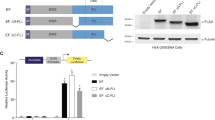
Abstract
Ewing sarcoma is characterized by the expression of the chimeric EWSR1-FLI1 transcription factor. Proteomic analyses indicate that the decrease of EWSR1-FLI1 expression leads to major changes in effectors of the dynamics of the actin cytoskeleton and the adhesion processes with a shift from cell-to-cell to cell-matrix adhesion. These changes are associated with a dramatic increase of in vivo cell migration and invasion potential. Importantly, EWSR1-FLI1 expression, evaluated by single-cell RT-ddPCR/immunofluorescence analyses, and activity, assessed by expression of EWSR1-FLI1 downstream targets, are heterogeneous in cell lines and in tumours and can fluctuate along time in a fully reversible process between EWSR1-FLI1high states, characterized by highly active cell proliferation, and EWSR1-FLI1low states where cells have a strong propensity to migrate, invade and metastasize. This new model of phenotypic plasticity proposes that the dynamic fluctuation of the expression level of a dominant oncogene is an intrinsic characteristic of its oncogenic potential.
Similar content being viewed by others
Introduction
Ewing sarcoma, the second most frequent primary bone tumour among teenagers and young adults, constitutes a highly aggressive tumour1 characterized by early metastatic spread.2 Although current treatment using chemotherapy in addition to local treatment has increased the 5-year survival rate to around 70%,3 the clinical outcome for patients who present metastatic disease, initially or at relapse, remains poor with a long-term survival rate of only 20%.4
In 85% of cases, Ewing sarcoma is characterized by the expression of the EWSR1-FLI1 chimeric protein resulting from the chromosomal translocation t(11;22)(q24:q12), which links the transcription regulating domain of EWSR1 to the ETS DNA-binding domain of FLI1.5 The EWSR1-FLI1 fusion protein behaves as an aberrant transcriptional factor modulating the expression of specific target genes.6 EWSR1-FLI1 expression promotes oncogenesis of Ewing sarcoma as demonstrated by its ability to transform NIH3T3 cells to form tumours in immunodeficient mice.7, 8 Moreover, the invalidation of EWSR1-FLI1 expression by specific si/shRNAs induces an arrest of Ewing sarcoma cell line proliferation in vitro and in vivo (reviewed in Toomey et al.9). However, very few reports have yet addressed the possible role of EWSR1-FLI1 on the mechanisms of metastatic spread and the metastatic process underlying sarcoma remains largely unknown.
Recently, two studies have proposed non-exclusive models of metastatic dissemination for Ewing sarcoma. Krook and colleagues10 have underlined the influence of various cell stresses such as hypoxia, pressure of microenvironment and privation of growth factors to upregulate the CXCR4 chemokine receptor and hence promote migration and invasion properties of Ewing cells. A second model reported as a ‘passive/stochastic metastasis model’ has also been proposed to account for the strong propensity of Ewing cells to disseminate.11, 12 The authors show that EWSR1-FLI1 expression loosens cell adhesion, and they therefore propose that poorly attached Ewing cells passively disseminate in the circulation.11, 13
Here we show that the heterogeneity of EWSR1-FLI1 expression may constitute an essential element of the multistep metastatic process. Indeed, we propose that the cell-to-cell fluctuations of EWSR1-FLI1 expression level constitute a major source of phenotypic heterogeneity and enable individual Ewing cells to switch from proliferation to migration states.
Results
Proteomic analyses of EWSR1-FLI1-regulated proteins
While numerous reports have investigated the transcriptional consequences of the modulation of EWSR1-FLI1 in Ewing sarcoma cells, the Ewing cell proteome upon EWSR1-FLI1 modulation still remains mostly unexplored. In this objective, we developed two proteomic approaches based on 2D-DIGE and SILAC to evaluate EWSR1-FLI1-dependent proteome by mass spectrometry (Supplementary Figures S1A and B). Two different cell systems were used. One consisted in lentivirus-mediated shRNA silencing of EWSR1-FLI1 in Ewing cell lines; the other was based on the construction of clones expressing a doxycycline (DOX) inducible shRNA against the EWSR1-FLI1 fusion. Inducible EWSR1-FLI1 expressing systems were obtained for the A673 (shA673-1c, described in Tirode et al14) and SK-N-MC cells lines (shSK-E17T). In the presence of DOX, the expression level of the EWSR1-FLI1 protein was decreased by 54 and 36% in the shA673-1c and shSK-E17T clones, respectively (Figure 1a). The inhibition of EWSR1-FLI1 is associated with a strong decrease of cell proliferation14 without cell mortality.15 EWSR1-FLI1 re-expression by removal of DOX restores cell proliferation (Supplementary Figures S2A and B).
EWSR1-FLI1low cells demonstrate increased migration and invasion in three-dimensional matrix and in zebrafish. (a) Western blot, (b) phalloidin-stained actin cytoskeleton and (c) measure of cell area of shA673-1c and shSK-E17T clones treated for 10 days (+DOX) with 1 μg/ml DOX or untreated (-DOX) (n=15 per group). Scale bars represent 20 μm. Error bars represent s.e.m. DOX-treated/untreated cells were compared using Welch t-test (***P<0.001). See also Supplementary Figures S1 and S2, Supplementary Tables S1 and S2. (d) Three-dimensional collagen-I multicellular spheroid invasion assay with shA673-1c and shSK-E17T spheroids prepared by the hanging droplet method. 24-h videos were acquired and three time-points are shown. Red dotted lines represent the initial spheroid perimeter. Scale bars represent 100 μm. See also Supplementary Movie S1. (e) Representative pictures of zebrafish xenotransplantation model 4 days post-injection and cumulative results of migration distance of shA673-1c/mCherry cells, DOX-treated or -untreated, from implantation site (X) into the yolk sac for all zebrafish embryos, where each colour represents an individual embryo, and each dot a cluster of one or more tumour cells. Scale bars represent 500 μm. (f) Mean cumulative distance of migration and relative tumour burden of shA673-1c/mCherry cells (n=22 to 27 fish per group), error bars represent 1-way ANOVA with Bonferroni post-test (*P<0.05, ***P<0.001).
Following lentivirus-mediated silencing of EWSR1-FLI1 in the A673 cell line, 2D-DIGE was performed comparing Cy3- and Cy5-labelled A673 infected with either a control shRNA (EWSR1-FLI1high cells) or an EWSR1-FLI1-specific shRNA (EWSR1-FLI1low cells), respectively. A total of 185 differential spots were picked and further investigated by MS/MS. We identified 175 EWSR1-FLI1-modulated proteins (fold change>1.5; P<0.01; Supplementary Table S1). Ingenuity Pathway Analysis was used to mine the data. It showed that the reorganization of actin cytoskeleton and the regulation of actin-based motility by Rho were among the most highly significant differential pathways between both conditions (Supplementary Figure S1C). Differentially expressed proteins include cofilin-1 (CFL1), profilin-2 (PFN2) and gelsolin (GSN) that are involved in the assemblage of actin cytoskeleton; α-actinin (ACTN4), moesin (MSN), radixin (RDX) and vinculin (VCL), which are actin-binding and cytoskeleton-linking proteins; and myosin light chain phosphorylatable (MYLPF), myosin light chain 12A (MYL12A) as well as myosin light chain 6 (MYL6) that participate to the contractile forces necessary to migration processes (Supplementary Figure S1D). These 2D-DIGE results were fully confirmed for the three proteins that were tested by western blot (CFL1, VCL, PFN2 in Supplementary Figure S1E).
In addition to 2D-DIGE data, SILAC experiments were performed on the shA673-1c clone in order to identify membrane proteins modulated by EWSR1-FLI1. Membrane proteins from untreated (EWSR1-FLI1high) or DOX-treated (EWSR1-FLI1low) shA673-1c cells were grown in the presence of heavy or light molecular weight forms of arginine and lysine, respectively. Ingenuity Pathway Analysis of the 131 SILAC-quantified proteins (Supplementary Table S2) allowed us to highlight an enrichment in proteins involved in cell morphology and migration in full agreement with 2D-DIGE data, but also proteins involved in cell interaction processes (Supplementary Figure S1F). Indeed, among this list, at least 10 EWSR1-FLI1-modulated proteins (FC>1.25) are known to be involved in cell–cell or cell–matrix interactions (Supplementary Figure S1G). Interestingly, EWSR1-FLI1 strongly upregulates plakophilin (PKP1) and desmoplakin (DSP), two essential components of the specialized cell-to-cell interaction desmosome structure. At the opposite, it considerably downregulates cell–matrix interaction proteins such as the integrins α1 (ITGA1), α4 (ITGA4), β1 (ITGB1) that are hence highly expressed in EWSR1-FLI1low cells. As for 2D-DIGE, a subset of the variations observed with the SILAC approach was confirmed by western blot in A673, SK-N-MC, EW7 or TC71 cells (Supplementary Figures S1H and I). Further investigating transcriptomics data,16 we also identified intercellular adhesion molecule 1 (ICAM1) and two tight junction proteins, claudin-1 (CLD1) and occludin (OCL), as highly downregulated by EWSR1-FLI1. Altogether, these data strongly suggest that EWSR1-FLI1low cells are less cohesive with more cell–matrix interactions and a stronger cytoskeleton organization as compared to EWSR1-FLI1high cells.
We also confirmed that these variations of protein levels are associated with profound modifications of the cytoskeleton.11, 17 Indeed, we compared the organization of the actin stress fibres in shA673-1c and shSK-E17T clones after 10 days of DOX treatment. Untreated cells have the small round cell morphology typical of the Ewing sarcoma, with thin, short and disorganized actin stress fibres. At the opposite, DOX-treated cells harbour a much larger area of spreading, associated to long and parallel actin stress fibres (Figures 1b and c).
To further explore the phenotype of EWSR1-FLI1low cells, we studied migration and invasion properties of Ewing cells in three-dimensional models. Multicellular spheroids, generated by the natural aggregation of shA673-1c and shSK-E17T cells, pretreated or not by DOX for 10 days, were included in a collagen-matrix and cell migration was monitored by time-lapse video microscopy (Figure 1d). After 24 h, DOX-untreated spheroids presented a stable surface with only very few cells (1 or 2 by field) escaping from the spheroid to migrate through the three-dimensional matrix (Supplementary Movie S1, upper panel). At the opposite, we observed a very rapid migration of cells outside the DOX-treated spheroids, with important protrusions (Supplementary Movie S1, lower panel). After 24 h, in DOX-treated conditions, the collagen-matrix was invaded by cells, through either individual (shA673-1c+DOX) or more collective cell movements (shSK-E17T+DOX).
To confirm three-dimensional migration and invasion in vivo, we used a recently established model of zebrafish embryo xenotransplantation18 and the shA673-1c clone, modified to stably express mCherry protein. After 10 days of DOX-treatment, EWSR1-FLI1low and EWSR1-FLI1high cells were injected into the yolk of 2-days-old embryos. After 4 days, in the presence (n=22 embryos) or absence (n=27) of DOX in the egg water, invasion and proliferation of tumour cells were evaluated by confocal microscopy and cumulative results of each embryo are represented in a scatter plot (Figure 1e), where each dot represents a fluorescent cluster of tumour cells. For invasion, DOX-treated cells displayed a fourfold increase in the cumulative migration distance from injection site (P<0.001). To determine tumour burden per embryo, the number of clusters was multiplied by the average size of clusters. Due to the increased spreading of the cells, more individual clusters were found in the DOX-treated group. However, cluster diameters of the DOX-treated versus untreated group were generally lower (from 27 to 51 μm and 33 to 78 μm, respectively). Resultantly, a significant reduction of the tumour burden was observed in DOX-treated as compared to untreated cells/embryos (Figure 1f).
Taking together with our molecular data, these in vitro and in vivo observations demonstrated clearly that EWSR1-FLI1low cells display proteomic changes associated with an active migration mechanism, which is uncoupled from proliferation of EWSR1-FLI1high cells.
EWSR1-FLI1 expression is heterogeneous in Ewing cell lines
Our results indicate that the experimental modification of EWSR1-FLI1 expression level is sufficient to control the proliferation and migration properties of Ewing cells. Thus, EWSR1-FLI1high cells proliferate, whereas EWSR1-FLI1low cells instead have a strong propensity to migrate. An important question is therefore to define how much this experimental observation is relevant in the context of the human malignancy.
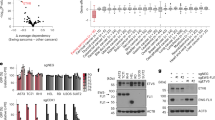
We first explored whether the expression levels of EWSR1-FLI1 transcript and protein are variable from one cell to the other in cell lines or in tumours. We quantified the absolute number of EWSR1-FLI1 and RPLP0 mRNA molecules at the single cell level through droplet digital PCR (single cell RT-ddPCR) in three Ewing cell lines (A673, SK-N-MC and TC71), as well as in DOX-treated/untreated shA673-1c and shSK-E17T clones (Supplementary Figure S3A). This showed that cells with an equivalent number of the housekeeping RPLP0 mRNA molecules may display important variations of EWSR1-FLI1 (Supplementary Figure S3B). Using RPLP0 to normalize EWSR1-FLI1 expression, we observed important variations of EWSR1-FLI1 expression within and across the different cell lines (Figure 2a). Importantly, highly significantly different EWSR1-FLI1 mean expression values (P<0.001) were observed between DOX-treated and untreated shA673-1c or shSKE-17T cells hence providing a validation of single cell measurements. Using the interquartile range of the normalized expression median (EWSR1-FLI1/RPLP0 ratio) of DOX-treated shA673-1c and shSK-E17T clones, we could define an upper threshold for EWSR1-FLI1low status. Interestingly, it is clearly apparent that EWSR1-FLI1low subpopulations of cells can be observed spontaneously in each of the A673, SK-N-MC and TC71 Ewing cell lines. The quantification of the EWSR1-FLI1 protein expression level was also performed using three-dimensional deconvolution microscopy associated with ImageJ software of Ewing cell lines immunostained with an FLI1 antibody (Figure 2b, Supplementary Figure S3C), knowing these Ewing do not express wild-type FLI1. For each cell, the FLI1 signal was normalized by the corresponding DNA signal obtained by DAPI staining.19, 20 As observed with the measurement of the EWSR1-FLI1 mRNA, this experiment fully confirmed the presence of an EWSR1-FLI1low subpopulation in A673 and TC71 cell lines (Figure 2c).
EWSR1-FLI1 expression is heterogeneous in Ewing cell lines. (a) Gene expression profiling at the single cell level by RT-ddPCR of EWSR1-FLI1 and RPLP0 mRNA. Representation of EWSR1-FLI1/RPLP0 ratio in A673, SK-N-MC and TC71 Ewing cell lines; and in shA673-1c, shSK-E17T, shA673-1c+DOX, and shSK-E17T+DOX clones (inducible systems), each dot represents one cell. Bars represent median with interquartile range. The EWSR1-FLI1low threshold is defined by the upper limit of the interquartile range of DOX-treated cells. DOX-treated/untreated cells were compared using Wilcoxon test (***P<0.001). See also Supplementary Figure S2. (b) Protein quantification by immunofluorescence using three-dimensional deconvolution microscopy completed with ImageJ quantification at the single cell level of EWSR1-FLI1 (FLI1) normalized to DNA content (DAPI). Three-dimensional microscopy was performed on Eclipse 90i upright microscope on Nikon Imaging centre (Institut Curie, Paris, France) with a 0.2 μm step for 3D stack and a X100 NA 1.4 oil-immersion objective. The deconvolution of each image stacks was performed automatically using an iterative 7 and algorithm Meinel50 by PICT-IBiSA imaging facility of the Curie Institut, Paris. Scale bar represents 10 μm (c) Representation of EWSR1-FLI1/DAPI ratio in A673 and TC71 Ewing cell lines; and in shA673-1c and shA673-1c+DOX clones (inducible systems), each dot represents one cell. Quantification experiments of the shA673 clone in the −/+ DOX conditions were repeated three times with fully consistent results showing highly significant differences in the quantification of the specific EWSR1-FLI1 staining between both conditions. Bars represent median with interquartile range. The EWSR1-FLI1low threshold is defined by the upper limit of the interquartile range of shA673-1c+DOX cells. DOX-treated/untreated cells were compared using Wilcoxon test (***P<0.001). (d) Characterization of ICAM1− and ICAM1+ subpopulations by FACS analysis of A673 cell line stained with APC anti-ICAM1 (1:500, Biolegend, APC anti-CD54 HA58) and (e) mRNA expression analysis by RT-qPCR of direct EWSR1-FLI1-targets, data are represented as mean +/−s.e.m. (f) Characterization of ICAM1− and ICAM1+ subpopulations at day 0, 20 and 38 days post-sorting.
We then wished to have a more direct measure of EWSR1-FLI1 activity. EWSR1-FLI1 is a transcription factor that can act as an activator or as a repressor depending on the regulated genes. We chose to use ICAM1, the expression of which is inhibited by EWSR1-FLI1 (Supplementary Figure S1H and in Tirode et al14). This protein can be very sensitively detected by FACS analysis and can be used for sorting living cells. A mostly negative expression of ICAM1 was observed by FACS in A673 cells with only a small component of cells being ICAM1 positive (Figure 2d). To assess whether this variable expression of ICAM1 could be related to a low EWSR1-FLI1 expression, we isolated the ICAM1− and ICAM+ subpopulations and analysed the mRNA expression of IGFBP3 and LOX, two targets that are known to be directly repressed by EWSR1-FLI121, 22, 23 (Supplementary Figure S4). Should ICAM1− expression be dependent on EWSR1-FLI1, we would expect IGFBP3 and LOX to be also downregulated in ICAM1− cells whereas an EWSR1-FLI1-independent variability of ICAM1 would not be associated with coordinated variation of LOX and IGFBP3. We observed that the ICAM1 but also the IGFBP3 and LOX transcripts are strongly upregulated in the ICAM1+ subpopulation, a result in full consistency with the three genes being coordinately regulated by EWSR1-FLI1 (Figure 2e). These data indicate that the expression levels of EWSR1-FLI1 downstream targets are heterogeneous in Ewing cell lines and further suggest that this heterogeneity results from cell-to-cell variation in EWSR1-FLI1 activity.
We then addressed the question of the ability of the Ewing cells to switch from an EWSR1-FLI1low to an EWSR1-FLI1high state and vice versa. ICAM1− and ICAM1+ subpopulations were flow-sorted and grown in culture. A new FACS analysis was performed 20 and 38 days after cell sorting and revealed that both cell populations have the ability to regenerate most of the initial heterogeneity therefore indicating that EWSR1-FLI1low can give rise to EWSR1-FLI1high cells and vice versa (Figure 2f).
As LOX is a known downstream repressed target of EWSR1-FLI123 (Supplementary Figure S4) that can be easily detected by immunohistochemistry, we used it as a surrogate marker of EWSR1-FLI1 activity to screen Ewing cell lines and primary tumours. As expected, in A673 and SK-N-MC cell lines, most cells do not express the LOX protein, and only very few cells (<0.1%) present intense LOX expression (Figure 3a). Interestingly, while LOX-negative cells form clusters of inter-adhesive cells, the LOX-positive cells are isolated and present an important area of spreading. A double staining of LOX and actin further showed that these LOX positive cells were characterized by the presence of strong and robust actin stress fibres and well-spread shape (Figures 1b and c), all features of EWSR1-FLI1low cells (Figure 3a). We next analysed LOX expression in shA673-1c-derived xenografts (described in Postel-Vinay et al.16 and Franzetti et al.24). When xenografted mice were treated with DOX for 10 days, tumours presented an intense LOX nuclear/peri-nuclear staining (Figure 3b) hence confirming in vivo the negative regulation of LOX expression by EWSR1-FLI1. In contrast, xenografts from untreated mice were globally negative for LOX staining, and only a very small number of LOX positive tumour cells was found. Collectively, these data confirmed the rarity of LOX positive cells in Ewing cell lines and xenograft models, and further suggested that these LOX-positive cells correspond to EWSR1-FLI1low cells. To further evaluate EWSR1-FLI1 heterogeneity in human tumours, we screened a large cohort of 294 Ewing sarcoma patient-derived samples on four series of tissue microarrays (TMAs) by immunohistochemistry. We detected LOX positive Ewing cells in 69 samples. Precise quantification on a subset of eight Ewing sarcoma samples revealed 0.97–1.6% of LOX positive cells (Figure 3c).
Downstream markers of EWSR1-FLI1 activity confirm the presence of EWSR1-FLI1low subpopulation. (a) Immunofluorescence of LOX (EWSR1-FLI1-downregulated target) and actin cytoskeleton in A673 and SK-N-MC cell lines. Images were acquired using upright widefield Apotome microscope (Zeiss, Marly-Le-Roi, France) equipped with a Coolsnap HQ2 camera through a x63 NA 1.4 oil-immersion objective lens and driven by Axiovision software (Zeiss). White arrows point the cells of interest, exhibiting LOX expression associated to an important cell-spreading and a well-organized actin stress fibre cytoskeleton. Scale bars represent 20 μm. (b) IHC of LOX in shA673-1c+/−DOX xenografts. Black arrows indicate the cells of interest. Right panels represent enlargement of left panels. Scale bars represent 50 μm. (c) IHC and quantification of LOX in eight representative samples from Ewing tumours TMA. Black arrows point cells expressing LOX. Scale bars represent 20 μm. Error bars mean s.d.
Altogether these experiments indicate that the presence and activity of EWSR1-FLI1 can quantitatively and qualitatively vary with time within Ewing cells. The assessment of EWSR1-FLI1 expression and activity also documents the spontaneous presence of small fraction of EWSR1-FLI1low cells in Ewing sarcoma cell lines, xenografts and tumours.
The EWSR1-FLI1low→high transition increases lung metastasis in vivo.
Above described experiments indicate that Ewing cell populations are characterized in vitro and in vivo by cell-to-cell heterogeneity of EWSR1-FLI1 expression. The observation that EWSR1-FLI1low cells have increased migration and invasion capacities, suggests they may have a critical role in the metastasis spread. Interestingly, Chaturvedi and colleagues11 have shown that EWSR1-FLI1-knocked-down Ewing cells have increased propensity for lung seeding in vivo. But the ability of these cells to form metastases was not tested. We therefore further evaluated the ability of EWSR1-FLI1low cells to develop in metastasis upon recovery of full EWSR1-FLI1 expression. We took advantage of the reversibility of the knock-down of EWSR1-FLI1 in shA673-1c cells upon DOX withdrawal (Supplementary Figure S2). Indeed, when DOX is removed from the medium of shA673-1c cells pre-treated by DOX during 10 days, full re-expression of EWSR1-FLI1 and recovery of proliferation potential were observed. To study the metastatic potential of cells with the EWSR1-FLI1low→high transition in vivo, shA673-1c cells stably expressing green fluorescent protein (GFP) were DOX-pretreated in vitro for 10 days to achieve a low EWSR1-FLI1 expression and then injected into mouse tail vein (EWSR1-FLI1low→high group; Figure 4a). To allow the re-expression of EWSR1-FLI1, no DOX was given to the drinking water of mice. As a control, the same experiment was conducted using DOX-untreated GFP expressing shA673-1c cells (EWSR1-FLI1high group). After 4 weeks, mice were sacrificed and lungs were collected then screened for GFP-positive nodules (Figure 4b). CD99 staining was also used as a positive surrogate marker of EWSR1-FLI1 expression.24 GFP/CD99/EWSR1-FLI1 cells were detected in 7/8 mice injected with EWSR1-FLI1low cells but in only 3/8 mice injected with EWSR1-FLI1high cells (Figure 4c). Moreover, the number and the size of nodules (Figure 4d) were very significantly higher in the EWSR1-FLI1low→high group as compared to the EWSR1-FLI1high group. Indeed, we counted 48 nodules (41 tumours >2000 μm2; 7 tumours <2000 μm2), against only six moderate-to-important nodules for the EWSR1-FLI1high group.
The EWSR1-FLI1low→high transition increases seeding in lung and colonization in vivo. (a) DOX-pre-treated (EWSR1-FLI1 low→high group) or untreated (EWSR1-FLI1high group) shA673-1c/GFP cells were injected in the mouse tail vein. During the experiments, no DOX was given to mice to enable EWSR1-FLI1 re-expression in the DOX-pre-treated group. (b) HES, GFP, CD99 and EWSR1-FLI1 staining on slides of lung from mice injected with shA673-1c/GFP-DOX-pre-treated clone and sacrificed after 4 weeks. Lung cryosections of 6 μm were prepared at different depth with a cryostat at −20 °C (Microm HM550). Ewing cells were revealed by positive GFP and CD99 immunostaining as described in Franzetti et al.24 Section was scanned with Ultra-Fast Scanner (UFS, Philips). Scale bars represent 50 μm. (c) Percentage of mice positive for GFP/CD99 lung nodules in EWSR1-FLI1high group or EWSR1-FLI1low→high group, compared using a Fisher’s exact test (*P<0.05, ***P<0.001). (d) Number of GFP/CD99 lung nodules by group with lung nodules size distribution, measured with Image Management System (Philips Digital Pathology) and compared using a Mann–Whitney test.
Collectively, these data indicate that a low EWSR1-FLI1 expression strongly increases cell seeding in the lung that may subsequently evolve into a metastasis upon EWSR1-FLI1 re-expression.
Discussion
In this study, differential quantitative mass spectrometry analyses based on 2D-DIGE and SILAC proteomic approaches reveal a strong influence of EWSR1-FLI1 on the expression of proteins involved in cell cytoskeleton structure and cell adhesion. Indeed, the knockdown of EWSR1-FLI1 is associated with the increase of actin-binding proteins implied in cell contractility (MYL6, MYL12A, MYLPF), cytoskeleton assembly and maintenance (ACTN4, CFL1, GSN, MSN, PFN2, RDX, VCL), but also the decrease of cell–cell adhesion proteins as tight junctions (CLD1, OCL) and desmosome (DSP, PKP1) family proteins. At the opposite, we observed the increase of integrins (ITGA1, A4, B1, B5) proteins, crucial component of cell–matrix interactions.
These modifications are associated to major phenotypic changes. Indeed, the invalidation of EWSR1-FLI1 expression (EWSR1-FLI1low cells) induced increased three-dimensional migration and invasion properties, as demonstrated here by spheroids culture embedded in three-dimensional collagen-matrix and zebrafish xenotransplantation models. Moreover, tail vein injections of EWSR1-FLI1low cells show that these cells can seed in the lung and then grow into metastases when allowed to re-express EWSR1-FLI1. This process is much less efficient when it is conducted with EWSR1-FLI1high cells. Our observation does not rule out previous hypotheses on the possible role of EWSR1-FLI1 to loosen cell–cell adhesion and facilitate detachment of Ewing cells from the main tumour mass to migrate into the blood stream.11 Obviously, the experimental procedure that is based on injection of cells into the tail vein omits this initial ‘detachment’ step of the metastatic process. Hence, passive detachment of EWSR1-FLI1high and active migration of EWSR1-FLI1low cells may concurrently lead to the presence of Ewing cells into the blood stream. Yet, our results indicate that exit from the blood stream and attachment in distant sites may be strongly facilitated in the context of EWSR1-FLI1low cells. Our experiments also show that EWSR1-FLI1low cells that can seed in the lung also maintain a potential for growth upon EWSR1-FLI1 re-induction. In vitro and in vivo zebrafish experiments clearly document the inverse relationship between migration and growth in Ewing sarcoma cells. While EWSR1-FLI1low cells migrate but have very slow proliferation rate, EWSR1-FLI1high cells proliferate but have a much slower capacity to migrate (as illustrated in Figure 5).
Model of Ewing cells dissemination based on cell-to-cell heterogeneity of EWSR1-FLI1 expression. In this model, undifferentiated EWSR1-FLI1high cells proliferate with robust cell–cell adhesion, whereas mesenchymal-like EWSR1-FLI1low cells migrate and invade environment through important cell–matrix interactions.
These experimental data led us to investigate the heterogeneity of EWSR1-FLI1 expression in Ewing cell lines and tumours. Single cell RT-ddPCR and immunofluorescence experiments in different Ewing cell lines clearly show that the levels of expression of the EWSR1-FLI1 transcript and protein are heterogeneous from one cell to the other. Interestingly, Ewing cell lines derived from metastasis (SK-N-MC25 and TC7126) displayed more EWSR1-FLI1low cells as compared to the A673 cell line derived from a local tumour.27 Moreover, the TC71 mouse xenografts are associated with metastases,28 to our knowledge no A673 metastases have ever been published nor have been observed in our laboratory.
To further investigate this heterogeneity, we reasoned that expression levels of well-documented EWSR1-FLI1 targets may constitute instrumental witnesses of EWSR1-FLI1 activity in each cell. We choose to use EWSR1-FLI1-repressed genes as their expression in EWSR1-FLI1low cells lead to very clear mRNA or protein signals. The strong positive co-staining of LOX with well-organized actin stress fibres, as well as the co-expression of ICAM1, LOX and IGFBP3, all markers that are inhibited by EWSR1-FLI1, document the low activity of this oncoprotein in a subset of cells. This heterogeneity of EWSR1-FLI1 activity is also highly suspected in tumours as a small subset of LOX-positive cells can be detected in a significant proportion of cases. Interestingly, LOX, which is used here as a surrogate of EWSR1-FLI1 expression, may also have an active role in the metastatic process. Indeed, LOX is described as a protein implicated in migration, cellular invasion and the metastasis.29 Recently, it was demonstrated in a subtype of breast tumours that LOX secretion by the cancer cells induces the formation of osteolytic lesions and may drive the formation of a pre-metastatic niche in bones facilitating the metastasis formation.30 It may be tempting to speculate that EWSR1-FLI1low cells expressing LOX could facilitate the degradation and invasion of bone, the usual primary site of Ewing sarcoma, and facilitate the formation of pre-metastatic lesions in lung. In this respect, it will be extremely interesting to assess whether the number of LOX-positive cells correlate with the presence of metastases once clinical data from the patients will be released.
An important issue is how much the EWSR1-FLI1low and EWSR1-FLI1high states are stable or can fluctuate from one to the other. Indeed, the slow proliferation of EWSR1-FLI1low cells constitutes a competitive disadvantage with respect to the high proliferation rate of EWSR1-FLI1high cells. The most likely hypothesis to account for the coexistence of these two states is that the expression level of EWSR1-FLI1 may dynamically fluctuate along time in the same cell. The isolation and regrowth of EWSR1-FLI1low and EWSR1-FLI1high cell populations through ICAM1 heterogeneity expression enable to observe in both cases a spontaneous regeneration of the initial heterogeneity of the cell population. Such dynamic and reversible switch between two different proliferative states has been clearly documented in human melanoma cells.31, 32 Dynamic variation of the expression of the JARID1B gene was shown to be responsible for the coexistence of a minor JARID1Bhigh low doubling time component and a main highly proliferative JARID1Blow compartment, both cell populations being essential to maintain tumour growth.32 Neuroblastoma, a pediatric tumour, also presents a reversible phenotypic heterogeneity from an adherent phenotype to a highly migratory and invasive cell type under stress exposure.33, 34
Another important question is the mechanisms of EWSR1-FLI1 fluctuation. Stochastic, cell-autonomous or non-cell-autonomous mechanisms may be involved. While the EWSR1-FLI1 variations spontaneously observed in cell lines, in the absence of any stromal cells, support a role for the two first mechanisms, the impact of the microenvironment may also be crucial in vivo. Indeed, tumour cell plasticity has been shown to enable tumour cells to adapt to their environment by reversible adaptive phenotypic transformation and/or turning on and off specific markers.35 In Ewing sarcoma, few studies have yet investigated the influence of micro-environmental factors on the expression of EWSR1-FLI1 transcript or protein. However, starvation of basic fibroblast growth factor by a neutralizing antibody induced a downregulation of EWSR1-FLI1 protein expression.36 Moreover, hypoxia transiently modulates the expression of EWSR1-FLI1 and the malignant properties of Ewing cells.37 Thus, according to cell localization and stress exposure in tumour, the EWSR1-FLI1 expression could be differentially modulated and consequently induce a phenotypic transition from proliferation to invasiveness.
In many tumour types the metastatic spread is suggested to be driven or at least favoured by the accumulation of genetic alterations that progressively provide the cancer cells the functional ability to escape from the primary mass and seed in distant sites. In Ewing sarcoma, different groups have recently described the paucity of secondary genetic alterations, in addition to the primary EWSR1-FLI1 genetic lesion.38, 39, 40 Though STAG2 and TP53 mutations are associated with a more aggressive disease, their low frequency cannot account for the much more frequent occurrence of metastases.41 The data presented here indicate that cell-to-cell variation of the expression level of EWSR1-FLI1 as a consequence of cell autonomous or non-autonomous mechanisms may constitute a critical component of the metastatic process. Importantly, this observation may be of high clinical importance as current therapies are directed against highly proliferative cells and as novel therapies that target EWSR1-FLI1 are currently being developed.42, 43, 44 These therapies are expected to be poorly efficient against the EWSR1-FLI1low cell population, which may suggest to associate to such treatments inhibitors of the migration and invasive processes.
Methods
Three-dimensional collagen I multicellular spheroid invasion assays
Multicellular spheroids of Ewing cells were prepared by the hanging droplets method45, 46 using 4 × 103 cells in 20-μl droplets of culture medium with 10% FBS. After 3 days of natural aggregation by gravity, spheroids were collected and embedded in 60-μl collagen matrix onto glass-bottom dishes.
Zebrafish embryo xenotransplantation
Around 500 shA673-1c/mCherry cells were injected into the yolk as described.18, 47 Larvae with regular developmental phenotypes were maintained at 34 °C in 60 μg/ml Instant Ocean eggwater, supplemented if necessary with 20 μg/ml DOX. After 4 days, embryos were fixed overnight in 4% paraformaldehyde at 4 °C. Embryos were imaged in glass-bottom 96-well plates using a NIKON3 confocal microscope. Image processing was performed with ImageJ 1.43 (NIH). ImagePro Analyzer 7.0 (Media Cybernetics, Rockville, MD, USA) analysis was performed as described previously.48 Zebrafish were handled compliant to local animal welfare regulations and maintained according to standard protocols (www.ZFIN.org).
Single cell reverse transcriptase droplet digital PCR (RT-ddPCR)
Cells were FACS-sorted (MoFlo Astrios, Beckman Coulter) and dispensed individually in 96-well plates (MicroAmp Optical 96-well plate, Lifetechnologies, Carlsbad, CA, USA) containing 5 μl Single Cell Lysis Buffer and 0.5 μl Single Cell DNaseI (from Single Cell lysis kit, Ambion, Carlsbad, CA, USA). Then, 2.5 μl of RT reaction mix (iScript Advanced cDNA Synthesis kit for RT-qPCR, Bio-Rad, Marnes-La-Coquette, France) was added to the single cell lysate. RT reaction was performed according to manufacturer instructions. Droplet digital PCR amplification of EWSR1-FLI1 (FAM) and RPLP0 (VIC) was performed in 25 μl reactions by adding the ddPCR Supermix for Probes No UDP (Bio-Rad), the FAM probes and the VIC probes to the cDNA. Droplets were generated using the QX100 Droplet Generator (Bio-Rad) with 50 μl Droplet Generation Oil (Bio-Rad) and were thermocycled under the following conditions: 95 °C for 10 min, then 40 cycles of 94 °C for 30 s, 59 °C for 60 s, and final enzyme deactivation at 98 °C for 10 min. The individual droplets were analysed using the QX100 Droplet Reader and Quantasoft software.
Immunohistochemistry on Ewing sarcoma tissue microarrays (TMA)
Four different series of Ewing tumour tissue microarray were used in this study: TMA1 containing 20 samples from CHR de Marseille Hopital de la Timone (C Bouvier), TMA2 containing 67 samples from CHR de Tours (G De Pinieux), TMA3 containing 23 samples from CHR de Lille (S Aubert), TMA4 containing 184 samples from University of Münster (Uta Dirksen/PROVABES). Paraffin-embedded sections of each TMA were stained with rabbit LOX antibody (PTGlab, #17958-1-AP, dil 1/100). Precise quantification of tumour sample subset was performed using Immunoratio ImageJ plugin.
Tail vein metastasis assay
Two million shA673-1c/GFP cells treated with 1 μg/ml of DOX during 10 days (EWSR1-FLI1low→high group) or untreated (EWSR1-FLI1high group) were injected into the tail vein of eight female Severe Combined Immune Deficiency mice for each group, 7-weeks old. A piece of each lung was fixed by immersion in 4% paraformaldehyde during 24 h, washed in 1X PBS and steeped into a solution of 20% sucrose until full infiltration, then embedded into optimal cutting temperature compound (Tissue-Tek, Sakura, Villeneuve d'Ascq, France) and cryofreezed. All experimental procedures and animal housing were performed in strict accordance with the recommendations of the European and National Regulations for the Protection of Vertebrate Animals used for Experimental and other Scientific Purposes (facility license #C75-05-18) for care and use of laboratory animals. All animal experiments were carried out under the supervision of an investigator authorized by the French Competent Veterinary Authority.
Ewing cell lines, EWSR1-FLI1 transient and inducible inhibition by shRNA
A673, SK-N-MC, EW7 and TC71 are patient-derived Ewing sarcoma cell lines. The shA673-1c clone was generated as previously described.14 Cell authentication was performed by comparison with STR profiling from ATCC (A673, SK-N-MC), DSMZ (TC71) and COSMIC (EW7). Cells were checked routinely by PCR for the absence of mycoplasma. The shSK-E17T clone was generated with SK-N-MC cell line as previously described for shA673-1c clone, but the H1 promoter-tet-O-shEF-1 cassette was cloned into pTRIP/GPF lentiviral expression vector. The transient inhibition of EWSR1-FLI1 was performed with lentiviral infection of pLKO lentiviral expression vectors carrying shRNA targeting the 3′UTR of FLI1.
The shA673-1c/mCherry and shA673-1c/GFP clones were constructed by lentiviral infection of the pCDH1-EF1-mcherry (a kind gift from Philippe Benaroch, U932, Institut Curie, Paris, France) and the pCDH1-EF1-copGFP plasmid (System Biosciences, Palo Alto, CA, USA) in shA673-1c clone.
Production of lentiviral particles
Packaging, production and titration of viral particles were performed as described in Lee et al.49
ShRNA, primers and antibodies
ShRNA for EWSR1-FLI1 inhibition, and primers, antibodies for RT-qPCR, western blot, IF and IHC experiments are detailed in Supplementary Table S3.
References
Ross KA, Smyth NA, Murawski CD, Kennedy JG . The biology of ewing sarcoma. ISRN Oncol 2013; 2013: 759725.
HaDuong JH, Martin AA, Skapek SX, Mascarenhas L . Sarcomas. Pediatr Clin North Am 2015; 62: 179–200.
Deley M-CL, Paulussen M, Lewis I, Brennan B, Ranft A, Whelan J et al. Cyclophosphamide compared with ifosfamide in consolidation treatment of standard-risk Ewing sarcoma: results of the randomized noninferiority euro-EWING99-R1 trial. J Clin Oncol 2014; 32: 2440–2448.
Kovar H, Alonso J, Aman P, Aryee DNT, Ban J, Burchill SA et al. The first European interdisciplinary Ewing sarcoma research summit. Front Oncol 2012; 2: 54.
Delattre O, Zucman J, Plougastel B, Desmaze C, Melot T, Peter M et al. Gene fusion with an ETS DNA-binding domain caused by chromosome translocation in human tumours. Nature 1992; 359: 162–165.
Lessnick SL, Ladanyi M . Molecular pathogenesis of Ewing sarcoma: new therapeutic and transcriptional targets. Annu Rev Pathol Mech Dis 2012; 7: 145–159.
May WA, Gishizky ML, Lessnick SL, Lunsford LB, Lewis BC, Delattre O et al. Ewing sarcoma 11;22 translocation produces a chimeric transcription factor that requires the DNA-binding domain encoded by FLI1 for transformation. Proc Natl Acad Sci USA 1993; 90: 5752–5756.
Braunreiter CL, Hancock JD, Coffin CM, Boucher KM, Lessnick SL . Expression of EWS-ETS fusions in NIH3T3 cells reveals significant differences to Ewing’s sarcoma. Cell Cycle 2006; 5: 2753–2759.
Toomey EC, Schiffman JD, Lessnick SL . Recent advances in the molecular pathogenesis of Ewing’s sarcoma. Oncogene 2010; 29: 4504–4516.
Krook MA, Nicholls LA, Scannell CA, Chugh R, Thomas DG, Lawlor ER . Stress-induced CXCR4 promotes migration and invasion of Ewing sarcoma. Mol Cancer Res MCR 2014; 12: 953–964.
Chaturvedi A, Hoffman LM, Welm AL, Lessnick SL, Beckerle MC . The EWS/FLI oncogene drives changes in cellular morphology, adhesion, and migration in Ewing sarcoma. Genes Cancer 2012; 3: 102–116.
Wiles ET, Bell R, Thomas D, Beckerle M, Lessnick SL . ZEB2 represses the epithelial phenotype and facilitates metastasis in Ewing sarcoma. Genes Cancer 2013; 4: 486–500.
Chaturvedi A, Hoffman LM, Jensen CC, Lin Y-C, Grossmann AH, Randall RL et al. Molecular dissection of the mechanism by which EWS/FLI expression compromises actin cytoskeletal integrity and cell adhesion in Ewing sarcoma. Mol Biol Cell 2014; 25: 2695–2709.
Tirode F, Laud-Duval K, Prieur A, Delorme B, Charbord P, Delattre O . Mesenchymal stem cell features of Ewing tumors. Cancer Cell 2007; 11: 421–429.
Stoll G, Surdez D, Tirode F, Laud K, Barillot E, Zinovyev A et al. Systems biology of Ewing sarcoma: a network model of EWS-FLI1 effect on proliferation and apoptosis. Nucleic Acids Res 2013; 41: 8853–8871.
Postel-Vinay S, Véron AS, Tirode F, Pierron G, Reynaud S, Kovar H et al. Common variants near TARDBP and EGR2 are associated with susceptibility to Ewing sarcoma. Nat Genet 2012; 44: 323–327.
Amsellem V, Kryszke M-H, Hervy M, Subra F, Athman R, Leh H et al. The actin cytoskeleton-associated protein zyxin acts as a tumor suppressor in Ewing tumor cells. Exp Cell Res 2005; 304: 443–456.
van der Ent W, Jochemsen AG, Teunisse AFAS, Krens SFG, Szuhai K, Spaink HP et al. Ewing sarcoma inhibition by disruption of EWSR1-FLI1 transcriptional activity and reactivation of p53. J Pathol 2014; 233: 415–424.
Noursadeghi M, Tsang J, Haustein T, Miller RF, Chain BM, Katz DR . Quantitative imaging assay for NF-kappaB nuclear translocation in primary human macrophages. J Immunol Methods 2008; 329: 194–200.
Barbáchano A, Ordóñez-Morán P, García JM, Sánchez A, Pereira F, Larriba MJ et al. SPROUTY-2 and E-cadherin regulate reciprocally and dictate colon cancer cell tumourigenicity. Oncogene 2010; 29: 4800–4813.
Prieur A, Tirode F, Cohen P, Delattre O . EWS/FLI-1 silencing and gene profiling of Ewing cells reveal downstream oncogenic pathways and a crucial role for repression of insulin-like growth factor binding protein 3. Mol Cell Biol 2004; 24: 7275–7283.
Agra N, Cidre F, García-García L, de la Parra J, Alonso J . Lysyl oxidase is downregulated by the EWS/FLI1 oncoprotein and its propeptide domain displays tumor supressor activities in Ewing sarcoma cells. PLoS ONE 2013, juin8: e66281.
Sankar S, Bell R, Stephens B, Zhuo R, Sharma S, Bearss DJ et al. Mechanism and relevance of EWS/FLI-mediated transcriptional repression in Ewing sarcoma. Oncogene 2013; 32: 5089–5100.
Franzetti G-A, Laud-Duval K, Bellanger D, Stern M-H, Sastre-Garau X, Delattre O . MiR-30a-5p connects EWS-FLI1 and CD99, two major therapeutic targets in Ewing tumor. Oncogene 2013; 32: 3915–3921.
Biedler JL, Helson L, Spengler BA . Morphology and growth, tumorigenicity, and cytogenetics of human neuroblastoma cells in continuous culture. Cancer Res 1973; 33: 2643–2652.
Whang-Peng J, Triche TJ, Knutsen T, Miser J, Douglass EC, Israel MA . Chromosome translocation in peripheral neuroepithelioma. N Engl J Med 1984; 311: 584–585.
Giard DJ, Aaronson SA, Todaro GJ, Arnstein P, Kersey JH, Dosik H et al. In vitro cultivation of human tumors: establishment of cell lines derived from a series of solid tumors. J Natl Cancer Inst 1973; 51: 1417–1423.
El-Naggar AM, Veinotte CJ, Cheng H, Grunewald TGP, Negri GL, Somasekharan SP et al. Translational activation of HIF1α by YB-1 promotes sarcoma metastasis. Cancer Cell 2015; 27: 682–697.
Erler JT, Bennewith KL, Nicolau M, Dornhöfer N, Kong C, Le Q-T et al. Lysyl oxidase is essential for hypoxia-induced metastasis. Nature 2006; 440: 1222–1226.
Cox TR, Rumney RMH, Schoof EM, Perryman L, Høye AM, Agrawal A et al. The hypoxic cancer secretome induces pre-metastatic bone lesions through lysyl oxidase. Nature 2015; 552: 106–110.
Hoek KS, Eichhoff OM, Schlegel NC, Döbbeling U, Kobert N, Schaerer L et al. In vivo switching of human melanoma cells between proliferative and invasive states. Cancer Res 2008; 68: 650–656.
Roesch A, Fukunaga-Kalabis M, Schmidt EC, Zabierowski SE, Brafford PA, Vultur A et al. A temporarily distinct subpopulation of slow-cycling melanoma cells is required for continuous tumor growth. Cell 2010; 141: 583–594.
Poomthavorn P, Wong SHX, Higgins S, Werther GA, Russo VC . Activation of a prometastatic gene expression program in hypoxic neuroblastoma cells. Endocr Relat Cancer 2009; 16: 991–1004.
Chakrabarti L, Abou-Antoun T, Vukmanovic S, Sandler AD . Reversible adaptive plasticity: a mechanism for neuroblastoma cell heterogeneity and chemo-resistance. Front Oncol 2012; 2: 82.
Meacham CE, Morrison SJ . Tumour heterogeneity and cancer cell plasticity. Nature 2013; 501: 328–337.
Girnita L, Girnita A, Wang M, Meis-Kindblom JM, Kindblom LG, Larsson O . A link between basic fibroblast growth factor (bFGF) and EWS/FLI-1 in Ewing’s sarcoma cells. Oncogene 2000; 19: 4298–4301.
Aryee DNT, Niedan S, Kauer M, Schwentner R, Bennani-Baiti IM, Ban J et al. Hypoxia modulates EWS-FLI1 transcriptional signature and enhances the malignant properties of Ewing’s sarcoma cells in vitro. Cancer Res 2010; 70: 4015–4023.
Brohl AS, Solomon DA, Chang W, Wang J, Song Y, Sindiri S et al. The genomic landscape of the Ewing sarcoma family of tumors reveals recurrent STAG2 mutation. PLoS Genet 2014; 10: e1004475.
Crompton BD, Stewart C, Taylor-Weiner A, Alexe G, Kurek KC, Calicchio ML et al. The genomic landscape of pediatric Ewing sarcoma. Cancer Discov 2014; 4: 1326–1341.
Tirode F, Surdez D, Ma X, Parker M, Le Deley MC, Bahrami A et al. Genomic landscape of Ewing sarcoma defines an aggressive subtype with co-association of STAG2 and TP53 mutations. Cancer Discov 2014; 4: 1342–1353.
Dahlin DC, Coventry MB, Scanlon PW . Ewing’s sarcoma. J Bone Jt Surg 1961; 43: 185–192.
Stegmaier K, Wong JS, Ross KN, Chow KT, Peck D, Wright RD et al. Signature-based small molecule screening identifies cytosine arabinoside as an EWS/FLI modulator in Ewing sarcoma. PLoS Med 2007; 4: e122.
Grohar PJ, Woldemichael GM, Griffin LB, Mendoza A, Chen Q-R, Yeung C et al. Identification of an inhibitor of the EWS-FLI1 oncogenic transcription factor by high-throughput screening. J Natl Cancer Inst 2011; 103: 962–978.
Sengupta A, Rahman M, Mateo-Lozano S, Tirado OM, Notario V . The dual inhibitory effect of thiostrepton on FoxM1 and EWS/FLI1 provides a novel therapeutic option for Ewing’s sarcoma. Int J Oncol 2013; 43: 803–812.
Kelm JM, Timmins NE, Brown CJ, Fussenegger M, Nielsen LK . Method for generation of homogeneous multicellular tumor spheroids applicable to a wide variety of cell types. Biotechnol Bioeng 2003; 83: 173–180.
Montagnac G, Meas-Yedid V, Irondelle M, Castro-Castro A, Franco M, Shida T et al. αTAT1 catalyses microtubule acetylation at clathrin-coated pits. Nature 2013; 502: 567–570.
Ban J, Aryee DNT, Fourtouna A, van der Ent W, Kauer M, Niedan S et al. Suppression of deacetylase SIRT1 mediates tumor-suppressive NOTCH response and offers a novel treatment option in metastatic Ewing sarcoma. Cancer Res 2014; 74: 6578–6588.
Ghotra VPS, He S, de Bont H, van der Ent W, Spaink HP, van de Water B et al. Automated whole animal bio-imaging assay for human cancer dissemination. PLoS ONE 2012; 7: e31281.
Lee M-TM, Coburn GA, McClure MO, Cullen BR . Inhibition of human immunodeficiency virus type 1 replication in primary macrophages by using Tat- or CCR5-specific small interfering RNAs expressed from a lentivirus vector. J Virol 2003; 77: 11964–11972.
Sibarita J . Deconvolution microscopy. Adv Biochem Eng-Biotechnol 2005; 95: 201–243.
Acknowledgements
This work was supported by grants from the Institut Curie, the INSERM, the Ligue Nationale Contre le Cancer (Equipe labellisée), the Institut National du Cancer (PLBIO14-237), the European PROVABES (ERA-649 NET TRANSCAN JTC-2011; 01KT1310), ASSET (FP7-HEALTH-2010-259348), and EEC (HEALTH-F2-2013-602856) projects, the German Cancer Aid (DKH 108128) and the following associations: the Société Française de lutte contre les Cancers et des leucémies de l’enfant et de l’adolescent, the Etoile de Martin, Courir pour Mathieu, Dans les pas du Géant, Olivier Chape, Les Bagouzamanon, Enfants et Santé, les Amis de Claire and M la vie avec Lisa. Georges-Alain Franzetti was supported by a fellowship from the Ministère de l’Enseignement Supérieur et de la Recherche (France) and from the FRM (FDT20130928190). Wietske van der Ent was supported by Stichting Kinderen Kankervrij (Grant No. 30677). The authors are grateful to Laura Brullé-Soumaré (INSERM U1143, CNRS UMR3666, Institut Curie, Paris, France) for her help with mice experiments. The authors would like to acknowledge the Nikon Imaging Center @Institut Curie-CNRS and the Cell and Tissue Imaging Platform (member of France-Bioimaging) of the Genetics and Developmental Biology Department (UMR3215/U934) of Institut Curie, and also the Cytometry Plateform of Tumor Biology Department of Institut Curie.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Franzetti, GA., Laud-Duval, K., van der Ent, W. et al. Cell-to-cell heterogeneity of EWSR1-FLI1 activity determines proliferation/migration choices in Ewing sarcoma cells. Oncogene 36, 3505–3514 (2017). https://doi.org/10.1038/onc.2016.498
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.498
This article is cited by
-
Targeting the EphA2 pathway: could it be the way for bone sarcomas?
Cell Communication and Signaling (2024)
-
Co-targeting JAK1/STAT6/GAS6/TAM signaling improves chemotherapy efficacy in Ewing sarcoma
Nature Communications (2024)
-
New dual inducible cellular model to investigate temporal control of oncogenic cooperating genes
Scientific Reports (2024)
-
Ganglioside SSEA-4 in Ewing sarcoma marks a tumor cell population with aggressive features and is a potential cell-surface immune target
Scientific Reports (2024)
-
DisP-seq reveals the genome-wide functional organization of DNA-associated disordered proteins
Nature Biotechnology (2024)