Abstract
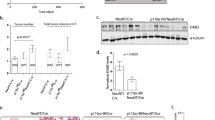
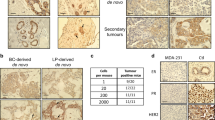
Breast cancer is the most common cancer among women and 30% of patients will be diagnosed with an ErbB2-positive tumor. Forty percent of ErbB2-positive breast tumors have an activating mutation in p110α, a catalytic subunit of phosphoinositide 3-kinase. Clinical and experimental data show that breast tumors treated with a p110α-specific inhibitor often circumvent inhibition and resume growth. To understand this mechanism of resistance, we crossed a p110α conditional (p110αflx/flx) mouse model with mice that overexpress the ErbB2/Neu-IRES-Cre transgene (NIC) specifically in the mammary epithelium. Although mammary-specific deletion of p110α dramatically delays tumor onset, tumors eventually arise and are dependent on p110β. Through biochemical analyses we find that a proportion of p110α-deficient tumors (23%) display downregulation of the Pten tumor suppressor. We further demonstrate that loss of one allele of PTEN is sufficient to shift isoform dependency from p110α to p110β in vivo. These results provide insight into the molecular mechanism by which ErbB2-positive breast cancer escapes p110α inhibition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hernandez-Aya LF, Gonzalez-Angulo AM . Targeting the phosphatidylinositol 3-kinase signaling pathway in breast cancer. Oncologist 2011; 16: 404–414.
Thorpe LM, Yuzugullu H, Zhao JJ . PI3K in cancer: divergent roles of isoforms, modes of activation and therapeutic targeting. Nat Rev Cancer 2015; 15: 7–24.
Engelman JA, Luo J, Cantley LC . The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat Rev Genet 2006; 7: 606–619.
Liu P, Cheng H, Roberts TM, Zhao JJ . Targeting the phosphoinositide 3-kinase pathway in cancer. Cancer Res 2009; 8: 627–644.
Vanhaesebroeck B, Guillermet-Guibert J, Graupera M, Bilanges B . The emerging mechanisms of isoform-specific PI3K signalling. Nat Rev Mol Cell Biol 2010; 11: 329–341.
Czech MP . PIP2 and PIP3: complex roles at the cell surface. Cell 2000; 100: 603–606.
Vanhaesebroeck B, Waterfield MD . Signaling by distinct classes of phosphoinositide 3-kinases. Exp Cell Res 1999; 253: 239–254.
Katso R, Okkenhaug K, Ahmadi K, White S, Timms J, Waterfield MD . Cellular function of phosphoinositide 3-kinases: implications for development, immunity, homeostasis, and cancer. Annu Rev Cell Dev Biol 2001; 17: 615–675.
Stoyanov B, Volinia S, Hanck T, Rubio I . Cloning and characterization of a G protein-activated human phosphoinositide-3 kinase. Science 1995; 269: 690.
Geering B, Cutillas PR, Nock G, Gharbi SI, Vanhaesebroeck B . Class IA phosphoinositide 3-kinases are obligate p85-p110 heterodimers. Proc Natl Acad Sci USA 2007; 104: 7809–7814.
Mellor P, Furber LA, Nyarko JNK, Anderson DH . Multiple roles for the p85α isoform in the regulation and function of PI3K signalling and receptor trafficking. Biochem J 2012; 441: 23–37.
Cantley LC . The phosphoinositide 3-kinase pathway. Science 2002; 296: 1655–1657.
Carpenter CL, Duckworth BC, Auger KR, Cohen B, Schaffhausen BS, Cantley LC . Purification and characterization of phosphoinositide 3-kinase from rat liver. J Biol Chem 1990; 265: 19704–19711.
Hiles ID, Otsu M, Volinia S, Fry MJ, Gout I, Dhand R et al. Phosphatidylinositol 3-kinase: structure and expression of the 110 kd catalytic subunit. Cell 1992; 70: 419–429.
Chantry D, Vojtek A, Kashishian A, Holtzman DA, Wood C, Gray PW et al. p110δ, a novel phosphatidylinositol 3-kinase catalytic subunit that associates with p85 and is expressed predominantly in leukocytes. J Biol Chem 1997; 272: 19236–19241.
Karakas B, Bachman KE, Park BH . Mutation of the PIK3CA oncogene in human cancers. Br J Cancer 2006; 94: 455–459.
Bellacosa A, Etro D, Neri LM . Mutations of the PIK3CA gene in ovarian and breast cancer. Women's Oncol Rev 2005; 5: 223–225.
Bachman KE, Argani P, Samuels Y, Silliman N, Ptak J, Szabo S et al. The PIK3CA gene is mutated with high frequency in human breast cancers. Cancer Biol Ther 2004; 3: 772–775.
Samuels Y, Wang Z, Bardelli A, Silliman N, Ptak J, Szabo S et al. High frequency of mutations of the PIK3CA gene in human cancers. Science 2004; 304: 554–554.
Wee S, Wiederschain D, Maira S-M, Loo A, Miller C, Stegmeier F et al. PTEN-deficient cancers depend on PIK3CB. Proc Natl Acad Sci USA 2008; 105: 13057–13062.
Ni J, Liu Q, Xie S, Carlson C, Von T, Vogel K et al. Functional characterization of an isoform-selective inhibitor of PI3K-p110β as a potential anticancer agent. Cancer Discov 2012; 2: 425–433.
Torbett NE, Luna-Moran A, Knight ZA, Houk A, Moasser M, Weiss W et al. A chemical screen in diverse breast cancer cell lines reveals genetic enhancers and suppressors of sensitivity to PI3K isoform-selective inhibition. Biochem J 2008; 415: 97–110.
Song MS, Salmena L, Pandolfi PP . The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol 2012; 13: 283–296.
Dbouk HA, Backer JM . A beta version of life: p110β takes center stage. Oncotarget 2010; 1: 729.
Wang Q, Liu P, Spangle JM, Von T, Roberts TM, Lin NU et al. PI3K-p110α mediates resistance to HER2-targeted therapy in HER2+, PTEN-deficient breast cancers. Oncogene 2015; 35: 3607–3612.
Schmit F, Utermark T, Zhang S, Wang Q, Von T, Roberts TM et al. PI3K isoform dependence of PTEN-deficient tumors can be altered by the genetic context. Proc Natl Acad Sci USA 2014; 111: 6395–6400.
Fruman DA, Rommel C . PI3K and cancer: lessons, challenges and opportunities. Nat Rev Drug Discov 2014; 13: 140–156.
Andrulis IL, Bull SB, Blackstein ME, Sutherland D, Mak C, Sidlofsky S et al. neu/erbB-2 amplification identifies a poor-prognosis group of women with node-negative breast cancer. Toronto Breast Cancer Study Group. J Clin Oncol 1998; 16: 1340–1349.
Costa C, Ebi H, Martini M, Beausoleil SA, Faber AC, Jakubik CT et al. Measurement of PIP 3 levels reveals an unexpected role for p110β in early adaptive responses to p110α-specific inhibitors in luminal breast cancer. Cancer Cell 2015; 27: 97–108.
Cheng H, Liu P, Ohlson C, Xu E, Symonds L, Isabella A, et al. PIK3CAH1047R-and Her2-initiated mammary tumors escape PI3K dependency by compensatory activation of MEK-ERK signaling. Oncogene 2015; 74: 15–23.
Juric D, Castel P, Griffith M, Griffith OL, Won HH, Ellis H et al. Convergent loss of PTEN leads to clinical resistance to a PI(3)K[agr] inhibitor. Nature 2015; 518: 240–244.
Utermark T, Rao T, Cheng H, Wang Q, Lee SH, Wang ZC et al. The p110α and p110β isoforms of PI3K play divergent roles in mammary gland development and tumorigenesis. Genes Dev 2012; 26: 1573–1586 mistry, 51(18), 5522–5532.
Zhao JJ, Cheng H, Jia S, Wang L, Gjoerup OV, Mikami A et al. The p110α isoform of PI3K is essential for proper growth factor signaling and oncogenic transformation. Proc Natl Acad Sci USA 2006; 103: 16296–16300.
Ursini Siegel J, Hardy WR, Zuo D, Lam SH, Sanguin Gendreau V, Cardiff RD et al. ShcA signalling is essential for tumour progression in mouse models of human breast cancer. EMBO J 2008; 27: 910–920.
Folkes AJ, Ahmadi K, Alderton WK, Alix S, Baker SJ, Box G et al. The identification of 2-(1 H-indazol-4-yl)-6-(4-methanesulfonyl-piperazin-1-ylmethyl)-4-morpholin-4-yl-thieno [3, 2-d] pyrimidine (GDC-0941) as a potent, selective, orally bioavailable inhibitor of class I PI3 kinase for the treatment of cancer. J Med Chem 2008; 51: 5522–5532.
Dourdin N, Schade B, Lesurf R, Hallett M, Munn RJ, Cardiff RD et al. Phosphatase and tensin homologue deleted on chromosome 10 deficiency accelerates tumor induction in a mouse model of ErbB-2 mammary tumorigenesis. Cancer Res 2008; 68: 2122–2131.
Jackson SP, Schoenwaelder SM, Goncalves I, Nesbitt WS, Yap CL, Wright CE et al. PI 3-kinase p110β: a new target for antithrombotic therapy. Nat Med 2005; 11: 507–514.
Beeton CA, Chance EM, Foukas LC, Shepherd PR . Comparison of the kinetic properties of the lipid-and protein-kinase activities of the p110α and p110β catalytic subunits of class-Ia phosphoinositide 3-kinases. Biochem J 2000; 350: 353–359.
Meier TI, Cook JA, Thomas JE, Radding JA, Horn C, Lingaraj T et al. Cloning, expression, purification, and characterization of the human Class Ia phosphoinositide 3-kinase isoforms. Protein Expr Purif 2004; 35: 218–224.
Martin V, Guillermet-Guibert J, Chicanne G, Cabou C, Jandrot-Perrus M, Plantavid M et al. Deletion of the p110β isoform of phosphoinositide 3-kinase in platelets reveals its central role in Akt activation and thrombus formation in vitro and in vivo. Blood 2010; 115: 2008–2013.
Schoenwaelder SM, Ono A, Nesbitt WS, Lim J, Jarman K, Jackson SP . Phosphoinositide 3-kinase p110β regulates integrin αIIbβ3 avidity and the cellular transmission of contractile forces. J Biol Chem 2010; 285: 2886–2896.
Dbouk HA, Vadas O, Shymanets A, Burke JE, Salamon RS, Khalil BD et al. G protein–coupled receptor–mediated activation of p110β by Gβγ is required for cellular transformation and invasiveness. Sci Signal 2012; 5 ra89.
Dbouk HA . PI3King the right partner: unique interactions and signaling by p110β. Postdoc J 2015; 3: 71.
Acknowledgements
We would like to thank Cynthia Lavoie for assistance with the GDC-0941 drug trial and Vasilios Papavasiliou for mammary fat pad injections and transplants; Colin Ratcliffe for technical support with the Metamorph software; Chen Ling for editorial assistance and all the students in the Muller laboratory for their constant feedback and support. Our work is supported by the Canadian Institutes of Health Research (CIHR): MOP-133706 (WJM), FDN-148373 (WJM), Terry Fox team program grant: RI X-242115 (WJM). WJM is supported by CRC Chair in Molecular Oncology. National Institutes of Health (NIH): P50 CA168504 (JJZ), CA187918-02 (JJZ), R35 CA210057 (JJZ), CA172461 (JJZ) and Breast Cancer Research Foundation (JJZ), Roland and Marcel Gosselin Fellowship (AMS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Simond, A., Rao, T., Zuo, D. et al. ErbB2-positive mammary tumors can escape PI3K-p110α loss through downregulation of the Pten tumor suppressor. Oncogene 36, 6059–6066 (2017). https://doi.org/10.1038/onc.2017.264
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2017.264
This article is cited by
-
Circadian gene ARNTL initiates circGUCY1A2 transcription to suppress non-small cell lung cancer progression via miR-200c-3p/PTEN signaling
Journal of Experimental & Clinical Cancer Research (2023)
-
PI3Kβ controls immune evasion in PTEN-deficient breast tumours
Nature (2023)