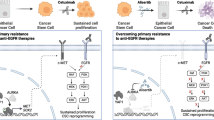
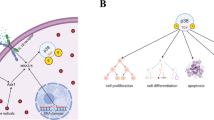
Abstract
Cancer stem cells (CSCs) are considered to be responsible for treatment relapse and have therefore become a major target in cancer research. Salinomycin is the most established CSC inhibitor. However, its primary mechanistic target is still unclear, impeding the discovery of compounds with similar anti-CSC activity. Here, we show that salinomycin very specifically interferes with the activity of K-ras4B, but not H-ras, by disrupting its nanoscale membrane organization. We found that caveolae negatively regulate the sensitivity to this drug. On the basis of this novel mechanistic insight, we defined a K-ras-associated and stem cell-derived gene expression signature that predicts the drug response of cancer cells to salinomycin. Consistent with therapy resistance of CSC, 8% of tumor samples in the TCGA-database displayed our signature and were associated with a significantly higher mortality. Using our K-ras-specific screening platform, we identified several new candidate CSC drugs. Two of these, ophiobolin A and conglobatin A, possessed a similar or higher potency than salinomycin. Finally, we established that the most potent compound, ophiobolin A, exerts its K-ras4B-specific activity through inactivation of calmodulin. Our data suggest that specific interference with the K-ras4B/calmodulin interaction selectively inhibits CSC.
Similar content being viewed by others
Introduction
Cancer stem cells (CSCs) are defined as tumor-initiating cells with a self-renewal capacity similar to that of normal stem cells.1 According to the CSC model, CSCs are at the top of the tumor cell hierarchy. Their defining molecular characteristics as well as whether they emerge from transformed stem/progenitor cells, dormant tumor cells or via epithelial–mesenchymal transition (EMT) are still a matter of intense debate.2, 3 CSCs are thought to be particularly resistant to standard chemotherapeutic agents and are considered to be responsible for relapse after therapy.4, 5 Therefore, CSCs have become the intense focus of more effective therapeutic strategies. Gupta et al.6 have identified salinomycin as being more than 100-fold more potent against human breast CSC than conventional chemotherapeutics. Subsequent studies supported that salinomycin could be a promising CSC inhibitor also for other cancer types, and its potential value in therapy-resistant tumors.7, 8
Different molecular targets have been proposed for the specific CSC activity of salinomycin, suggesting significant polypharmacology.7 Notably, the P-glycoprotein, which is implicated in multi-drug resistance,9 and the Wnt pathway, a canonical stem cell signaling pathway,10 are plausible targets that may ultimately lead to the induction of apoptosis.11 However, these targets have not helped to establish the exact mechanism of action of salinomycin. In particular, it remains unclear which cancer types and patients would show the highest benefit from the salinomycin therapy.
It is largely unknown how the Ras signaling pathway is wired within CSC or embryonal stem cells (ESC) and whether it could be specifically targeted by small molecules. Mutations in the Ras signaling pathway are a hallmark of cancer.12 For Ras itself to function, it must be anchored to the plasma membrane.13 Notably, interference with Ras membrane organization has experimentally been by far more successful than its direct small molecule targeting.14 On the membrane, Ras isoforms H-ras and K-ras4B (hereafter K-ras) are laterally segregated into submicroscopic signaling hubs, termed nanocluster.15 Nanocluster concentrate six to eight Ras proteins and are indispensable for its signaling, as they represent sites for the recruitment of downstream effectors, such as Raf-kinases, that have a high off-rate.16, 17 Nanoclustering of K-ras, but not H-ras, depends on phosphatidylserine (PS).18 Moreover, either the expression of H-rasG12V or the presence of caveolae can critically perturb the PS distribution, thus decreasing K-ras nanoclustering and signaling output.18, 19
Because of the critical role of nanoclustering for Ras signaling, we have previously developed a cell-based assay that exploits the emergence of FRET from fluorescently tagged Ras proteins, when they are tightly packed in nanocluster (nanoclustering-FRET).20 This assay allowed us to identify the modulators of nanoclustering20 as well as the inhibitors of processes upstream of nanoclustering, such as membrane targeting and lipid modifications (for example, statins or farnesylation inhibitors).21, 22, 23 These results illustrate that modulation of the nanoclustering-FRET signal can be due to multiple reasons that may involve direct modulation of the membrane,20 lateral segregation changes of the FRET-probe,24 conformational changes25 or indirect, intracellular processes.22
Here, we utilized this nanoclustering-FRET assay to differentially screen chemical libraries with K- and H-ras-derived FRET biosensors. We identified previously established CSC inhibitors such as salinomycin, and found that salinomycin affects PS nanoscale distribution, thus decreasing K-ras nanoclustering and its effector recruitment. On the basis of this mechanistic insight, we derived a K-ras-associated gene expression signature that is predictive of the salinomycin potency in cancer cell lines and mortality in patients. Importantly, our mechanism-based screening platform enabled us to identify a number of new candidate CSC inhibitors. We demonstrate that the highly potent ophiobolin A interferes with K-ras activity via the inhibition of calmodulin (CaM) as its direct target. Our results suggest that our differential screening approach is highly suitable to identify novel candidate CSC inhibitors.
Results
Differential screening with K- and H-ras nanocluster FRET biosensors identifies CSC inhibitors
The nanoscale packing of Ras proteins on the plasma membrane, termed nanoclustering, is necessary for Ras signaling.17, 26 In specific cancer cases, nanoclustering can be exploited to hyperactivate Ras,27 therefore representing a potential target to suppress hyperactive Ras in cancer. We recently identified ionophoric macrotetrolides/nactins as disruptors of H-ras associated nanocluster.20 Owing to their ability to insert into the plasma membrane, ionophores may be good candidates for nanocluster disruptors. We therefore tested here a small collection of 13 ionophoric compounds using our previously established NANOcluster and Prenylation Sensors (NANOPS) FRET assay in mammalian cells. In this assay, we exploit the FRET, which arises because of nanoclustering of membrane-anchored fluorescent proteins (nanoclustering-FRET) (Figure 1a).
Differential screening identifies CSC inhibitors. (a) Schematic representation of the FRET assay, which reports on nanoclustering and subcellular distribution changes of Ras-derived biosensors (NANOPS). NANOPS exploit the high FRET associated with nanoclustering of fluorescently tagged membrane anchors of H-ras or K-ras4B. (b) Scheme of chemical screen workflow. (c) Scatter plot of the chemical screen with BHK cells expressing K- and H-ras-NANOPS. Emax values of cells treated for 24 h with 1 μg/ml (black dots) and 2 μg/ml (green dots) of compounds (see Supplementary Table 1) are shown. Bars denote the s.e.m. from three independent experiments. Block line indicates the average Emax (K-ras-NANOPS, 0.32; H-ras-NANOPS, 0.40), dotted lines mark the hit selection threshold of 15% change from the average Emax. Arrows indicate the hits. Identity, compound number and chemical structures of the hits are represented below.
H- and K-ras were shown to segregate into distinct nanodomains, from where distinct signaling output emanates.28 To identify Ras-isoform-specific inhibitors and to filter out more trivial activities, such as inhibition of prenylation by statins, we performed differential screening with two Ras-derived biosensors H-ras-NANOPS (H-ras derived20) and K-ras-NANOPS (K-ras derived29).
BHK cells transiently expressing either of these biosensors were incubated with compounds and then analyzed using our well-established flow cytometry FRET assay (Figure 1b).20, 21, 23, 29, 30 Three compounds, salinomycin, nigericin and lasalocid, changed the nanoclustering-associated FRET-parameter Emax by at least 15% specifically for K-ras-NANOPS and were thus selected as hits for further evaluation (Figure 1c). Interestingly, the compound activity did not seem to correlate with the ion selectivity, as compounds with similar ion selectivity were not identified as hits (Supplementary Table 1). This suggests a mechanism of action that does not relate to the ionophoric activity.
CSC inhibitors perturb the nanoscale membrane organization of PS and Ras
The three hit compounds are known and in use for their antibiotic/anticoccidial activity.31 More interestingly in the cancer context is that salinomycin and nigericin have been identified as CSC inhibitors.6 CSCs are necessary for tumor initiation and are linked to relapse after therapy, because of their exquisite resistance against standard antiproliferative cancer drugs. Therefore, CSC inhibitors bear an exceptional significance as future anti-cancer drugs. However, their exact molecular target remains controversial. Likewise, it is not resolved whether they truly target a ‘stemness’ feature and which patients would benefit from treatment.
To explore a K-ras associated mechanism of action of our hit compounds, we validated their specific activity on full length K- or H-rasG12V. For both proteins, FRET pairs were generated and co-expressed in BHK cells to measure isoform-specific nanoclustering-FRET using fluorescence lifetime imaging microscopy (FLIM). Salinomycin and lasalocid specifically reduced nanoclustering-FRET of K-rasG12V (Figures 2a and c), while nigericin also reduced FRET of H-rasG12V (Figures 2b and c).
CSC inhibitors affect K-rasG12V nanoclustering in a PS-dependent manner. (a and b) Nanoclustering-FRET analysis (illustrated in schemes) in BHK cells co-expressing mGFP- and mCherry-tagged (a) K-rasG12V or (b) H-rasG12V. Cells were treated for 24 h with either DMSO control, 1.3 μM salinomycin, 1.3 μM nigericin or 1.7 μM lasalocid. The numbers in the bars indicate the number of analyzed cells. (c) Examples of FLIM-FRET images of BHK cells from nanoclustering-FRET experiments as indicated. Image color look-up table on the right shows fluorescence lifetimes. (d) Confocal images of MDCK cells stably expressing mGFP-K-rasG12V or mGFP-H-rasG12V treated with the inhibitors for 24 h. Representative images of treatments from three independent experiments are shown. Scale bar, 20 μm. (e and f) Co-clustering FRET analysis (illustrated in schemes) using FLIM on BHK cells expressing mGFP-LactC2 and (e) mCherry-K-rasG12V or (f) mCherry-H-rasG12V and treated like in (a and b). In all graphs (a, b, e and f), the apparent FRET efficiency was calculated from FLIM data (mean±s.e.m., n=3). The numbers in the bars indicate the number of analyzed cells. Statistical significance of differences between control and treated cells were examined using one-way ANOVA tests (n.s., not significant; **P<0.01; ***P<0.001; ****P<0.0001).
In our nanoclustering-FRET assays, the loss of FRET can be due to changes in Ras nanoclustering, subcellular redistribution or loss of membrane anchorage (Figure 1a). Confocal imaging of MDCK cells stably expressing mGFP-tagged K- or H-rasG12V revealed that all compounds led to a strong loss of plasma membrane labeling and perinuclear redistribution of both Ras isoforms (Figure 2d). Similar observations were made in transiently transfected BHK cells (Supplementary Figure 1a).
Recently, low nanomolar concentrations of staurosporine (STS) were described to affect nanoclustering and subcellular distribution of K-ras32 (Supplementary Figures 1a and b). This activity was reminiscent of that of our hit compounds. Mechanistically, STS redistributes cellular PS from the plasma membrane to endomembranes.32 The negatively charged PS is critical for plasma membrane anchorage and clustering of K-ras, which possesses a polybasic (positively charged) C-terminal membrane anchor.18 We therefore assayed for a similar effect of our hit compounds on PS nanoscale organization and coclustering with Ras. To this end, we used the fluorescently tagged C2 domain of lactadherin (LactC2), which binds to PS.33 All compounds significantly decreased nanoclustering-FRET of PS in BHK cells (Supplementary Figure 1b). Interestingly, only salinomycin had a specific effect on coclustering of PS and K-rasG12V (Figure 2e), but not on PS/H-rasG12V (Figure 2f), while the other compounds, including the tool compound STS, decreased PS coclustering FRET with both K- and H-rasG12V. This K-ras-specific activity of salinomycin may be significant, as the overexpression of H-rasG12V negatively regulates K-ras nanoclustering via segregation of PS.18 Thus, H-rasG12V nanoclustering antagonizes an increase in K-ras nanoclustering and signaling.
We conclude that our hit compounds predominantly disrupt K-ras nanoscale membrane organization by decreasing nanoscale PS clustering, while they also affect compound-specifically PS/Ras coclustering.
Salinomycin most specifically reduces K-ras activity
We next followed up on the downstream consequences of K-ras nanoscale disruption by the hit compounds. An essential step in Ras/MAPK signal propagation is the recruitment of the effector Raf to the plasma membrane by active, nanoclustered Ras.16 We have previously established a FRET assay, where we co-expressed mGFP-tagged RasG12V with the mRFP-tagged Ras-binding domain (RBD) of the effector C-Raf in BHK cells, to quantify the effector recruitment step.17, 29, 34
All three hit compounds significantly and specifically reduced FRET between K-rasG12V and the RBD (Figure 3a), but not between H-rasG12V and the RBD (Figure 3b). This was associated with a specific reduction in downstream MAPK signaling output of K-rasG12V (Figure 3c)- but not H-rasG12V (Figure 3d)-transfected BHK cells by the hit compounds. In line with a significant role of MAPK signaling in cell growth, proliferation of non-transfected BHK cells was efficiently suppressed by salinomycin and nigericin (Figure 3e).
Salinomycin most specifically attenuates K-ras-effector recruitment, -MAPK signaling and cell proliferation. (a and b) Effector-recruitment FRET assay (illustrated in schemes) in BHK cells expressing (a) mGFP-K-rasG12V or (b) mGFP-H-rasG12V and mRFP-RBD. Cells were treated for 24 h with either DMSO control, 1.3 μM salinomycin, 1.3 μM nigericin or 1.7 μM lasalocid. The apparent FRET efficiency was calculated from FLIM data (mean±s.e.m., n=3). The numbers in the bars indicate the number of analyzed cells. Statistical significance of differences between control and treated cells was examined using one-way ANOVA (****P<0.0001). (c) Left, representative western blots from mGFP-K-rasG12V transfected BHK cells serum-starved for 5 h and subsequently treated for 24 h with inhibitors. Right, quantification of western blots showing relative ppERK levels from six independent experiments. (d) Left, representative western blots from mGFP-H-rasG12V-transfected BHK cells serum-starved for 5 h and subsequently treated for 24 h with inhibitors. Right, quantification of western blots showing relative ppERK levels from three independent experiments. Statistical significance was examined using one-way ANOVA (**P<0.01). (e) Proliferation of BHK cells treated for 72 h with the inhibitors. The graph shows mean fold change compared with DMSO control at each time point±s.e.m. (n=6).
These data suggest that salinomycin, the most potent CSC inhibitor with potential efficacy in human cancers,8 specifically disrupts K-ras nanoscale membrane organization. This effectively reduces effector recruitment to K-ras, which then compromises at least MAPK signaling and proliferation.
High caveola levels decrease the sensitivity to salinomycin
When validating the activity of our hits in HEK cells using Ras-NANOPS, the activity of the hit compounds was lost (Supplementary Figures 2a and b). To explain this, we focused our attention on the different abundance of caveolae in these cell lines as it was recently shown that downregulation of caveolae perturbs the lipid composition of the plasma membrane. This leads to an increase in PS clustering and as a consequence of K-ras nanoclustering and signaling output, while H-ras nanoclustering remains unaffected19 (Supplementary Figure 2c). Caveolin-1 (Cav-1) and PTRF/cavin-1 are the two major protein components of cellular caveolae.35, 36 We therefore tested whether reexpression of Cav-1 and PTRF in HEK cells could restore the K-ras-directed activity of the hit compounds that was observed in BHK cells.
Caveolae are 50–80 nm in size, which required us to quantify caveolae levels in HEK and BHK cells using stimulated emission depletion (STED)-superresolution microscopy37 (Figure 4a). HEK cells had a significantly lower number of caveolae than BHK cells (Figure 4b). As expected, transient reexpression of Cav-1 together with PTRF increased the number of caveolae in HEK cells to the same level as that found in BHK cells. In agreement with the data by Ariotti et al.,19 this treatment significantly reduced K-rasG12V nanoclustering-FRET (Figure 4c), while H-rasG12V nanoclustering remained unchanged (Figure 4d). Owing to its lower potency (Figure 3e), we excluded lasalocid from subsequent experiments. Reexpression of the caveolar proteins did not significantly affect the H-ras nanoclustering-FRET response to either salinomycin or nigericin. By contrast, the K-ras nanoclustering-FRET was highly significantly reduced by salinomycin, while that of nigericin remained at the same low level as in control HEK cells. This effect of caveolar protein reexpression to specifically sensitize K-ras but not H-ras activity to salinomycin was also seen on the level of effector recruitment (Figures 4e and f). In agreement with these FRET data, Cav-1 and PTRF expression specifically desensitized HEK cells (Figure 4g), but not BHK cells (Figure 4h) to salinomycin. This effect is consistent with our observations in Figure 3e, as also H-rasG12V negatively regulates PS and therefore K-ras-nanoclustering18 similar to caveolae.
Caveolar protein reexpression desensitizes HEK cells to salinomycin. (a) Imaging of caveolae in HEK, HEK overexpressing Cav-1+PTRF and BHK cells with confocal- and STED-microscopy. Scale bars, confocal and STED 2.5 μm. (b) Quantification of caveolae-like structures from HEK, HEK-overexpressing Cav-1+PTRF and BHK cells. Shown are the mean number (± s.e.m.) of caveolae-like structures per μm2 of plasma membrane from at least 10 cells. (c and d) Nanoclustering-FRET analysis in HEK and Cav-1+PTRF-overexpressing HEK cells co-expressing mGFP- and mCherry-tagged (c) K-rasG12V or (d) H-rasG12V. Cells were treated for 24 h with DMSO control, 1.3 μM -salinomycin or 1.3 μM nigericin. (e and f) Effector-recruitment FRET assay in HEK and Cav-1+PTRF-overexpressing HEK cells expressing (e) mGFP-K-rasG12V or (f) mGFP-H-rasG12V and mRFP-RBD and treated like in (c and d). In graphs (c–f), the apparent FRET efficiency was calculated from FLIM data (mean±s.e.m., n=3). The numbers in the bars indicate the number of analyzed cells. (b–f) Statistical significance of differences between control and treated cells were examined using one-way ANOVA tests (***P<0.001; ****P<0.0001). (g and h) Dose–response analysis of either (g) HEK or (h) BHK cells with or without Cav-1+PTRF overexpression for 24 h with salinomycin. Data represent the relative cell viability±s.e.m. from three independent experiments performed in triplicate.
Thus, salinomycin but not nigericin clearly discriminates between normal HEK cells and those with caveolar protein overexpression.
A K-ras-nanocluster gene expression signature is predictive for sensitivity to salinomycin and patient survival
It is important to distinguish between the ease to observe the K-ras isoform specificity of salinomycin by FRET (in caveolae-rich BHK cells, with little free PS) and the dependence of the cell proliferation on a high K-ras signaling level (in caveolae-poor HEK cells, with relatively higher PS and higher K-ras nanoclustering). The latter renders HEK cells ∼8 times more sensitive to the PS/K-ras coclustering disrupting salinomycin (IC50 (HEK)=0.63 μM) as compared with BHK cells (IC50 (BHK)=4.9 μM).
In an effort to apply this insight to cancer cells, we wanted to establish a relationship between the absolute potency of salinomycin and factors such as caveolae that are known to impact on the nanoscale membrane organization (nanoclustering) of K-ras (Supplementary Table 2).
We therefore first examined the expression patterns of genes that are associated with Ras and their nanoclustering in ESC, assuming that CSC inhibitors might target molecular features of a normal ESC. We compared the expression of all three Ras isoforms, the EGFR38 and the six genes of known K-ras nanocluster modulators (Supplementary Table 2) in fibroblasts and human ESCs, using gene expression profiles from the stem cell database ESTOOLS.39 We found that all fibroblasts display a relatively similar expression signature, which is basically inverted in a large set of ESCs. Altogether, we recognized three distinct sets of ESC expression signatures characteristic for naive ESC, primed ESC40 and those that appeared to be in transit between these two states (Supplementary Figure 3a).
We next extended these expression signatures by searching the ESTOOLS database for additional genes that showed a co-expression pattern with the initial set of 10 genes (see Online Methods for details). From this list of 32 co-expressed genes (Supplementary Table 3), we selected vimentin (VIM), caveolin-2 (Cav-2) and integrin αV (ITGA5) as the most plausible genes to be associated with K-ras nanocluster and stemness regulation.
Strikingly, unsupervised clustering of this extended panel of genes separated them consistent with K-ras nanoclustering being high in ES cells and low in differentiated cells/fibroblasts (Figure 5a; Supplementary Figure 3a). These results prompted us to test whether salinomycin targets K-ras in cancer cell lines in a gene expression context similar to that of primed human ESCs. Conversely, such cell lines should be more resistant to conventional chemotherapeutics.6
A K-ras-nanoclustering signature predicts cancer cell line sensitivity to salinomycin and patient survival. (a) List of definitive K-ras-nanoclustering-associated genes and their relative expression in fibroblasts and ESCs. Bottom, expected K-ras signaling outcome. (b) Expression profiles of cancer cell lines with either ESC-like or fibroblast-like K-ras-nanoclustering signature. (c and d) Drug sensitivity analysis of cancer cell lines treated with salinomycin (c) or STS (d) for 72 h. The bars show the mean IC50 values from triplicate measurements. (e) Spearman correlation coefficient between the drug response and gene expression profiles. Statistical significance was calculated on the basis of exact null distribution using Ryser’s formula (*P<0.05). (f) Overall survival analysis between patient tumor samples from TCGA with gene expression signatures similar to ESCs (n=605) vs fibroblasts (n=402).
By correlating the expression profiles of our set of 13 predominantly K-ras-nanoclustering genes (hereafter K-ras-nanoclustering signature) from cancer cell lines in the cancer cell line encyclopedia (CCLE) database,41 ESCs and fibroblasts, we identified specific cancer cell lines whose K-ras-nanoclustering signature was either more ESC-like or more fibroblast-like (Figure 5b, Supplementary Figure 4).
Sensitivity profiles of salinomycin and a selected conventional chemotherapeutic drug (STS) across 15 cancer cell lines revealed that those cell lines predicted to have a more ESC-like signature typically showed a higher sensitivity to salinomycin than the cell lines with a fibroblast-like signature (Figure 5c; P=0.051, Wilcoxon rank sum test). Consistent with our prediction, the opposite was found for their sensitivity to STS (Figure 5d); P=0.067. This was further supported by correlating the drug response profiles with the gene expression signature, which confirmed that the salinomycin response showed a positive trend of association with KRAS expression, but negative in particular with the expression of caveolar genes (Cav-1, Cav-2, PTRF) (Figure 5e). Strikingly, the STS response showed predominantly the opposite correlation pattern with the gene expression signature (Figure 5e; P=0.0024, Wilcoxon signed rank test). These data suggested that our K-ras-nanoclustering signature could predict the sensitivity of cancer cell lines to the CSC inhibitor salinomycin.
Interestingly, a substantial fraction ~8% (605/7536) of the patient tumor samples in The Cancer Genome Atlas (TCGA) database exhibited an ESC-like and therefore CSC inhibitor sensitive K-ras-nanoclustering signature (at Spearman rank correlation coefficient cutoff of 0.6; see Methods for details), which was also associated with a significant negative effect on the overall survival (Figure 5f, Cox proportional hazard ratio (95% confidence interval)=1.36 (1.06–1.73), P=0.013). Interestingly, this effect was even more significant, when the ESC-like cancer cell line expression data were used (Supplementary Figure 3b, Cox proportional hazard ratio (95% confidence interval)=1.27 (1.10–1.46), P=8.8 × 10−4). This is consistent with our assumption that tumors with the K-ras-nanoclustering signature contain a high number of CSC or CSC-like cells, which are refractory to conventional therapy and/or mediating more aggressive cancers. Intriguingly, among these tumor samples with an ESC-like signature, we found enrichment for cancer types of the female reproductive tissue with the highest proportion in breast cancer, as well as a relatively high proportion and high enrichment for acute myeloid leukemia (Supplementary Table 4).
In summary, we have identified a panel of only 13 genes (the K-ras-nanoclustering signature) that can predict the sensitivity of cancer cell lines to the CSC inhibitor salinomycin. Importantly, this gene expression signature was also associated with poorer survival outcome in patients with cancers in particular of the female reproductive tissue.
Additional compound screening identifies ophiobolins and conglobatin A as new candidate CSC inhibitors
To demonstrate that our K-ras-focused screening platform allows for the identification of novel compounds with anti-CSC activity, we conducted a second screen with a chemically more diverse library of ~400 microbial metabolites (Supplementary Figure 5a).
In concordance with our previous screening results,20 nactins/macrotetrolides were again identified as H-ras-NANOPS specific hits (Supplementary Table 5). Moreover, among the hit compounds that positively affected the K-ras-NANOPS FRET were a number of STS derivatives, consistent with our observations with STS as a tool compound that affects PS nanoclustering and PS/Ras coclustering (Figure 2, Supplementary Figure 1b). The positive deviation of the NANOPS FRET is specific to the cytometer FRET method and may relate to the distribution of the biosensor to internal membranes. Importantly, we found avermectin, which was also identified in the CSC inhibitor screen by Gupta et al. as abamectin (a mixture of avermectin B1a and B1b), and its derivative ivermectin. Thus, out of the four compounds that were more closely studied by Gupta et al., we have identified three.
Despite the relatively small size of our compound library, five substance classes (leptomycins, avermectins, ophiobolins, conglobatin and streptonigrin) with some selectivity for the K-ras-NANOPS were found (Supplementary Table 5). We included at least one representative from each substance class for validation, thus ending up with a selection of seven new compounds that were assessed for their potential as CSC inhibitors (Supplementary Table 6). First, we validated their Ras-isoform selectivity. All compounds significantly decreased K-rasG12V nanoclustering-FRET (Figure 6a). In contrast, H-rasG12V nanoclustering-FRET remained largely unaffected, except under leptomycin B and streptonigrin treatment (Figure 6b). This activity profile was basically also reflected by the effect of the compounds on effector recruitment-FRET, with almost all compounds significantly reducing FRET of K-rasG12V/RBD, but not of H-rasG12V/RBD (Supplementary Figure 5b). Unlike with the ionophores that were found in the first screen (Figure 2d), subcellular distribution of K-rasG12V or H-rasG12V remained apparently unaffected (Supplementary Figure 5c).
K-ras-specific screening identifies new candidate CSC inhibitors. (a and b) Nanoclustering-FRET analysis in BHK cells co-expressing mGFP- and mCherry-tagged (a) K-rasG12V or (b) H-rasG12V. The apparent FRET efficiency was calculated from FLIM data (mean±s.e.m., n=3). Cells were treated for 24 h with either DMSO control, 1.3 μM salinomycin or 0.2 μM of avermectin, ivermectin, conglobatin A, ophiobolin A, kazusamycin B, leptomycin B or streptonigrin. Inhibitors are color-coded by chemical class as in Supplementary Table 4. Red dotted line denotes the average response with salinomycin as positive control. Numbers in the bars indicate the number of analyzed cells. Statistical significance of differences between control and treated cells was examined using one-way ANOVA (**P<0.01; ***P<0.001; ****P<0.0001). (c) Left, CD44/CD24 FACS profiles are shown for MDA-MB-231 after compound treatments. The green ellipse denotes the CD44+/CD24− breast CSC fraction and the dotted-black ellipse marks the CD44+/CD24+ fraction. Right, shown is the average percentage of CD44+/CD24− MDA-MB-231 cells after treatment for 4 days with either DMSO control, 100 ng/μl doxorubicin, 10 nM STS or 2.0 μM of salinomycin, avermectin, ivermectin, conglobatin A, ophiobolin A, leptomycin B, kazusamycin B and streptonigrin. Error bars denote the s.e.m. from three independent experiments performed in duplicate. Statistical significance of differences between doxorubicin treated and cells treated with other compounds were examined using one-way ANOVA complemented with Tukey’s test (***P<0.001). (d) Left, mammosphere formation efficiency of MDA-MB-231, MDA-MB-436, Hs578T and MCF7 cells grown in non-adherent conditions. Mammospheres were allowed to form for 6 days and treated for 3 additional days with either DMSO control or the indicated compounds (mean±s.e.m., n=4). Right, representative images of mammospheres as indicated. Bar represents 1000 μm. Statistical significance of differences between doxorubicin-treated and cells treated with other compounds were examined using one-way ANOVA complemented with Tukey’s test (n.s., not significant; **P<0.01; ***P<0.001; ****P<0.0001).
To evaluate the potential of these compounds as CSC inhibitors, we next studied their effect on the CD44+/CD24− (invasive, mesenchymal) CSC population that is naturally found in the MDA-MB-231 breast cancer cell line, using a fluorescence-activated cell sorting (FACS)-based assay.42 Both conglobatin A and ophiobolin A decreased the CD44+/CD24− population to a comparable or higher extent than salinomycin (Figure 6c).
This CSC-specific activity was confirmed by growing mammary cancer cells under non-adherent conditions as mammospheres, a well-established model for CSC.43 MDA-MB-231, MDA-MB-436, Hs578T and MCF7 grown under non-adherent conditions formed mammospheres with comparable efficiencies, albeit they contained different fractions of CD44+/CD24− cells as established by others (MDA-MB-231≈Hs578T>MDA-MB-436, none in MCF7) (Supplementary Table 7).42
In agreement with the different stemness characteristics of these cell lines, we found that they expressed stemness marker Oct3/4 at high levels already when grown under adherent conditions (MDA-MB-231, Hs578T) or only when grown as mammospheres (MCF7). Stemness marker Sox2 was detected only at low levels under both conditions, while also NANOG expression somewhat increased for MDA-MB-436 and MCF7 cells, while expression remained low for the two other cell lines (Supplementary Figure 6a).
Specific stemness characteristics were also reflected in the response to CSC drug treatment. Only mammospheres that were derived from cell lines with a high CD44+/CD24− population (MDA-MB-231, MDA-MB-436, Hs578T) responded selectively to salinomycin, conglobatin A and ophiobolin A as compared with doxorubicin (Figure 6d). Interestingly, ophiobolin A was most potent in both the K-rasG13D-mutated MDA-MB-231 and in the H-rasG12D-mutated Hs578T mammospheres. Both of these cell lines belong to the highly aggressive basal-like, claudin-low, triple negative type (Supplementary Table 7). Note that the apparent potency of ophiobolin A in MDA-MB-231 cells grown under adherent conditions was only slightly lower as compared with that in mammospheres (Supplementary Figure 6b), which is consistent with the high fraction of CD44+/CD24− cells in that cell line.
Collectively, these results demonstrate that ophiobolin A is a highly potent inhibitor of mammospheres derived from the most aggressive breast cancer cell lines.
Identification of CaM as the K-ras-directed target of ophiobolin A
Ophiobolin A is a potent CaM inhibitor, which modifies lysines 75 or 77 and 148 by covalent Schiff base bond formation.44 The McCormick group45 very recently proposed that blocking of the K-ras/CaM interaction might represent a novel approach to specifically interfere with K-ras signaling in cancer. We therefore tested, whether the inhibition of CaM can suppress mammosphere formation. Indeed, CaM inhibitors W7 and with high efficiency also calmidazolium reduced the growth of MDA-MB-231-derived mammospheres (Figure 7a). This effect was phenocopied by siRNA-mediated depletion of CaM (Figure 7a; Supplementary Figure 6c).
Inhibition of CaM reduces mammosphere formation and K-ras nanoclustering-FRET. (a) Left, mammosphere formation efficiency of MDA-MB-231 cells or MDA-MB-231 cells pre-transfected with CaM siRNA (siCaM1), grown in non-adherent conditions was determined. Mammospheres without knockdown were allowed to form for 6 days and then treated for three additional days with either DMSO control or 2 μM of the indicated compounds before being counted (mean±s.e.m., n=3). Right, representative images of mammospheres as indicated. Bar represents 1000 μm. Statistical significance of differences between doxorubicin-treated or control (as indicated) and cells treated with other compounds were examined using one-way ANOVA complemented with Tukey’s test (n.s., non significant; ****P<0.0001). (b) Nanoclustering-FRET analysis in HEK cells co-expressing mGFP- and mCherry-tagged K-rasG12V. After cells were transfected with scrambled or CaM siRNA and plasmids for overexpressing wild-type or mutant CaM to rescue the knockdown, they were treated with either DMSO control or 0.2 μM ophiobolin A for 24 h. The apparent FRET efficiency was calculated from FLIM data (mean±s.e.m., n=5). Statistical significance of differences between control and treated cells was examined using one-way ANOVA. (c) Mammosphere formation efficiency of MDA-MB-231 cells grown in non-adherent conditions. Mammospheres were allowed to form for 6 days and treated for three additional days with either DMSO control or the indicated concentrations of ophiobolin A and conglobatin A before being counted (mean±s.e.m., n=3). Statistical significance of differences between doxorubicin-treated and cells treated with other compounds were examined using one-way ANOVA complemented with Tukey’s test (****P<0.0001). (d) Left, MDA-MB-231 cells were treated for 3 days with ophiobolin analogs and conglobatin A, and then their CD44/CD24 population was assessed by flow cytometry. Shown is the percentage of CD44+/CD24− breast CSC subpopulation after treatment with inhibitors at indicated concentrations (mean±s.e.m., n=3). Statistical significance of differences between control and treated cells were examined using one-way ANOVA complemented with Tukey’s test (****P<0.0001). Right, chemical structures of conglobatin A and ophiobolin analogs A, B and C are presented. The red, dashed square marks the oxygen 13 that correlated with activity of ophiobolins A and B, but is absent in ophiobolin C.
To provide proof for CaM as the decisive target of ophiobolin A, we evaluated the effect of ophiobolin A on K-ras nanoclustering-FRET in HEK cells expressing wild-type or ophiobolin resistant CaM. First, siRNA-mediated down-modulation of CaM led to a loss of FRET identical to that of cells treated with ophiobolin A (Figure 7b). This loss of FRET could be fully rescued by both wild-type CaM and CaM carrying mutations K75Q, K77Q, K148Q (mut-CaM), which render it ophiobolin A-resistant.44 Correspondingly, only rescued cells expressing the wild-type CaM responded to ophiobolin A, while those rescued with mut-CaM were not showing any decrease of FRET (Figure 7b). Importantly, the same experiment carried out with conglobatin A showed that mut-CaM rescued cells still responded with a loss of FRET to it, demonstrating the ophiobolin A specificity and indicating that conglobatin A has a different mechanism of action and probably an entirely different target than ophiobolin A (Supplementary Figure 6d). In summary, these results suggest that the anti-CSC potential of ophiobolin A is due to its inhibitory effect on CaM-supported K-ras membrane organization.
Direct comparison of ophiobolin A and conglobatin A indicated that ophiobolin A is more potent in inhibiting mammosphere formation (Figure 7c). Remarkably, ophiobolin A showed a drug-response profile with very similar trends of association with gene expression as salinomycin (P=0.039, Wilcoxon signed rank test, for pairwise difference between the expression correlation of ophiobolin A and STS over the 13 K-ras-nanoclustering signature genes; Supplementary Figure 6e). In particular, the negative trend with negative regulators of K-ras nanoclustering (HRAS, PTRF, Cav-1, Cav-2) was similar to what we found for salinomycin (Figure 5e). In addition, also a positive trend with positive regulators (NCL, NPM1) was seen, which may relate to its higher potency.
To understand the structural determinants for the potency of ophiobolin A, we established a small structure activity relationship series by testing two additional ophiobolins. While ophiobolin A and B had a similar potential in reducing the CD44+/CD24− CSC population, ophiobolin C had almost lost its activity (Figure 7d). These results therefore tentatively suggest that oxygen 13 is required for the anti-CSC activity of ophiobolin A and B.
In conclusion, our differential nanoclustering-FRET-based screening approach enabled us to identify ophiobolin A as a highly potent CSC inhibitor. We furthermore show that CaM levels affect mammosphere formation and that CaM mediates the ophiobolin A effect on K-ras.
Discussion
We have established that salinomycin and other known and potentially new CSC inhibitors specifically block K-ras4B, but not H-ras activity. The potency of two of these drugs, salinomcyin and ophiobolin A, depends on the expression of other factors that impact on the nanoscale membrane organization of K-ras4B, in particular on caveola levels. Our most potent novel inhibitor, ophiobolin A, mediates its anti-CSC potential by inhibiting CaM-dependent K-ras4B membrane organization, strongly supporting a critical role for the complex of CaM and K-ras4B alone or in complex with PI3Kα as a specific vulnerability in cancer.45, 46
Our data suggest that CSC inhibitors need to discriminate between H-ras and K-ras4B. This is in line with distinct roles of K-ras4B and H-ras in stem cells, with the former governing stem cell self-renewal and the latter differentiation.47 Importantly, this activity was associated with the Ras C-terminus, which also dictates their nanoscale lateral segregation.29 Lateral segregation is maintained within a very defined PS-concentration range.18 It is tempting to speculate that the PS-mediated K-ras4B inhibitory activity of GTP-H-ras (H-rasG12V) affects stem cell fate. This molecular mechanism could even provide a new lead to understand the dominating role of mutant K-ras in cancers. Curiously, N-ras, which is the second most frequently mutated Ras isoform, appears to be neutral for stem cell differentiation.47 However, recent data indicate that N-ras bimodally stimulates symmetrical self-renewal of hematopoetic stem cells in one subset of cells and in another subset asymmetric division.48 Thus, N-ras would combine functionalities of K-ras4B and H-ras. Moreover, based on our Ras-isoform specificity analysis,34 we would ascribe a similar function to K-ras4A. In line with this idea, the two-state function of K-ras4A could be due to different lipid modification states that either mimic K-ras4B or N-ras.49 Though direct experimental validation is still missing, it can be expected that such a bimodal activity of K-ras4A would effectively be masked, considering that it is equally expressed with K-ras4B.50
There is still considerable debate about whether CSC emerge from stem cells or through EMT. Clearly, EMT induction can generate cells with (cancer) stem cell characteristics.2, 6, 51 It is possible that our K-ras-centric mechanism would apply also in the EMT context, as K-ras signaling is at the apex of EMT, where it controls Wnt pathway induction.52 This may also explain why the Wnt pathway was previously reported as a salinomycin target.10 On the other hand, very recent data demonstrate that suppression of the non-canonical Wnt/Ca2+ pathway is essential for the tumorigenic potential of mutant K-ras.45
Our second important finding implies that loss of caveolae can sensitize cells in a specific (K-ras-nanoclustering) gene expression context not only to salinomycin, but also to ophiobolin A, which does not act via PS but via CaM on K-ras membrane organization. This significant role of caveolae is consistent with either an EMT or stem cell model for CSC, as both cells that underwent EMT,53 as well as stem cells, possess low Cav-1 or caveolae levels.54 In fact, loss of Cav-1 in stromal breast tumors can expand the stem cell compartment.55 The loss of stromal Cav-1 is also associated with a poorer cancer prognosis,56 consistent with a critical role of caveolae in CSCs. A similar situation exists in prostate cancer, where PTRF/cavin-1, the other major caveolar protein, is typically lost with higher aggressiveness. Cav-1 is downregulated in tumor cells at early stages enhancing proliferation and later upregulated in metastatic stages. By contrast, it is lost in stromal cells, which correlates also with higher Gleason score.57 These genetic observations support a predominant role of caveolae (via PS) and not of individual caveolar proteins in cancer. It may therefore also be timely to reconsider the ascribed inhibitory function of Cav-1, which is in addition not plausible from a structural point of view.58
Of note, salinomycin, nigericin and lasalocid are insecticidal, antihelmintic and anticoccidial (active against protozoa), as are many of the compounds that were found as primary hits in our second screen. Intriguingly, this toxicity spectrum seems to correlate with the absence of caveolae.59 This may also explain the relatively high hit rate of ∼8% that we have observed with our library of microbial metabolites and related semi-synthetics, as microbes share an eco-niche with nematodes and protozoa and will therefore have evolved defensive strategies against them. While the compounds in the library displayed diverse biological activities, a high proportion displays no chemotherapeutic activity. In line with this, toxicity per se does not qualify a potent compound in our screen, because metabolites with little cell toxicity such as the avermectins and conglobatin A are active.
Unfortunately, the lack of caveolae in stem cells is also suggestive for a general stem cell toxicity of CSC inhibitors. This has actually been observed for salinomycin,60 and consistently salinomycin has a low therapeutic index.61 However, our K-ras-nanoclustering signature may be of particular relevance for the stratification of a significant fraction of patients (at least 8% based on our TCGA analysis) that would be predisposed to a higher mortality and be the most promising responders to salinomycin and other CSC inhibitors. Analysis of the actual tumor types that were characterized by an ESC-like expression signature, revealed an enrichment of this signature in particular in breast invasive carcinoma, ovarian serous cystadenocarcinoma and uterine corpus endometrial carcinoma and carcinosarcomas, all tumors associated with the reproductive system of women (Supplementary Table 4). This is surprising, given that the signature originated from ESCs, but is in excellent agreement with the screening approach that identified salinomycin using mammary-derived cell lines.6 The only other cancer type that reached a similarly high and also specific proportion in our ESC-like set was acute myeloid leukemia. In line with this, a stemness gene signature that is shared between CSC of acute myeloid leukemia and hematopoetic stem cells was shown to predict patient survival.62 Interestingly, this signature shared Cav-1 with our signature.
Owing to their defining role for cell fate, transcription factors have taken center stage in defining cell lineages and stemness.63 However, signaling processes, which give rise to transcriptional changes and would therefore be highly relevant for chemical reprogramming, have lagged behind. The K-ras-associated, ESC-derived gene set, which defined CSC inhibitor sensitivity, may furthermore have profound implications for our understanding of cellular differentiation. It implies that loss of caveolae through membrane stretching,64 which may occur for instance during morphogenetic events in the embryo, could contribute to adhesion matrix-dependent programming of stem cells.65 Our newly identified candidate CSC inhibitors may therefore not only represent new starting points for K-ras-drug and CSC inhibitor development, but also for chemical reprogramming factors.
Materials and methods
DNA constructs and molecular cloning
Plasmids encoding the C-terminal hypervariable region of human H-ras (H-ras NANOPS; previously referred to as CTH) and C-terminal hypervariable region of K-ras (K-ras NANOPS; previously referred to as CTK) genetically fused to monomeric Cyan Fluorescent Protein and monomeric Citrine (mCitrine) have been previously described.20, 29 pmGFP-H-rasG12V, pmGFP-K-ras4BG12V and mRFP-RBD of C-Raf have been previously described.29, 34 pmCherry-H-rasG12V and pmCherry-K-rasG12V constructs were generated by replacing pmGFP from pmGFP-H-rasG12V and pmGFP-K-rasG12V with pmCherry from pmCherry-C1 vector (Clontech Laboratories Inc., Mountain View, CA, USA) using NheI and BsrGI restriction sites. Plasmid pcDNA3.1+N-HA with wild-type human Calmodulin1 (NM 006888) (wild-type-CaM) and mutant CaM plasmid (mut-CaM) with substitutions K75Q, K77Q and K148Q were generated by GenScript USA Inc. (Piscataway, NJ, USA). GFP-LactC2 and RFP-LactC2 plasmids were kind gifts from Prof John Hancock (University of Texas, Texas). Untagged PTRF-Flag and Cav-1-HA constructs have been described elsewhere.35
Cell lines and cell culture
Baby Hamster Kidney (BHK21), HEK293 EBNA,66 MDA-MB-436, Hs578T and MCF7 cells were cultured in Dulbecco’s modified Eagle’s medium (Sigma–Aldrich, Helsinki, Finland, Cat. No. D6171), containing 10% fetal bovine serum (PromoCell, Heidelberg, Germany, Cat. No. C-37360), 100 U/ml penicillin (Sigma–Aldrich), 100 μg/ml streptomycin (Sigma–Aldrich), L-glutamine (Sigma–Aldrich, Cat. No. G7513). MDA-MB-231 were cultured in RPMI supplemented with 10% horse serum, 5% fetal bovine serum, L-glutamine, penicillin (100 U/ml) and streptomycin (100 μg/ml). All cells were and incubated at 37 °C with 5% CO2. Cells were grown to confluency of 80% (8 × 107 cells) and subcultured every 2–3 days.
MDCK (Madin-Darby canine kidney) cells stably expressing either mGFP-K- or H-rasG12V were kind gifts from Prof John Hancock (University of Texas, Texas).32 Cancer cell lines used for sensitivity testing were cultured and maintained by GenScript human tumor cell line profiling services.
Chemical screening with Ras-NANOPS, data acquisition and analysis
Chemical screens were performed as recently described.20, 21 In brief, BHK cells were seeded in six-well plates and transfected with Ras-NANOPS FRET-pair plasmids using jetPRIME (Polyplus transfection, Illkrich-Graffenstaden, France) following the manufacturer’s instructions. Eighteen hours post transfection, cells were transferred to clear flat bottom 96-well plates with 50 000 cells per well with complete Dulbecco’s modified Eagle’s medium. After the cells have attached (usually 5–7 h), they were treated for 24 h with compounds. The compound stocks were typically prepared in 100% DMSO (Sigma, D2438) and stored at −20 °C until use, unless indicated otherwise. Prior to use, the compounds were brought to room temperature and were diluted to the final concentration in the medium with a final DMSO concentration of less than 0.1%. The ionophore collection was screened at a final drug concentration of 1 μg/ml and the Microbial Screening Technologies (MST) Library was screened at a final concentration of 0.25 μg/ml in the growth medium. After treatment, cells were detached with 75 μl of 10 mM EDTA in phosphate-buffered saline and fixed using an equal volume of 4% paraformaldehyde (PFA; Sigma, Cat. No. P6148) for 15 min at room temperature. The samples were stored at 4 °C until analysis. The libraries were all screened in three independent experiments and each 96-well plate was designed to have an internal compactin control. The cross-validation screen with HEK cells was performed as described above. Fixed cells were analyzed using the FACS LSR II (BD Biosciences, San Jose, CA, USA) using a high throughput sampler with the following filter settings for donor (405 nm excitation, 440/40 nm emission), acceptor (488 nm excitation, 530/30 nm emission) and FRET (405 nm excitation, 584/42 nm emission) filter. The acquired cytometry data were analyzed using a custom written procedure in IgorPro5 (Wavemetrics, Portland, OR, USA).29, 30 FITC beads (Bangs Laboratories, Fishers, IN, USA, MESF, pH 7.2, Cat. No. 1187) with a size (7.56 μm) and defined 1 401 242 FITC-equivalents were used for normalized acceptor level calibration. The FITC-beads fluorescence intensity at 488 nm excitation was correlated with mCitrine intensity of the FRET sample. The monomeric Cyan Fluorescent Protein-mCitrine fusion protein was used to calibrate the FRET efficiency and donor–acceptor ratio. The FRET Emax was calculated as earlier described.29, 30 The FRET Emax values were averaged over the plate and a 15% change in the FRET Emax compared with plate average was used as the primary criteria for hit selection.
Confocal and STED microscopy
MDCK cells stably expressing either mGFP-H-rasG12V or mGFP-K-rasG12V and BHK cells transiently expressing either mGFP-H-rasG12V or mGFP-K-rasG12V were treated for 24 h with 1.3 μM salinomycin, 1.3 μM nigericin and 1.7 μM lasalocid sodium, 10 nM STS or 0.2 μM avermectin, ivermectin, conglobatin A, ophiobolin A, leptomycin B, kazusamycin B and streptonigrin. Cells were fixed 24 h after treatments with 4% PFA and mounted with Mowiol 4–88 (Sigma–Aldrich). Cells were imaged using a Zeiss LSM 780 confocal microscope (Carl Zeiss AG, Jena, Germany) with a 488 nm wavelength Argon laser (2% of nominal power 35 mW) used for EGFP (excitation: 488 nm, detection: 493–575 nm). Bidirectional scanning and 8 × line averaging was used on a 1024 × 1024 resolution with zoom adjusted according to the cell of interest. Images were treated using ImageJ 1.49n (National Institutes of Health, Bethesda, MD, USA) to adjust for contrast and to display them on an inverted gray-scale. For STED microscopy of Cav-1-positive caveolar structures, cells were fixed in 4% PFA and permabilized with 0.2% saponin and blocked with 0.2% bovine serum albumin before incubation with anti-Cav-1 antibody (BD Biosciences, Cat. No. 610060). Cells were then incubated with anti-rabbit IgG-Star-635 (Abberior GmbH, Göttingen, Germany, Cat. No. 2-0012-002-7) secondary antibody and mounted in mowiol 4–88 (Sigma–Aldrich). Images were acquired on an inverted confocal laser-scanning microscope (Leica SP5 STED, Leica Microsystems GmbH, Wetzlar, Germany). Excitation source was a 635 nm pulsed laser (PicoQuant LDH-P-C- 640B, Berlin, Germany) and fluorescence was collected with an avalanche photodiode (APD) detector at 665–705 nm range. Depletion was carried out with a pulsed Titanium:Sapphire laser MaiTai HP (Spectra-Physics, Santa Clara, CA, USA) delivering femtosecond pulses at a rate of 80 MHz with an output power of 2.2 watts for the selected wavelength of 740 nm. Images were acquired with a silicone oil immersion objective lens (Leica NA1.4 × 100) and the confocal pinhole was set to one airy unit, with a scan speed of 700 Hz, and line averaging of 8. Sampling was carried out on a 1024 × 1024 format and using a × 10 zoom allowing sufficient magnification to observe individual caveolae while keeping the region of interest large enough for quantification of a significant area of the cell membrane. Caveolae number quantification was carried out with ImageJ 1.49n (National Institutes of Health) using the built-in functions of threshold and analyze particles. For particle detection, we set a size range of 0.005–0.3 μm2 that excludes free molecules of Cav-1 and large agglomerates that do not correspond to actual caveolae. Number of caveolae were quantified and divided by the membrane area included in each image to obtain the density of caveolae. For each condition, a minimum of 10 cells were imaged and quantified.
FRET-imaging using fluorescence lifetime microscopy (FLIM)
Cells were transfected using jetPRIME transfection reagent (Polyplus) with the donor alone (mGFP-tagged constructs) in control samples, or mGFP-tagged together with the acceptor. Acceptors were mCherry-tagged Ras constructs in nanoclustering-FRET experiments (pmGFP:pmCherry plasmids at 1:3 ratio, 2 μg total plasmid), mRFP-RBD in C-Raf-RBD-recruitment (1:3 plasmid ratio, 2 μg total plasmid) FRET experiments. Caveolae level manipulations were performed by co-expressing untagged PTRF-Flag and Cav-1-HA plasmids (1:3:1.5:1.5 plasmid ratio, 3.5 μg total plasmid). For CaM level manipulation experiments, HEK cells were transfected with pmGFP- and pmCherry-K-rasG12V with either pcDNA3.1-wild-type-CaM or pcDNA3.1-mut-CaM (1:3:3 plasmid ratio, 3.5 μg total plasmid) together with 60 nM of CaM siRNA (Qiagen, Germantown, MD, USA, Cat. No. 1027416) targeting the non-coding region of Calmodulin 1. Cells were treated 24 h after transfection with either DMSO control, 1.3 μM salinomycin, 1.3 μM nigericin, 1.7 μM lasalocid, 10 nM STS or 0.2 μM avermectin, ivermectin, conglobatin A, ophiobolin A, leptomycin B, kazusamycin B and streptonigrin. The final DMSO concentration in cells was kept under 0.1%. Cells were fixed 24 h after treatments with 4% PFA and mounted with Mowiol 4–88 (Sigma–Aldrich). The mGFP fluorescence lifetime was measured using a fluorescence lifetime imaging attachment (Lambert Instruments, Groningen, The Netherlands) on an inverted microscope (Zeiss AXIO Observer D1). For the sample excitation, sinusoidally modulated 3 W, 497 nm LED at 40 MHz under epi-illumination was used. Cells were imaged using the × 63, NA1.4 oil objective with the GFP filter set (excitation: BP 470/40, beam splitter: FT 495, emission: BP 525/50). The phase and modulation were determined using the manufacturer’s software from images acquired at 12 phase settings. Fluorescein at 0.01 mM, pH 9.0 was used as a lifetime reference standard.67 For each treatment condition, the fluorescence lifetime was measured typically for >60 cells from three biological repeats. From the obtained fluorescence lifetimes, the apparent FRET efficiency was calculated as described.17
Western blotting analysis
BHK21 cells were transfected with mGFP-K-rasG12V or mGFP-H-rasG12V using jetPRIME transfection reagent. Twenty-four hours after transfection, cells were serum-starved for 5 h and treated with either DMSO control, 1.3 μM salinomycin, 1.3 μM nigericin or 1.7 μM lasalocid for 24 h. For Calmodulin siRNA experiments, MDA-MB-231 cells were transfected with CaM siRNA (Qiagen, Cat. No. 1027416) using Lipofectamine 3000 (Thermo-Fischer Scientific, Grand Island, NY, USA) according to the manufacturer's instructions for 24 h. Following treatment or transfection, cells were harvested in Laemmli buffer with 0.1% bromophenol blue and resolved on a 10% gel and transferred to Protran nitrocellulose membrane (Perkin Elmer, Waltham, MA, USA). Membranes were blocked with 5% milk in TBS+0.1% Tween-20 and immunolabelled using primary antibodies against phospho-p44/42 MAP kinase (Thr2202/Tyr204; Cell Signaling, Danvers, MA, USA, Cat. No. 9101), p44/42 MAP kinase (Cell Signaling, Cat. No. 9102), GFP (BioVision, Milpitas, CA, USA, Cat. No. 3999-100), Oct3/4 (Santa Cruz, Paso Robles, CA, USA, sc-9081), Sox2 (Cat. No. C70B1, Cell Signaling), NANOG (R&D systems, Wiesbaden, Germany, Cat. No. AF1997) and Calmodulin (Cell Signaling, Cat. No. 4830). β-Actin (clone AC-15; Sigma–Aldrich, Cat. No. A1978) was used as a loading control. Goat-anti-rabbit immunoglobulins/HRP (Santa Cruz, Cat. No. sc-2004) and chicken-anti-mouse immunoglobulins/HRP (Santa Cruz, Cat. No. sc-2954) were used as secondary antibodies. The bands were detected with enhanced chemiluminescence (Bio-Rad, Helsinki, Finland) and densitometrically analyzed with ImageLab Software (Bio-Rad).
Cell proliferation and viability assays
For proliferation assays of HEK or BHK, cells were seeded in 96-well plates and treated after 24 h with either DMSO control, 1.3 μM salinomycin, 1.3 μM nigericin or 1.7 μM lasalocid for 3 days. DMSO control samples were matched with the highest DMSO concentration of the inhibitors and typically the final DMSO concentration in cell experiments was kept below 0.1%. Cell proliferation was measured as change in fluorescence intensity every 24 h using alamarBlue (Invitrogen, Carlsbad, CA, USA, Cat. No. DAL1100) following the manufacturer’s instructions. Fluorescence intensity was measured at 570 nm excitation and 590 nm emission on a Synergy H1 Multi-Mode Reader (BioTek, Winooski, VT, USA). Each treatment condition consisted of at least three replicate wells and was repeated three times. The fluorescence intensities at each time point was compared with the DMSO control and data were expressed as change in cell proliferation relative to the control. For dose–response assays, cells were seeded in 96-well plates and treated with various concentrations of the inhibitors. For each inhibitor concentration, three to six replicate wells were measured and were repeated three times. After incubation, cell viability was measured using alamarBlue. The fluorescence intensities of treated cells were compared with the DMSO control and the data were expressed as cell viability relative to the control. Drug sensitivity profiling of tumor cell lines was performed through GenScript human tumor cell line profiling services. Briefly, cancer cell lines were plated on 384-well plates and treated for 3 days with various concentrations of the inhibitors. After incubation, viability was measured using CellTiter-Glo (Promega BioSciences, San Luis Obispo, CA, USA) reagent following the manufacturer's instructions. Luminescence was measured on a PHERAstar Plus microplate reader (BMG Labtech, Ortenberg, Germany). Data analysis was performed using GraphPad Prism.
Flow cytometric analysis of CD44+/CD24− cells
MDA-MB-231 cells treated with inhibitors were stained with APC-conjugated anti-CD44 (clone G44-26, BD Biosciences) antibody and PE-conjugated anti-CD24 antibody (clone ML5, BD Biosciences) following the manufacturer’s instructions. Briefly, treated cells were washed with phosphate-buffered saline and harvested in 0.05% trypsin/0.025% EDTA. Detached cells were washed with phosphate-buffered saline containing 2% fetal calf serum and 0.1% sodium azide (FACS buffer 1) and resuspended in the FACS buffer 1 at 106 cells/100 μl. Fluorochrome-conjugated monoclonal antibodies were added to the cell suspensions at concentrations recommended by the manufacturer and incubated at 4 °C in the dark for 30–40 min. After staining, cells were washed with phosphate-buffered saline containing 0.1% sodium azide (FACS buffer 2), fixed using 4% PFA and cytometric analysis was performed in a FACS LSR II (BD Biosciences) cytometer in accordance with the manufacturer’s protocols. In brief, unstained cells, CD44, CD24 and double-stained control cells were used to mark the four quadrants in a dot-plot for unstained, CD44+, CD24+ and double-positive populations. The change in percentage of CD44+/CD24− cells (top left quadrant) was measured in DMSO control and inhibitor treated samples.
Mammosphere assay
Mammosphere formation assays were performed using MDA-MB-231, MDA-MB-436, Hs578T and MCF7 cells. The cells were cultured in 48-well suspension culture plates (Cellstar, Greiner Bio-One, Frickenhausen, Germany) at an initial density of 2000 cells per well in serum-free media supplemented with 1 × B27 (Gibco, Thermo Fisher Scientific), 25 ng/ml EGF (Sigma) and 25 ng/ml FGF (Sigma). After 6 days in culture, cells were treated for additional 3 days with the indicated inhibitors. For CaM RNAi experiments, cells were first seeded in six-well plates and transfected with either 60 nM scrambled siRNA or siRNA targeting Calmodulin 1 (Qiagen, Cat. No. 1027416) and 24 h later, they were transferred to 48-well suspension culture plates and grown as mentioned above. The mammospheres were analyzed in an Evos FL microscope (Thermo Fisher Scientific) and spheres with a minimum size of at least 50 μm were counted. The sphere formation under different treatments was expressed as percentage normalized to the vehicle treated control.
Statistical analysis
Unless otherwise stated, statistical differences were determined using an analysis of variance (ANOVA) complemented by Tukey’s honest significant difference test (Tukey’s HSD). The GraphPad PRISM software was used to perform these analyses. Statistical significance levels are annotated as n.s.=non significant, *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001.
Dose–response analysis
The half maximal inhibitory concentrations (IC50) of the compounds was calculated in GraphPad Prism using the nonlinear regression analysis of log (inhibitor concentration) vs drug response (four parameter logistic function) with the equation:

where y is the parameter quantifying the relative cell viability, x is the log of inhibitor dose or concentration. The fitting parameters a and b correspond to the lower and upper limits of the cell viability, respectively. IC50 values of tumor cell lines used for drug sensitivity profiling were acquired and analyzed by GenScript using the same equation as above.
Selection of co-expressed genes
Whole-genome gene expression data for ESCs and Fibroblast cells were obtained from two meta-sets: GeneChip Human Genome U133 Plus 2.0 Array (408 samples) and Illumina (San Diego, CA, USA) Sentrix HumanRef-8 Expression BeadChip (245 samples) available in the ESTOOLs database (http://estools.cs.tut.fi/index.php).39 Gene expression data were normalized for batch effects using ComBat package available in R. Spearman correlation analysis was performed to find additional set of genes co-expressed with the initial set of 10 genes. Briefly, we first selected the 40 most correlated genes in expression pattern with each of the 10 genes in ESCs and fibroblasts extracted from ESTOOLS database. This set of genes was filtered to keep genes that were correlated with more than one gene in the initial set, resulting in a total of 32 co-expressed genes. Unsupervised hierarchical clustering was performed using the selected set of genes representing K-ras-nanoclustering signature, with distance measure as Pearson correlation and using the average linkage algorithm in R version 3.1.2.
Identification of cancer cell lines with K-ras-nanoclustering signature
Gene expression data for cancer cell lines was extracted from the Cancer Cell Line Encyclopedia (CCLE) resource (http://www.broadinstitute.org/ccle/home).41 On the basis of gene expression data of the 13 genes representing the K-ras-nanoclustering signature, Spearman correlation coefficients were calculated between the cancer cell lines and the ESCs and fibroblasts. Rank product analysis was performed to find the most consistently top-ranked (positively correlated) cancer cell lines for ESCs and fibroblasts, separately. Top 30 most correlated cancer cell lines for the two groups were considered further, and 15 cancer cell lines (9 ESC-like and 6 fibroblast-like) were selected for drug testing.
Identification of clinical samples with K-ras-nanoclustering signature
Whole-genome gene expression and clinical data for patients was obtained from The Cancer Genome Atlas project using TCGA Assembler.68 For selecting samples exhibiting the ESC-like K-ras-nanoclustering signature pattern, the patient’s expression signature was correlated (Spearman correlation) with 20 randomly chosen primed ESC cell lines from the ESTOOLs database and averaged. A total of 605 patients had average correlation⩾0.6 with ESC-like-K-ras-nanoclustering signature pattern (8% of the TCGA samples). Similar correlation analysis was performed for the fibroblast-like-K-ras-nanoclustering signature pattern, resulting in 402 subjects with average correlation⩾0.6 with Fibroblast-like-K-ras-nanoclustering signature pattern. Kaplan–Meier analysis and Univariate Cox proportional hazard test was performed using R-package survival to assess the difference in overall survival between the two groups of TCGA samples: ESC-like and Fibroblast-like. Similarly, we also compared subjects that exhibited expression pattern similar to those observed in the ESC-like-cancer cell lines and fibroblast-like-cancer cell lines among the 15 cell lines screened. Subjects that had average Spearman correlation⩾0.6 with the ESC-like-cancer cell lines (n=1401) were compared with subjects that had average spearman correlation⩾0.6 with the fibroblast-like cancer cell lines (n=2631).
References
Kreso A, Dick JE . Evolution of the cancer stem cell model. Cell Stem Cell 2014; 14: 275–291.
Ansieau S . EMT in breast cancer stem cell generation. Cancer Lett 2013; 338: 63–68.
Medema JP . Cancer stem cells: The challenges ahead. Nat Cell Biol 2013; 15: 338–344.
Dean M, Fojo T, Bates S . Tumour stem cells and drug resistance. Nat Rev Cancer 2005; 5: 275–284.
Singh A, Settleman J . EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene 2010; 29: 4741–4751.
Gupta PB, Onder TT, Jiang G, Tao K, Kuperwasser C, Weinberg RA et al. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 2009; 138: 645–659.
Zhou S, Wang F, Wong ET, Fonkem E, Hsieh T-C, Wu JM et al. Salinomycin: a novel anti-cancer agent with known anti-coccidial activities. Curr Med Chem 2013; 20: 4095–4101.
Naujokat C, Steinhart R . Salinomycin as a drug for targeting human cancer stem cells. J Biomed Biotechnol 2012; 2012: 950658.
Riccioni R, Dupuis ML, Bernabei M, Petrucci E, Pasquini L, Mariani G et al. The cancer stem cell selective inhibitor salinomycin is a p-glycoprotein inhibitor. Blood Cells Mol Dis 2010; 45: 86–92.
Lu D, Choi MY, Yu J, Castro JE, Kipps TJ, Carson DA . Salinomycin inhibits Wnt signaling and selectively induces apoptosis in chronic lymphocytic leukemia cells. Proc Natl Acad Sci USA 2011; 108: 13253–13257.
Fuchs D, Heinold A, Opelz G, Daniel V, Naujokat C . Salinomycin induces apoptosis and overcomes apoptosis resistance in human cancer cells. Biochem Biophys Res Commun 2009; 390: 743–749.
Hanahan D, Weinberg RA . Hallmarks of cancer: the next generation. Cell 2011; 144: 646–674.
Hancock JF . Ras proteins: different signals from different locations. Nat Rev Mol Cell Biol 2003; 4: 373–385.
Spiegel J, Cromm PM, Zimmermann G, Grossmann TN, Waldmann H . Small-molecule modulation of Ras signaling. Nat Chem Biol 2014; 10: 613–622.
Abankwa D, Gorfe AA, Hancock JF . Ras nanoclusters: molecular structure and assembly. Semin Cell Dev Biol 2007; 18: 599–607.
Rotblat B, Belanis L, Liang H, Haklai R, Elad-Zefadia G, Hancock JF et al. H-Ras nanocluster stability regulates the magnitude of MAPK signal output. PLoS One 2010; 5: e11991.
Guzmán C, Solman M, Ligabue A, Blazevitš O, Andrade DM, Reymond L et al. The efficacy of Raf kinase recruitment to the GTPase H-ras depends on H-ras membrane conformer-specific nanoclustering. J Biol Chem 2014; 289: 9519–9533.
Zhou Y, Liang H, Rodkey T, Ariotti N, Parton RG, Hancock JF . Signal integration by lipid-mediated spatial cross talk between Ras nanoclusters. Mol Cell Biol 2014; 34: 862–876.
Ariotti N, Fernández-Rojo MA, Zhou Y, Hill MM, Rodkey TL, Inder KL et al. Caveolae regulate the nanoscale organization of the plasma membrane to remotely control Ras signaling. J Cell Biol 2014; 204: 777–792.
Köhnke M, Schmitt S, Ariotti N, Piggott AM, Parton RG, Lacey E et al. Design and application of in vivo FRET biosensors to identify protein prenylation and nanoclustering inhibitors. Chem Biol 2012; 19: 866–874.
Najumudeen AK, Köhnke M, Solman M, Alexandrov K, Abankwa D . Cellular FRET-biosensors to detect membrane targeting inhibitors of N-myristoylated proteins. PLoS One 2013; 8: e66425.
Najumudeen AK, Posada IMD, Lectez B, Zhou Y, Landor SK-J, Fallarero A et al. Phenotypic screening identifies protein synthesis inhibitors as H-Ras-nanocluster-increasing tumor growth inducers. Biochemistry 2015; 54: 7212–7221.
Coxon F, Joachimiak Ł, Najumudeen AK, Breen G, Gmach J, Oetken-Lindholm C et al. Synthesis and characterization of novel phosphonocarboxylate inhibitors of RGGT. Eur J Med Chem 2014; 84: 77–89.
Crouthamel M, Abankwa D, Zhang L, DiLizio C, Manning DR, Hancock JF et al. An N-terminal polybasic motif of Gαq is required for signaling and influences membrane nanodomain distribution. Mol Pharmacol 2010; 78: 767–777.
Sykes AM, Palstra N, Abankwa D, Hill JM, Skeldal S, Matusica D et al. The effects of transmembrane sequence and dimerization on cleavage of the p75 neurotrophin receptor by gamma-secretase. J Biol Chem 2012; 287: 43810–43824.
Tian T, Harding A, Inder K, Plowman S, Parton RG, Hancock JF . Plasma membrane nanoswitches generate high-fidelity Ras signal transduction. Nat Cell Biol 2007; 9: 905–914.
Solman M, Ligabue A, Blazevitš O, Jaiswal A, Zhou Y, Liang H et al. Specific cancer-associated mutations in the switch III region of Ras increase tumorigenicity by nanocluster augmentation. eLife 2015; 4: e08905.
Zhou Y, Hancock JF . Ras nanoclusters: Versatile lipid-based signaling platforms. Biochim Biophys Acta 2015; 1853: 841–849.
Abankwa D, Hanzal-Bayer M, Ariotti N, Plowman SJ, Gorfe AA, Parton RG et al. A novel switch region regulates H-ras membrane orientation and signal output. EMBO J 2008; 27: 727–735.
Abankwa D, Vogel H . A FRET map of membrane anchors suggests distinct microdomains of heterotrimeric G proteins. J Cell Sci 2007; 120: 2953–2962.
Huczyński A . Polyether ionophores—promising bioactive molecules for cancer therapy. Bioorg Med Chem Lett 2012; 22: 7002–7010.
Cho K-J, Park J-H, Piggott AM, Salim AA, Gorfe AA, Parton RG et al. Staurosporines disrupt phosphatidylserine trafficking and mislocalize Ras proteins. J Biol Chem 2012; 287: 43573–43584.
Fairn GD, Schieber NL, Ariotti N, Murphy S, Kuerschner L, Webb RI et al. High-resolution mapping reveals topologically distinct cellular pools of phosphatidylserine. J Cell Biol 2011; 194: 257–275.
Abankwa D, Gorfe AA, Inder K, Hancock JF . Ras membrane orientation and nanodomain localization generate isoform diversity. Proc Natl Acad Sci USA 2010; 107: 1130–1135.
Hill MM, Bastiani M, Luetterforst R, Kirkham M, Kirkham A, Nixon SJ et al. PTRF-Cavin, a conserved cytoplasmic protein required for caveola formation and function. Cell 2008; 132: 113–124.
Bastiani M, Liu L, Hill MM, Jedrychowski MP, Nixon SJ, Lo HP et al. MURC/Cavin-4 and cavin family members form tissue-specific caveolar complexes. J Cell Biol 2009; 185: 1259–1273.
Hell SW . Far-field optical nanoscopy. Science (New York, NY) 2007; 316: 1153–1158.
Seguin L, Kato S, Franovic A, Camargo MF, Lesperance J, Elliott KC et al. An integrin β3-KRAS-RalB complex drives tumour stemness and resistance to EGFR inhibition. Nat Cell Biol 2014; 16: 457–468.
Kong L, Aho K-L, Granberg K, Lund R, Järvenpää L, Seppälä J et al. ESTOOLS Data@Hand: human stem cell gene expression resource. Nat Methods 2013; 10: 814–815.
Blair K, Wray J, Smith A . The liberation of embryonic stem cells. PLoS Genet 2011; 7: e1002019.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012; 483: 603–607.
Sheridan C, Kishimoto H, Fuchs RK, Mehrotra S, Bhat-Nakshatri P, Turner CH et al. CD44+/CD24-breast cancer cells exhibit enhanced invasive properties: an early step necessary for metastasis. Breast Cancer Res 2006; 8: R59.
Dontu G, Abdallah WM, Foley JM, Jackson KW, Clarke MF, Kawamura MJ et al. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev 2003; 17: 1253–1270.
Kong Au T, Chow Leung P . Identification of the binding and inhibition sites in the calmodulin molecule for ophiobolin A by site-directed mutagenesis. Plant Physiol 1998; 118: 965–973.
Wang M-T, Holderfield M, Galeas J, Delrosario R, To MD, Balmain A et al. K-Ras promotes tumorigenicity through suppression of non-canonical Wnt signaling. Cell 2015; 163: 1237–1251.
Nussinov R, Muratcioglu S, Tsai CJ, Jang H, Gursoy A, Keskin O . The key role of calmodulin in KRAS-driven adenocarcinomas. Mol Cancer Res 2015; 13: 1265–1273.
Quinlan MP, Quatela SE, Philips MR, Settleman J . Activated Kras, but not Hras or Nras, may initiate tumors of endodermal origin via stem cell expansion. Mol Cell Biol 2008; 28: 2659–2674.
Li Q, Bohin N, Wen T, Ng V, Magee J, Chen S-C et al. Oncogenic Nras has bimodal effects on stem cells that sustainably increase competitiveness. Nature 2013; 504: 143–147.
Nussinov R, Tsai CJ, Chakrabarti M, Jang H . A new view of Ras isoforms in cancers. Cancer Res 2016; 76: 18–23.
Tsai FD, Lopes MS, Zhou M, Court H, Ponce O, Fiordalisi JJ et al. K-Ras4A splice variant is widely expressed in cancer and uses a hybrid membrane-targeting motif. Proc Natl Acad Sci USA 2015; 112: 779–784.
Mani SA, Guo W, Liao M-J, Eaton EN, Ayyanan A, Zhou AY et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 2008; 133: 704–715.
Lemieux E, Cagnol S, Beaudry K, Carrier J, Rivard N . Oncogenic KRAS signalling promotes the Wnt/[beta]-catenin pathway through LRP6 in colorectal cancer. Oncogene 2014; 34: 4914–4927.
Lu Z, Ghosh S, Wang Z, Hunter T . Downregulation of caveolin-1 function by EGF leads to the loss of E-cadherin, increased transcriptional activity of β-catenin, and enhanced tumor cell invasion. Cancer Cell 2003; 4: 499–515.
Baker N, Tuan RS . The less-often-traveled surface of stem cells: caveolin-1 and caveolae in stem cells, tissue repair and regeneration. Stem Cell Res Ther 2013; 4: 1–1.
Sotgia F, Williams TM, Cohen AW, Minetti C, Pestell RG, Lisanti MP . Caveolin-1-deficient mice have an increased mammary stem cell population with upregulation of Wnt/beta-catenin signaling. Cell Cycle 2005; 4: 1808–1816.
Witkiewicz AK, Dasgupta A, Sotgia F, Mercier I, Pestell RG, Sabel M et al. An absence of stromal caveolin-1 expression predicts early tumor recurrence and poor clinical outcome in human breast cancers. Am J Pathol 2009; 174: 2023–2034.
Nassar ZD, Hill MM, Parton RG, Parat M-O . Caveola-forming proteins caveolin-1 and PTRF in prostate cancer. Nat Rev Urol 2013; 10: 529–536.
Collins BM, Davis MJ, Hancock JF, Parton RG . Structure-based reassessment of the caveolin signaling model: do caveolae regulate signaling through caveolin-protein interactions? Dev Cell 2012; 23: 11–20.
Kirkham M, Nixon SJ, Howes MT, Abi-Rached L, Wakeham DE, Hanzal-Bayer M et al. Evolutionary analysis and molecular dissection of caveola biogenesis. J Cell Sci 2008; 121: 2075–2086.
Sachlos E, Risueño RM, Laronde S, Shapovalova Z, Lee J-H, Russell J et al. Identification of drugs including a dopamine receptor antagonist that selectively target cancer stem cells. Cell 2012; 149: 1284–1297.
Boehmerle W, Muenzfeld H, Springer A, Huehnchen P, Endres M . Specific targeting of neurotoxic side effects and pharmacological profile of the novel cancer stem cell drug salinomycin in mice. J Mol Med 2014; 92: 889–900.
Eppert K, Takenaka K, Lechman ER, Waldron L, Nilsson B, van Galen P et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat Med 2011; 17: 1086–1093.
Buganim Y, Faddah DA, Jaenisch R . Mechanisms and models of somatic cell reprogramming. Nat Rev Genet 2013; 14: 427–439.
Sinha B, Köster D, Ruez R, Gonnord P, Bastiani M, Abankwa D et al. Cells respond to mechanical stress by rapid disassembly of caveolae. Cell 2011; 144: 402–413.
Engler AJ, Sen S, Sweeney HL, Discher DE . Matrix elasticity directs stem cell lineage specification. Cell 2006; 126: 677–689.
Meissner P, Pick H, Kulangara A, Chatellard P, Friedrich K, Wurm FM . Transient gene expression: recombinant protein production with suspension-adapted HEK293-EBNA cells. Biotechnol Bioeng 2001; 75: 197–203.
Guzmán C, Oetken-Lindholm C, Abankwa D . Automated High-Throughput Fluorescence Lifetime Imaging Microscopy to Detect Protein-Protein Interactions. J Lab Autom e-pub ahead of print 17 September 2015 doi:10.1177/2211068215606048.
Zhu Y, Qiu P, Ji Y . TCGA-Assembler: open-source software for retrieving and processing TCGA data. Nat Methods 2014; 11: 599–600.
Acknowledgements
Plasmids encoding tagged LactC2 and MDCK cells stably expressing either mGFP-K- or H-rasG12V were kindly provided by Prof John Hancock. Plasmids encoding PTRF and Cav-1 were kindly provided by Prof Rob Parton. We thank Lingjia Kong for help with the ESTOOLS database. We thank Jasmin Varjonen for experimental support. This work was supported by the Academy of Finland fellowship grant, the Sigrid Juselius Foundation, the Cancer Society of Finland, the Marie-Curie Reintegration Grant and the Jane and Aatos Erkko Foundation grants to DA, as well as Academy of Finland (grants 272437, 269862, 279163) and the Cancer Society of Finland grants to TA. AKN was supported by the ISB graduate school and AJ by the Integrative Life Science Doctoral Program (ILS).
Author contributions
DA conceived the project. AKN acquired and analyzed FRET and cellular imaging and proliferation data. AJ carried out the bioinformatic and statistical analyses and cancer genome database mining. BL performed and analyzed mammosphere data. CO-L acquired and analyzed cellular and protein expression data. CG acquired and analyzed confocal and STED microscopy data and assisted in statistical analysis. ES performed western blot experiments for MAPK signaling. IP acquired and analyzed FRET experiments for CaM manipulation. EL provided compounds and advised on compound selection. TA supervised and contributed to the bioinformatic analyses. All authors contributed to writing and discussion of the manuscript. AKN and DA designed the experiments and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Najumudeen, A., Jaiswal, A., Lectez, B. et al. Cancer stem cell drugs target K-ras signaling in a stemness context. Oncogene 35, 5248–5262 (2016). https://doi.org/10.1038/onc.2016.59
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.59
This article is cited by
-
An overview of recent advancements in small molecules suppression of oncogenic signaling of K-RAS: an updated review
Molecular Diversity (2024)
-
Epithelial-mesenchymal transition sensitizes breast cancer cells to cell death via the fungus-derived sesterterpenoid ophiobolin A
Scientific Reports (2021)
-
Salinomycin effectively eliminates cancer stem-like cells and obviates hepatic metastasis in uveal melanoma
Molecular Cancer (2019)
-
PIMKL: Pathway-Induced Multiple Kernel Learning
npj Systems Biology and Applications (2019)
-
Cytoprotective effect of neuropeptides on cancer stem cells: vasoactive intestinal peptide-induced antiapoptotic signaling
Cell Death & Disease (2017)