–195.
Pierard-Franchimont C, Hermanns-Le T, Delvenne P, Pierard GE . Dormancy of growth-stunted malignant melanoma: sustainable and smoldering patterns. Oncol Rev 2014; 8: 252.
Rocken M . Early tumor dissemination, but late metastasis: insights into tumor dormancy. J Clin Invest 2010; 120: 1800–1803.
Faries MB, Steen S, Ye X, Sim M, Morton DL . Late recurrence in melanoma: clinical implications of lost dormancy. J Am Coll Surg 2013; 217: 27.
Scott IC, Blitz IL, Pappano WN, Imamura Y, Clark TG, Steiglitz BM et al. Mammalian BMP-1/Tolloid-related metalloproteinases, including novel family member mammalian Tolloid-like 2, have differential enzymatic activities and distributions of expression relevant to patterning and skeletogenesis. Dev Biol 1999; 213: 283–300.
Amano S, Scott IC, Takahara K, Koch M, Champliaud MF et al. Bone morphogenetic protein 1 is an extracellular processing enzyme of the laminin 5 gamma 2 chain. J Biol Chem 2000; 275: 22728–22735.
Garrigue-Antar L, Barker C, Kadler KE . Identification of amino acid residues in bone morphogenetic protein-1 important for procollagen C-proteinase activity. J Biol Chem 2001; 276: 26237–26242.
Wu X, Liu T, Fang O, Leach LJ, Hu X, Luo Z . miR-194 suppresses metastasis of non-small cell lung cancer through regulating expression of BMP1 and p27(kip1). Oncogene 2014; 33: 1506–1514.
Chou CT, Li YJ, Chang CC, Yang CN, Li PS, Jeng YM et al. Prognostic significance of CDCP1 expression in colorectal cancer and effect of its inhibition on invasion and migration. Ann Surg Oncol 2015; 22: 4335–4343.
Leroy C, Shen Q, Strande V, Meyer R, McLaughlin ME, Lezan E et al. CUB-domain-containing protein 1 overexpression in solid cancers promotes cancer cell growth by activating Src family kinases. Oncogene 2015; 34: 5593–5598.
Bouton MC, Boulaftali Y, Richard B, Arocas V, Michel JB, Jandrot-Perrus M . Emerging role of serpinE2/protease nexin-1 in hemostasis and vascular biology. Blood 2012; 119: 2452–2457.
Scott CF, Carrell RW, Glaser CB, Lewis JH, Colman RW . Alpha 1-antitrypsin-Pittsburgh: a potent inhibitor of human plasma factor XIa, kallikrein, and factor XIIf. Trans Assoc Am Phys 1985; 98: 344–351.
Evans DL, McGrogan M, Scott RW, Carrell RW . Protease specificity and heparin binding and activation of recombinant protease nexin I. J Biol Chem 1991; 266: 22307–22312.
Heit C, Jackson BC, McAndrews M, Wright MW, Thompson DC, Silverman GA et al. Update of the human and mouse SERPIN gene superfamily. Hum Genomics 2013; 7: 22.
Silverman GA, Bird PI, Carrell RW, Church FC, Coughlin PB, Gettins PG et al. The serpins are an expanding superfamily of structurally similar but functionally diverse proteins. Evolution, mechanism of inhibition, novel functions, and a revised nomenclature. J Biol Chem 2001; 276: 33293–33296.
Gettins PG . Serpin structure, mechanism, and function. Chem Rev 2002; 102: 4751–4804.
Strickland DK, Ranganathan S . Diverse role of LDL receptor-related protein in the clearance of proteases and in signaling. J Thromb Haemost 2003; 1: 1663–1670.
Kousted TM, Jensen JK, Gao S, Andreasen PA Protease nexin-1 – a serpin with a possible proinvasive role in cancerIn: Behrendt N ed. Matrix Proteases in Health and Disease. Wiley-VCH Verlag GmbH & Co. KGaA: Weinheim, Germany, 2012, pp 251–282.
Buchholz M, Biebl A, Neesse A, Wagner M, Iwamura T, Leder G et al. SERPINE2 (protease nexin I) promotes extracellular matrix production and local invasion of pancreatic tumors in vivo. Cancer Res 2003; 63: 4945–4951.
Neesse A, Wagner M, Ellenrieder V, Bachem M, Gress TM, Buchholz M . Pancreatic stellate cells potentiate proinvasive effects of SERPINE2 expression in pancreatic cancer xenograft tumors. Pancreatology 2007; 7: 380–385.
Nagahara A, Nakayama M, Oka D, Tsuchiya M, Kawashima A, Mukai M et al. SERPINE2 is a possible candidate promotor for lymph node metastasis in testicular cancer. Biochem Biophys Res Commun 2010; 391: 1641–1646.
Fayard B, Bianchi F, Dey J, Moreno E, Djaffer S, Hynes NE et al. The serine protease inhibitor protease nexin-1 controls mammary cancer metastasis through LRP-1-mediated MMP-9 expression. Cancer Res 2009; 69: 5690–5698.
Xu D, McKee CM, Cao Y, Ding Y, Kessler BM, Muschel RJ . Matrix metalloproteinase-9 regulates tumor cell invasion through cleavage of protease nexin-1. Cancer Res 2010; 70: 6988–6998.
Yang S, Dong Q, Yao M, Shi M, Ye J, Zhao L et al. Establishment of an experimental human lung adenocarcinoma cell line SPC-A-1BM with high bone metastases potency by (99 m)Tc-MDP bone scintigraphy. Nucl Med Biol 2009; 36: 313–321.
Pagliara V, Adornetto A, Mammi M, Masullo M, Sarnataro D, Pietropaolo C et al. Protease Nexin-1 affects the migration and invasion of C6 glioma cells through the regulation of urokinase plasminogen activator and matrix metalloproteinase-9/2. Biochem Biophys Acta 2014; 1843: 2631–2644.
Bergeron S, Lemieux E, Durand V, Cagnol S, Carrier JC, Lussier JG et al. The serine protease inhibitor serpinE2 is a novel target of ERK signaling involved in human colorectal tumorigenesis. Mol Cancer 2010; 19: 271.
Vaillant C, Valdivieso P, Nuciforo S, Kool M, Schwarzentruber-Schauerte A, Méreau H et al. Serpine2/PN-1 is required for proliferative expansion of pre-neoplastic lesions and malignant progression to medulloblastoma. PLoS One 2015; 10: e0124870.
Wu QW . Serpine2, a potential novel target for combating melanoma metastasis. Am J Transl Res 2016; 8: 1985–1997.
Valiente M, Obenauf AC, Jin X, Chen Q, Zhang XH, Lee DJ et al. Serpins promote cancer cell survival and vascular co-option in brain metastasis. Cell 2014; 156: 1002–1016.
Wagenblast E, Soto M, Gutierrez-Angel S, Hartl CA, Gable AL, Maceli AR et al. A model of breast cancer heterogeneity reveals vascular mimicry as a driver of metastasis. Nature 2015; 520: 358–362.
Chapman A, Fernandez del Ama L, Ferguson J, Kamarashev J, Wellbrock C, Hurlstone A . Heterogeneous tumor subpopulations cooperate to drive invasion. Cell Rep 2014; 8: 688–695.
Vaillant C, Michos O, Orolicki S, Brellier F, Taieb S, Moreno E et al. Protease nexin 1 and its receptor LRP modulate SHH signaling during cerebellar development. Development 2007; 134: 1745–1754.
Czekay RP, Loskutoff DJ . (2009). Plasminogen activator inhibitors regulate cell adhesion through a uPAR-dependent mechanism. J Cell Physiol 2009; 220: 655–663.
Rossignol P, Ho-Tin-Noe B, Vranckx R, Bouton MC, Meilhac O, Lijnen HR et al. Protease nexin-1 inhibits plasminogen activation-induced apoptosis of adherent cells. J Biol Chem 2004; 279: 10346–10356.
Wang K, Wang B, Xing AY, Xu KS, Li GX, Yu ZH . Prognostic significance of SERPINE2 in gastric cancer and its biological function in SGC7901 cells. J Cancer Res Clin Oncol 2015; 141: 805–812.
Selzer-Plon J, Bornholdt J, Friis S, Bisgaard HC, Lothe IM, Tveit KM et al. Expression of prostasin and its inhibitors during colorectal cancer carcinogenesis. BMC Cancer 2009; 9: 201.
Gao S, Krogdahl A, Sorensen JA, Kousted TM, Dabelsteen E, Andreasen PA . Overexpression of protease nexin-1 mRNA and protein in oral squamous cell carcinomas. Oral Oncol 2008; 44: 309–313.
Maurer M, Müller AC, Wagner C, Huber ML, Rudashevskaya EL, Wagner SN et al. Combining filter-aided sample preparation and pseudoshotgun technology to profile the proteome of a low number of early passage human melanoma cells. J Proteome Res 2013; 12: 1040–1048.
Maurer M, Müller AC, Parapatics K, Pickl WF, Wagner C, Rudashevskaya EL et al. Comprehensive comparative and semiquantitative proteome of a very low number of native and matched epstein-barr-virus-transformed B lymphocytes infiltrating human melanoma. J Proteome Res 2014; 13: 2830–2845.
Manza LL, Stamer SL, Ham AJ, Codreanu SG, Liebler DC . Sample preparation and digestion for proteomic analyses using spin filters. Proteomics 2005; 5: 1742–1745.
Wisniewski JR, Zougman A, Mann M . Combination of FASP and StageTip-based fractionation allows in-depth analysis of the hippocampal membrane proteome. J Proteome Res 2009; 8: 5674–5678.
Olsen R, Sagredo C, Ovrebo S, Lundanes E, Greibrokk T, Molander P . Determination of benzo[a]pyrene tetrols by column-switching capillary liquid chromatography with fluorescence and micro-electrospray ionization mass spectrometric detection. Analyst 2005; 130: 941–947.
Kersey P, Hermjakob H, Apweiler R . VARSPLIC: alternatively-spliced protein sequences derived from SWISS-PROT and TrEMBL. Bioinformatics 2000; 16: 1048–1049.
Colinge J, Masselot A, Carbonell P, Appel RD . InSilicoSpectro: an open-source proteomics library. J Proteome Res 2006; 5: 619–624.
Breitwieser FP, Muller A, Dayon L, Kocher T, Hainard A, Pichler P et al. General statistical modeling of data from protein relative expression isobaric tags. J Proteome Res 2011; 10: 2758–2766.
Acknowledgements
We thank Jeffrey Faust, Scott Weiss and Christopher Corcoran from Wistar’s Flow Cytometry Facility; James Hayden from Wistar’s Imaging Facility and Dmitri Gourevitch from Wistar’s Histopathology Core Facility for technical support; and Rachel E Locke and Jessica L Kohn for editorial assistance. This work was supported by NIH grants P01 CA114046, P01 CA025874, P30 CA010815, R01 CA047159, CA076674, CA182890, by the Melanoma Research Alliance and by the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation, Vienna Science and Technology Fund (WWTF through project LS11-045 to SNW), Vienna Hans Mayr-Fund, and by APART fellowship of the Austrian Academy of Sciences (ÖAW to MM).
Author contributions
MP, RS and MH made intellectual contributions to the conception and/or design of the study. MP, MM, JXW, SS, LL, DH, SS and SL conducted the experiments. SNW, AR, ACM, KP, KLB, JKJ, FK and XX were involved in acquisition, analysis and interpretation of the data. XX was responsible for patients and tumor selection. JKJ was responsible of developing and providing SerpinE2 neutralizing antibody. All authors were involved in drafting and/or critical revision of the manuscript and approved the final submitted version.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Perego, M., Maurer, M., Wang, J. et al. A slow-cycling subpopulation of melanoma cells with highly invasive properties. Oncogene 37, 302–312 (2018). https://doi.org/10.1038/onc.2017.341
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.341
This article is cited by
-
A novel amphiphilic squalene-based compound with open-chain polyethers reduces malignant melanoma metastasis in-vitro and in-vivo
Cell Communication and Signaling (2024)
-
The glutathione S-transferase Gstt1 drives survival and dissemination in metastases
Nature Cell Biology (2024)
-
Targeting tumor exosomal circular RNA cSERPINE2 suppresses breast cancer progression by modulating MALT1-NF-𝜅B-IL-6 axis of tumor-associated macrophages
Journal of Experimental & Clinical Cancer Research (2023)
-
Role of dual specificity phosphatases (DUSPs) in melanoma cellular plasticity and drug resistance
Scientific Reports (2022)
-
Multi-color clonal tracking reveals intra-stage proliferative heterogeneity during mammary tumor progression
Oncogene (2021)