Abstract
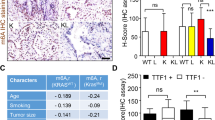
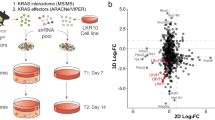
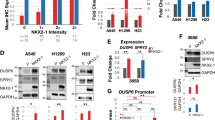
Overexpression of DNMT1 and KIT is prevalent in lung cancer, yet the underlying molecular mechanisms are poorly understood. While the deregulated activation of DNMT1 or KIT has been implicated in lung cancer pathogenesis, whether and how DNMT1 and KIT orchestrate lung tumorigenesis are unclear. Here, using human lung cancer tissue microarrays and fresh frozen tissues, we found that the overexpression of DNMT1 is positively correlated with the upregulation of KIT in tumor tissues. We demonstrated that DNMT1 and KIT form a positive regulatory loop, in which ectopic DNMT1 expression increases, whereas targeted DNMT1 depletion abrogates KIT signaling cascade through Sp1/miR-29b network. Conversely, an increase of KIT levels augments, but a reduction of KIT expression ablates DNMT1 transcription by STAT3 pathway leading to in-parallel modification of the DNA methylation profiles. We provided evidence that KIT inactivation induces global DNA hypomethylation, restores the expression of tumor suppressor p15INK4B through promoter demethylation; in turn, DNMT1 dysfunction impairs KIT kinase signaling. Functionally, KIT and DNMT1 co-expression promotes, whereas dual inactivation of them suppresses, lung cancer cell proliferation and metastatic growth in vitro and in vivo, in a synergistic manner. These findings demonstrate the regulatory and functional interplay between DNA methylation and tyrosine kinase signaling in propelling tumorigenesis, providing a widely applicable approach for targeting lung cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Okano M, Bell DW, Haber DA, Li E . DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 1999; 99: 247–257.
Gowher H, Stockdale CJ, Goyal R, Ferreira H, Owen-Hughes T, Jeltsch A . De novo methylation of nucleosomal DNA by the mammalian Dnmt1 and Dnmt3A DNA methyltransferases. Biochemistry 2005; 44: 9899–9904.
Yu W, Jin C, Lou X, Han X, Li L, He Y et al. Global analysis of DNA methylation by methyl-capture sequencing reveals epigenetic control of cisplatin resistance in ovarian cancer cell. PloS one 2011; 6: e29450.
Arand J, Spieler D, Karius T, Branco MR, Meilinger D, Meissner A et al. In vivo control of CpG and non-CpG DNA methylation by DNA methyltransferases. PLoS genet 2012; 8: e1002750.
Jeltsch A, Jurkowska RZ . New concepts in DNA methylation. Trends biochem sci 2014; 39: 310–318.
Bauer C, Gobel K, Nagaraj N, Colantuoni C, Wang M, Muller U et al. Phosphorylation of TET proteins is regulated via O-GlcNAcylation by the glycosyltransferase OGT. J Biol Chem 2015; 290: 4801–4812.
Wu J, Issa JP, Herman J, Bassett DE Jr, Nelkin BD, Baylin SB . Expression of an exogenous eukaryotic DNA methyltransferase gene induces transformation of NIH 3T3 cells. Proc Natl Acad Sci USA 1993; 90: 8891–8895.
Garzon R, Liu S, Fabbri M, Liu Z, Heaphy CE, Callegari E et al. MicroRNA -29b induces global DNA hypomethylation and tumor suppressor gene re-expression in acute myeloid leukemia by targeting directly DNMT3A and 3B and indirectly DNMT1. Blood 2009; 113: 6411–6418.
Liu S, Liu Z, Xie Z, Pang J, Yu J, Lehmann E et al. Bortezomib induces DNA hypomethylation and silenced gene transcription by interfering with Sp1/NF-kappaB-dependent DNA methyltransferase activity in acute myeloid leukemia. Blood 2008; 111: 2364–2373.
Miller AA, Pang H, Hodgson L, Ramnath N, Otterson GA, Kelley MJ et al. A phase II study of dasatinib in patients with chemosensitive relapsed small cell lung cancer (Cancer and Leukemia Group B 30602). J thorac oncol 2010; 5: 380–384.
Fabbri M, Garzon R, Cimmino A, Liu Z, Zanesi N, Callegari E et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci USA 2007; 104: 15805–15810.
Lin RK, Hsu HS, Chang JW, Chen CY, Chen JT, Wang YC . Alteration of DNA methyltransferases contributes to 5'CpG methylation and poor prognosis in lung cancer. Lung Cancer 2007; 55: 205–213.
Tamborini E, Bonadiman L, Negri T, Greco A, Staurengo S, Bidoli P et al. Detection of overexpressed and phosphorylated wild-type kit receptor in surgical specimens of small cell lung cancer. Clin Cancer Res 2004; 10: 8214–8219.
Litz J, Krystal GW . Imatinib inhibits c-Kit-induced hypoxia-inducible factor-1alpha activity and vascular endothelial growth factor expression in small cell lung cancer cells. Mol cancer therapeut 2006; 5: 1415–1422.
McCabe MT, Davis JN, Day ML . Regulation of DNA methyltransferase 1 by the pRb/E2F1 pathway. Cancer Res 2005; 65: 3624–3632.
Kimura H, Nakamura T, Ogawa T, Tanaka S, Shiota K . Transcription of mouse DNA methyltransferase 1 (Dnmt1) is regulated by both E2F-Rb-HDAC-dependent and -independent pathways. Nucleic Acids Res 2003; 31: 3101–3113.
Jafri NF, Ma PC, Maulik G, Salgia R . Mechanisms of metastasis as related to receptor tyrosine kinases in small-cell lung cancer. J environ pathol toxicol oncol 2003; 22: 147–165.
Levina V, Marrangoni A, Wang T, Parikh S, Su Y, Herberman R et al. Elimination of human lung cancer stem cells through targeting of the stem cell factor-c-kit autocrine signaling loop. Cancer Res 2010; 70: 338–346.
Tsuura Y, Hiraki H, Watanabe K, Igarashi S, Shimamura K, Fukuda T et al. Preferential localization of c-kit product in tissue mast cells, basal cells of skin, epithelial cells of breast, small cell lung carcinoma and seminoma/dysgerminoma in human: immunohistochemical study on formalin-fixed, paraffin-embedded tissues. Virchows Archiv 1994; 424: 135–141.
Boldrini L, Ursino S, Gisfredi S, Faviana P, Donati V, Camacci T et al. Expression and mutational status of c-kit in small-cell lung cancer: prognostic relevance. Clin Cancer Res 2004; 10: 4101–4108.
Jemal A, Siegel R, Ward E, Murray T, Xu J, Thun MJ . Cancer statistics, 2007. CA: cancer j clin 2007; 57: 43–66.
Carlomagno F, Santoro M . Receptor tyrosine kinases as targets for anticancer therapeutics. Curr Med Chem 2005; 12: 1773–1781.
Rossi A, Maione P, Colantuoni G, Ferrara C, Rossi E, Guerriero C et al. Recent developments of targeted therapies in the treatment of non-small cell lung cancer. Curr Drug Discov Technol 2009; 6: 91–102.
Ansari J, Palmer DH, Rea DW, Hussain SA . Role of tyrosine kinase inhibitors in lung cancer. Anticancer Agents Med Chem 2009; 9: 569–575.
Monnerat C, Henriksson R, Le Chevalier T, Novello S, Berthaud P, Faivre S et al. Phase I study of PKC412 (N-benzoyl-staurosporine), a novel oral protein kinase C inhibitor, combined with gemcitabine and cisplatin in patients with non-small-cell lung cancer. Ann Oncol 2004; 15: 316–323.
Schrump DS, Fischette MR, Nguyen DM, Zhao M, Li X, Kunst TF et al. Phase I study of decitabine-mediated gene expression in patients with cancers involving the lungs, esophagus, or pleura. Clin Cancer Res 2006; 12: 5777–5785.
Gauler TC, Christoph DC, Fischer J, Frickhofen N, Huber R, Gonschorek C et al. Phase-I study of sagopilone in combination with cisplatin in chemotherapy-naive patients with metastasised small-cell lung cancer. Eur J Cancer 2013; 49: 2461–2468.
Yuan WC, Lee YR, Huang SF, Lin YM, Chen TY, Chung HC et al. A Cullin3-KLHL20 ubiquitin ligase-dependent pathway targets PML to potentiate HIF-1 signaling and prostate cancer progression. Cancer Cell 2011; 20: 214–228.
Yan F, Shen N, Pang J, Molina JR, Yang P, Liu S . The DNA methyltransferase DNMT1 and tyrosine-protein kinase KIT cooperatively promote resistance to 5-Aza-2'-deoxycytidine (decitabine) and midostaurin (PKC412) in lung cancer cells. J Biol Chem 2015; 290: 18480–18494.
Mott JL, Kurita S, Cazanave SC, Bronk SF, Werneburg NW, Fernandez-Zapico ME . Transcriptional suppression of mir-29b-1/mir-29a promoter by c-Myc, hedgehog, and NF-kappaB. J cell biochem 2010; 110: 1155–1164.
Liu S, Wu LC, Pang J, Santhanam R, Schwind S, Wu YZ et al. Sp1/NFkappaB/HDAC/miR-29b regulatory network in KIT-driven myeloid leukemia. Cancer Cell 2010; 17: 333–347.
Li W, Melton DW . Cisplatin regulates the MAPK kinase pathway to induce increased expression of DNA repair gene ERCC1 and increase melanoma chemoresistance. Oncogene 2012; 31: 2412–2422.
Crivellari G, Monfardini S, Stragliotto S, Marino D, Aversa SM . Increasing chemotherapy in small-cell lung cancer: from dose intensity and density to megadoses. oncologist 2007; 12: 79–89.
Abidin AZ, Garassino MC, Califano R, Harle A, Blackhall F . Targeted therapies in small cell lung cancer: a review. Therapeut adv med oncol 2010; 2: 25–37.
Poirier JT, Gardner EE, Connis N, Moreira AL, de Stanchina E, Hann CL et al. DNA methylation in small cell lung cancer defines distinct disease subtypes and correlates with high expression of EZH2. Oncogene 2015; 34: 5869–5878.
Lin RK, Wu CY, Chang JW, Juan LJ, Hsu HS, Chen CY et al. Dysregulation of p53/Sp1 control leads to DNA methyltransferase-1 overexpression in lung cancer. Cancer Res 2010; 70: 5807–5817.
Itamochi H, Nishimura M, Oumi N, Kato M, Oishi T, Shimada M et al. Checkpoint kinase inhibitor AZD7762 overcomes cisplatin resistance in clear cell carcinoma of the ovary. Int j gynecol cancer 2014; 24: 61–69.
Liu LZ, Zhou XD, Qian G, Shi X, Fang J, Jiang BH . AKT1 amplification regulates cisplatin resistance in human lung cancer cells through the mammalian target of rapamycin/p70S6K1 pathway. Cancer Res 2007; 67: 6325–6332.
Schneider BJ, Kalemkerian GP, Ramnath N, Kraut MJ, Wozniak AJ, Worden FP et al. Phase II trial of imatinib maintenance therapy after irinotecan and cisplatin in patients with c-Kit-positive, extensive-stage small-cell lung cancer. Clin lung cancer 2010; 11: 223–227.
Yan F, Shen N, Pang JX, Zhang YW, Rao EY, Bode AM et al. Fatty acid-binding protein FABP4 mechanistically links obesity with aggressive AML by enhancing aberrant DNA methylation in AML cells. Leukemia 2016; 31: 1434–1442.
Yan F, Shen N, Pang J, Xie D, Deng B, Molina JR et al. Restoration of miR-101 suppresses lung tumorigenesis through inhibition of DNMT3a-dependent DNA methylation. Cell death dis 2014; 5: e1413.
Gao XN, Yan F, Lin J, Gao L, Lu XL, Wei SC et al. AML1/ETO cooperates with HIF1alpha to promote leukemogenesis through DNMT3a transactivation. Leukemia 2015; 29: 1730–1740.
Acknowledgements
This work was supported in part by Hormel Foundation (S. Liu) and National Cancer Institute (Bethesda, MD, USA) grants R01CA149623 (S. Liu) and R21CA155915 (S. Liu).
Author contributions
SL designed the research; FY, NS, JP, NZ and BD performed research; YY, PY and JRM provided the patient samples; FY, BL, PY, JRM and SL analyzed data and wrote the paper; SL conceived the idea and supervised the whole project. All authors discussed the results and commented on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Yan, F., Shen, N., Pang, J. et al. A regulatory circuit composed of DNA methyltransferases and receptor tyrosine kinases controls lung cancer cell aggressiveness. Oncogene 36, 6919–6928 (2017). https://doi.org/10.1038/onc.2017.305
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.305
This article is cited by
-
Coordinated activation of DNMT3a and TET2 in cancer stem cell-like cells initiates and sustains drug resistance in hepatocellular carcinoma
Cancer Cell International (2024)
-
KLRK1 as a prognostic biomarker for lung adenocarcinoma cancer
Scientific Reports (2022)
-
Regulation of DNA methylation machinery by epi-miRNAs in human cancer: emerging new targets in cancer therapy
Cancer Gene Therapy (2021)
-
Methylation of RILP in lung cancer promotes tumor cell proliferation and invasion
Molecular and Cellular Biochemistry (2021)
-
A novel c-Kit/phospho-prohibitin axis enhances ovarian cancer stemness and chemoresistance via Notch3—PBX1 and β-catenin—ABCG2 signaling
Journal of Biomedical Science (2020)