Abstract
The introduction of new therapies against particular genetic mutations in non-small-cell lung cancer is a promising avenue for improving patient survival, but the target population is small. There is a need to discover new potential actionable genetic lesions, to which end, non-conventional cancer pathways, such as RNA editing, are worth exploring. Herein we show that the adenosine-to-inosine editing enzyme ADAR1 undergoes gene amplification in non-small cancer cell lines and primary tumors in association with higher levels of the corresponding mRNA and protein. From a growth and invasion standpoint, the depletion of ADAR1 expression in amplified cells reduces their tumorigenic potential in cell culture and mouse models, whereas its overexpression has the opposite effects. From a functional perspective, ADAR1 overexpression enhances the editing frequencies of target transcripts such as NEIL1 and miR-381. In the clinical setting, patients with early-stage lung cancer, but harboring ADAR1 gene amplification, have poor outcomes. Overall, our results indicate a role for ADAR1 as a lung cancer oncogene undergoing gene amplification-associated activation that affects downstream RNA editing patterns and patient prognosis.
Similar content being viewed by others
Introduction
Non-small-cell lung cancer (NSCLC) is the primary cause of cancer-related death.1 The poor outcome of patients with NSCLC is associated with several factors, among which are late disease diagnosis and the small number of effective drugs. The absence of validated prognostic biomarkers may also be relevant, because even patients with stage I NSCLC who undergo potentially curative surgical resection have a 5-year relapse rate of 35–50%.1 Comprehensive genomic analyses have revealed a few genetically altered therapeutic targets in lung adenocarcinoma, such as EGFR mutations and ALK translocations,2 but many more drug-actionable genetic hits are necessary if the clinical course of this tumor type is to be significantly improved. In this regard, beyond the classical signaling pathways, less explored molecular and cellular networks can also contribute to lung tumorigenesis and provide promising targets for new anticancer compounds. One particular candidate is RNA editing.3
RNA editing is defined as chemical modifications in the RNA transcript after synthesis by RNA polymerases.3 These specific changes in RNA sequence can dramatically alter the amino acid composition and properties of the messenger RNA-derived protein. Recent studies indicate that many thousands of transcripts are affected by RNA editing, including mRNAs and non-coding RNAs.4, 5, 6, 7 Two families of proteins carry out editing by deamination: apolipoprotein B mRNA editing complexes, including the apolipoprotein B mRNA editing complex ortholog activation-induced cytidine deaminase), which convert cytosine to uracil (C-to-U), and the adenosine deaminases acting on RNA (ADARs), which convert adenosine to inosine (A-to-I).3 Analysis of the spectrum of mutations has implicated apolipoprotein B mRNA editing complexes-mediated editing in many cancers8 and the contribution of activation-induced cytidine deaminase to lymphomagenesis is well established.9 However, the role of ADARs in tumorigenesis has been less extensively investigated. There are three ADAR genes in humans: ADAR1, ADAR2 and ADAR3. The first two of these are competent at editing and ubiquitously expressed, whereas ADAR3 expression is restricted to the brain and no editing activity has been described.3 Direct involvement of ADAR1 A-to-I-mediated editing in cancer development has been described in hepatocellular carcinoma, where ADAR1 upregulation stabilizes antizyme inhibitor 1 (AZIN1) by increasing its editing frequency, leading to increased amounts of downstream oncogenic proteins.10 In addition to liver cancer,11 different expression levels of ADAR1 have been related to other malignancies such as melanoma,12 esophageal cancer13 and chronic myeloid leukemia.14
These findings prompted us to investigate the presence of ADAR1 and ADAR2 expression changes, and their possible gene amplification, in lung cancer cells and primary tumors. Extra copies of a candidate oncogene may give tumor cells a growth advantage as well being potentially useful as a biomarker for cancer diagnosis and prognosis, and ultimately for designing targeted therapies. Once we identified ADAR1 gene amplification-mediated overexpression, we also studied its consequences for in vitro and in vivo tumoral growth, for RNA editing levels of target transcripts and its potential value as a biomarker for clinical outcome in lung tumors.
Results and discussion
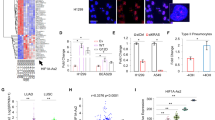
We first screened a collection of nine human NSCLC cell lines for the RNA expression levels of the two main ADARs, ADAR1 and ADAR2. These cell lines were A549, NCI-H1975, NCI-H1993, NCI-H1437, Cal12T, NCI-H1650, NCI-H1703, NCI-H441 and NCI-H1395. We also assessed ADAR1 gene expression in the human bronchial epithelial cells immortalized with telomerase and Cdk4-mediated p16 bypass (HBEC3KT) and a HBEC3KT tumorigenic-derived clone in which p53 has been knocked down and an oncogenic K-Ras mutation has been introduced (V12) (HBEC3KT-p53-K-Ras).15 The lung cancer cell lines were purchased from the American Type Culture Collection (Rockville, MD, USA) and were grown and maintained in 10% fetal bovine serum in Roswell Park Memorial Institute medium. Using quantitative reverse transcription-PCR we observed that ADAR2 expression did not show any significant differences within the described cell lines (Figure 1a and Supplementary Figure S1); however, a different pattern was found in ADAR1. Three NSCLC lines, NCI-H1395, NCI-H1993 and NCI-H1437, expressed significantly higher levels of the ADAR1 transcript than did all other cell lines (Figure 1a and Supplementary Figure S1). ADAR1 overexpression was further validated at the protein level: western blot analyses confirmed its enhanced expression in NCI-H1395, NCI-H1437 and NCI-H1993 cells relative to the other cell lines (Figure 1b). Immunofluorescence further confirmed the existence of ADAR1 overexpression in these three NSCLC cell lines (Figure 1c). As expected, the pattern of staining reflected the nuclear localization of the ADAR1 protein (Figure 1c), confirming that ADAR1 is in constant flux in and out of the nucleolus as it has been previously reported16 (Supplementary Figure S2).
Determination of ADAR1 RNA and protein overexpression in lung cancer cell lines and its association with gene amplification. (a) Assessment of ADAR1 and ADAR2 expression by quantitative reverse-transcription PCR. Determination of ADAR1 protein levels by western blot (AMAb90535; Atlas Antibody AB, Stockholm, Sweden) (b) and immunofluorescence (ab88574; Abcam, Cambridge, UK) (c) confirms overexpression of the ADAR1 protein in NCI-H1395, NCI-H1437 and NCI-H1993 cells. (d) Fluorescence in situ hybridization of the ADAR1 gene. The UCSC genome browser (http://www.genome.ucsc.edu) was used to select the bacterial artificial chromosome (BAC) clone spanning the 1q21.3 region of the ADAR1 gene: RP11-498A2 (1q21.3–1q22). A telomeric BAC clone located in the telomeric 1p36.23 region was used as a control. The BACs were obtained from the BACPAC Resource Center of the Children’s Hospital Oakland Research Institute (Oakland, CA, USA). ADAR1 and telomeric probes were labeled with Spectrum Green and Red dUTP (Abbott, Wiesbaden, Germany), respectively, using a CGH Nick Translation Reagent Kit (Abbott Molecular Inc., Des Plaines, IL, USA). The samples were counterstained with 4′,6-diamidino-2-phenylindole in Vectashield antifade solution (Burlingame, CA, USA). Gene amplification was observed in the interphases of NCI-H1395, NCI-H1437 and NCI-H1993 cells. Probes were verified to give a single signal on normal commercial lymphocyte metaphase slides (CGH Reagents; Abbott). (e) Assessment of ADAR1 copy number by quantitative genomic PCR. Amplification frequency of ADAR1 (evaluated with SYBR Green; Bio-Rad, Hercules, CA, USA) was calculated by the standard curve method using the 7900HT SDS program. To define an internal control gene, we chose chromosome 1p36.11 because it is the least aneuploid region among our cell lines (RPL11 gene). Primers are available upon request. DNA from the normal lung was used as the reference standard. Results are reported as n-fold copy number increase relative to the RPL11 gene. Gene amplification was observed in the NCI-H1395, NCI-H1437 and NCI-H1993 cell lines. (f) Multiplex ligation-dependent probe amplification (MLPA) assay. Two probemixes contain one probe for exons 4, 8 and 14 of the ADAR1 gene (in light blue). Twenty-one reference probes are included (in green). MLPA images from one of the two probemixes are shown. Values greater than 1 (equivalent to two copies) were considered to be extra copies. NCI-H1395, NCI-H1437 and NCI-H1993 show ADAR1 gene amplification, while A549, Cal12T and HBEC3KT-p53-K-ras are presented as examples of two ADAR1 copy number cells. (g) Graph depicting the ADAR1 amplified region in 1q21.3, identified with the Illumina Infinium HumanOmni5 microarray that interrogates 4301332 single-nucleotide polymorphisms in the entire genome. NCI-H1395 cell line has the largest amplified region (1502410 bp) that encompasses the ADAR1 gene. NCI-H1437 and NCI-H1993 have an amplified region of 767 275 and 759 573 bp, respectively. For the three cell lines there is a minimal common region of 662 085 bp where ADAR1 is included.
We next considered the possible existence of aberrant extra copies of the ADAR1 gene, a common mechanism of activation of oncogenes such as ERBB2 and MYC, that could lead to the described overexpression of the ADAR1 RNA transcript and protein. Adopting fluorescence in situ hybridization (Figure 1d) and a quantitative genomic PCR approach (Figure 1e), we observed ADAR1 gene amplification in the three ADAR1-overexpressing NSCLC lines mentioned above. This increase was particularly important in the cancer cell line H1395 (Figures 1d and e). The remaining NSCLC cell lines and the HBEC3KT/HBEC3KT-p53-K-Ras pair showed no apparent change in ADAR1 gene copy number (i.e., amplification, half gene dosage or homozygous deletion). Multiplex ligation-dependent probe amplification further confirmed the presence of ADAR1 gene amplification in NCI-H1395, NCI-H1437 and NCI-H1993 (Figure 1f). Multiplex ligation-dependent probe amplification also validated the finding of a normal ADAR1 gene copy number in A549, Cal12T and HBEC3KT/HBEC3KT-p53-K-Ras cells, which are used as illustrative examples (Figure 1f). Finally, using the Illumina Infinium HumanOmni5 microarray, which interrogates 4 301 332 single-nucleotide polymorphisms per sample, we determined the size of the amplicon containing ADAR1 in the three lung cancer cell lines studied here: NCI-H1993 (759 573 bp), NCI-H1437 (767 275 bp) and NCI-H1395 (1 502 410 bp). Genomically, ADAR1 (45 923 bp) was located in the middle of the smallest identified region of recurrent amplification (662 085 bp) (Figure 1g).
Once we had demonstrated the presence of ADAR1 gene amplification and its associated overexpression in the lung cancer cell lines, we examined its contribution to the tumorigenic phenotype in vitro and in vivo. We first analyzed the effect of ADAR1 depletion in lung cancer cells harboring its gene amplification and associated overexpression. Six ADAR1 shRNA-depleted clones were established for NCI-H1993 cells and two clones were established for NCI-H1395. Experiments for each clone were performed in triplicate. We observed that the reduction of ADAR1 expression in the gene-amplified cells had cancer growth-inhibitory features. Upon stable transfection of shRNAs against ADAR1 in the gene-amplified NCI-H1993 and NCI-H1395 lung cancer cell lines and efficient depletion of the ADAR1 protein (Figure 2a), the cells proved less viable in the 3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2H-tetrazolium bromide assay (Figure 2b) and had a markedly reduced percentage colony-formation density in the assay developed on plastic plates (Figure 2c). Transfection of the scramble shRNAdid not reduce cell viability (Figure 2b) and had no impact on the colony formation assay (Figure 2c). The shRNA-mediated depletion of ADAR1 in both cell lines using identical shRNA sequences that targeted ADAR1 was also associated with growth inhibition (Supplementary Figure S3). The used ADAR1 shRNA sequences did not target ADAR2 (Supplementary Figure S4). We next tested the ability of ADAR1 shRNA-depleted NCI-H1993 cells to form subcutaneous tumors in nude mice compared with scramble shRNA-transfected cells (Figure 2d). NCI-H1993 scramble shRNAcells formed tumors rapidly, but cells with shRNA-mediated depletion of ADAR1 had much lower tumorigenicity measured in terms of tumor volume and weight (Figure 2d). We then performed an orthotopic growth study, implanting equal-sized tumor pieces from the subcutaneous model in the lung. We observed that orthotopic ADAR1 shRNA-depleted tumors were significantly smaller and lighter than the scramble shRNA-derived tumors (Figure 2e). Most important, we also used a robust in vivo model of metastasis that it is not influenced by tumor burden: direct spleen injection. We observed that direct spleen injection of ADAR1 shRNA-depleted NCI-H1993 cells induced a lower level of formation of liver metastases than did scramble-shRNANCI-H1933 cells (Fisher's exact test, P<0.001) (Figure 2f).
Growth-promoting effects of ADAR1 in lung cancer. (a) Stable downregulation of the ADAR1 gene by short hairpins using two different target sequences for NCI-H1993 (clones sh1_4, sh1_5, sh1_29, sh4_24, sh4_25 and sh4_27) and NCI-H1395 (sh1_24 and sh2_6). ADAR1 shRNA sequences are available upon request. (b) The short hairpin ADAR1-depleted cells were less viable in the 3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2H-tetrazolium bromide (MTT) assay than in the untransfected or scrambled shRNA-transfected cells. Probabilities are those from permutation test. (c) The colony formation assay showed that NCI-H1993 and NCI-H1395 cells stably transfected with the shRNA against ADAR1 formed significantly fewer colonies than did scrambled shRNA-transfected cells. Probabilities are those from Student’s t-test. Results are presented as the mean±s.d., n=8. (d) Effect of ADAR1 shRNA-mediated depletion on the growth of NCI-H1993 xenografts in nude mice. Upper right panel: western-blot showing effective depletion of the ADAR1 protein upon shRNA targeting. Tumor volume was monitored over time and the tumor was excised and weighed at 49 days. There was a significant decrease in tumor weight and volume in the ADAR1 shRNA-stably transfected cells. Probabilities are those from Student’s t-test. Results are presented as the mean±s.e.m., n=20. (e) Left: illustrative tumor samples obtained at the end point of the orthotopic growth nude mouse experiment from shRNA scramble and shRNA-ADAR1-depleted NCI-H1993 cells. Right: mean±s.e.m. of tumor weights and volumes of orthotopic tumors for both groups at 60 days. Student’s t-test: **P<0.05. (f) Right: direct spleen injection of ADAR1 shRNA-depleted NCI-H1993 cells (n=24 mice) showed fewer liver metastases than with scramble-shRNAcells (n=7 mice). Left: illustrative images of macroscopic and microscopic metastases. (g) Top left: western blot showing ADAR1 protein overexpression upon transfection. Bottom left: effect of ADAR1 on the invasion potential of A549 cells determined by the matrigel invasion assay. Statistical significance was derived from ANOVAs. *P<0.05. Right: ADAR1 transfection in A549 also increased colony formation (top) and migration capacity in the wound-healing assay (below). (h) Right: direct spleen injection of ADAR1-transfected A549 cells (n=10 mice) showed increased liver metastases than with empty-vector transfected cells (n=5 mice). Left: illustrative images of macroscopic and microscopic metastases. (i) Top left: western blot showing ADAR1 protein overexpression upon transfection. Stable transfection of ADAR1 in HBEC3KT-p53-K-ras cells also caused increased colony formation (bottom left) and enhanced migration capability (right). Results are presented as the mean±s.d. from three independent experiments (Student’s t-test P<0.01 vs empty vector in all experiments). ***P<0.01.
We also studied the oncogenic potential of ADAR1 by evaluating its ability to enhance cell growth and invasion in the lung carcinoma cell line A549, which does not feature ADAR1 gene amplification. To this end, we transfected the plasmids driving the expression of the p110 isoform of ADAR1 and performed matrigel invasion (Figure 2g) and colony formation (Figure 2g) assays. We observed a significant increase in the number of invasive cells and colonies upon ADAR1 transfection compared with empty vector-transfected cells in the two described assays. Similar results were obtained from the wound-healing assay. We observed that stable transfection of ADAR1 in A549 cells caused a significant increase in the migration rate (Student’s t-test: P<0.01; Figure 2g). Importantly, we found that direct spleen injection of ADAR1-transfected A549 cells induced a higher number of liver metastases than empty-vector transfected cells (Fisher’s exact test, P<0.001; Figure 2h). Importantly, ADAR1 stable transfection in another unamplified cell line (HBEC3KT-p53-K-Ras) mimicked the effects observed in A549: the induced overexpression of ADAR1 increased colony formation and the migration rate (Figure 2i).
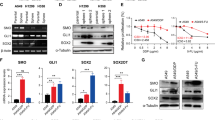
We also wondered about gene targets whose RNA editing could be altered in ADAR1-amplified lung cancer cell lines and that could further explain the aforementioned impact on cell growth and invasiveness. Whole-transcriptome sequence analysis in human tissues indicates thousands of possible A-to-I editing sites in nonrepeat sequences.4, 5, 6, 7 Among these involves the DNA repair enzyme NEIL1, a finding that was validated and studied in greater detail.17 Herein, we have studied the existence of possible different A-to-I editing levels in the identified site of the NEIL1 RNA (lysine 242 AAA codon) according to the ADAR1 gene dosage. We have found that the ADAR1 gene-amplified lung cancer cell lines NCI-H1395 and NCI-H1993, relative to the ADAR1 normal copy number cells A549 and HBEC3KT-p53-K-ras, show a higher frequency of A-to-I editing. NEIL1 AAA-to-AIA editing is observed in 92 and 82% of the sequenced transcripts in NCI-H1993 and NCI-H1395, respectively, whereas it is occurs in only 28 and 45% of the NEIL1 RNAs from A549 and HBEC3KT-p53-K-ras (Figure 3a). The editing of a second adenine (AAA-to-AII) for the lysine 242 codon of NEIL1 was also observed in our sequencing analyses, where it was also enriched in the lung cancer cell lines harboring ADAR1 gene amplification (Figure 3a). In addition, we further proved the role of ADAR1 in the editing levels of these NEIL1 RNA sites by external intervention: stable transfection of ADAR1 in A549 and HBEC3KT-p53-K-ras cells increased lysine 242 AAA codon editing (Figure 3b), whereas ADAR1 shRNA-mediated depletion in the amplified NCI-H1993 and NCI-H1395 cell lines reduced the levels of the NEIL1 transcripts containing the AIA and AII codons (Figure 3b). shRNA-mediated depletion of ADAR2 did not affect NEIL1 editing (Supplementary Figure S4).
Effect of ADAR1 on RNA-editing lung cancer cells. (a) Quantification of A-to-I editing at the lysine 242 AAA codon of the NEIL1 transcript determined by cDNA sequencing. Position 1, AIA; Position 2, AII. Left: ADAR1 gene-amplified cells (NCI-H1993 and NCI-H1395) show higher basal levels of NEIL1 editing than ADAR1 non-amplified cells (A549 and HBEC3KT-p53-K-ras). (b) Transfection-mediated overexpression of ADAR1 in A549 and HBEC3KT-p53-K-ras cells increases NEIL1 editing (left), whereas ADAR1-shRNA-mediated depletion in NCI-H1993 and NCI-H1395 cells reduces NEIL1 editing (right). P-values obtained by Fisher’s exact test. (c) Depiction of the minigene strategy to study the editing effect of ADAR1 in miR-381. The minigene is cloned in pLVX-shRNA2 vector and inserted into the genome. After its transcription, complementary sequences of the intron fall into a local stem loop that enhance ADAR1 binding and editing on the neighboring pri-miR-381 sequence. The two edited positions are highlighted in blue. (d) ADAR1-shRNA-mediated depletion in NCI-H1993 reduces miR-381 A-to-I editing.
We used a second strategy to verify ADAR1-candidate RNA targets by studying a minigene construct formed by the fusion of an RNA-editing inducer18 and miR-381, a target of editing in glioblastoma19 (Figure 3c). Lentiviral particles containing the described contruct were used to infect the ADAR1-amplified NCI-H1993 cell line. Upon ADAR1 shRNA-mediated depletion, we observed a significant decrease in the A-to-I editing of miR-381 in these infected cells (Figure 3d). Importantly, we observed that the A-to-I editing on the proposed targets contributed to the phenotype of lung cancer cells: we found that the overexpression of the edited NEIL1 and miR-381 transcripts significantly enhances the growth of A459 cells in comparison with the transfection of the unedited transcripts (permutation test, P<0.001) (Supplementary Figure S5). Overall, these findings suggest that, in the studied lung cancer cells, ADAR1 controls the A-to-I editing levels of critical target genes, having shown the example of NEIL1, a DNA repair gene involved in the removal of a wide array of modified bases20 and miR-381, a microRNA implicated in stemness, chemoresistance and other cancer-relevant pathways.21, 22 Importantly, numerous other transcripts are known to be edited and those in aggregate might need to be considered for a proper functional understanding of the role of ADAR1 gene amplification-associated overexpression in lung tumorigenesis.
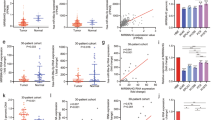
Finally, we sought to demonstrate that the presence of ADAR1 gene amplification was not a specific feature of in vitro-grown lung cancer cell lines and that it also occurred in primary tumors of lung cancer patients. In this regard, recent genomic data using single-nucleotide polymorphism microarrays confirm the gain of the 1q21 chromosomic region, where ADAR1 is located, in primary lung tumors.23 In the present study, we performed multiplex ligation-dependent probe amplification of the ADAR1 locus using a collection of 162 primary non-small-cell lung tumors comprising 147 lung adenocarcinomas and 15 squamous cell carcinomas. We identified ADAR1 gene amplification in 16 tumors comprising 10.2% (15 of 147) and 6.6% (1 of 15) adenocarcinoma and squamous lung cancer cases, respectively (Figure 4a). Most importantly, when we performed ADAR1 immunohistochemical staining for 12 primary lung cancer cases for which ADAR1 copy number had been determined by multiplex ligation-dependent probe amplification, we found that the presence of extra copies of ADAR1 was associated with overexpression of the ADAR1 protein (Figure 4b). Interestingly, in those lung cancer patients for whom we have clinical information and a long follow-up over different stages (n=128), the presence of the ADAR1 gene amplification (in 8.5%, 11 of 128, cases) was associated with a shorter relapse-free survival that was very nearly statistically significant (log-rank test: HR=2.29; 95% CI=0.96–5.48; P=0.054) (Figure 4c). The clinical stage (tumor, node, metastasis) of the lung cancer is known to be an independent prognostic factor, including in our cases (Cox multivariate regression: HR=2.22; 95% CI: 1.14–4.31; P=0.02) (Figure 4c). Therefore, we examined whether ADAR1 extra-dosage marked those patients with stage I tumors at high risk of recurrent disease, and found this to be the case. We observed that ADAR1 gene amplification, which was present in 8 of 90 (8.8%) stage I cases, was associated with shorter relapse-free survival (log-rank test: HR=3.31; 95% CI=1.24–8.83; P=0.011) (Figure 4d) and it was an independent prognostic factor in this set of patients (Figure 4d).
Detection of ADAR1 gene amplification, its associated overexpression and clinical impact in primary tumors from lung cancer patients. (a) Examples of assessment of ADAR1 copy number by MLPA in NSCLC patients with (top) and without (bottom) gene amplification. (b) Association between ADAR1 gene amplification and ADAR1 protein expression measured by immunohistochemistry (Ab126755; Abcam) in 12 studied cases. Fisher’s exact test, significant for two-tailed values of P=0.0182. Illustrative immunohistochemistry images for ADAR1 expression in two ADAR1 amplified (1 and 2) and unamplified (3 and 4) cases are shown. x100 magnification. (c) Left: Kaplan–Meier analysis of progression-free survival (PFS) among all clinical stages of NSCLC cases by ADAR1 genomic status. ADAR1 gene amplification is marginally associated with a shorter PFS. Number of events (progression) from 10 to 60 months in both groups. Right: forest plot of Cox multivariate regression, taking clinical features and ADAR1 copy number into account. Parameters with an associated value of P<0.05 were considered to be independent prognostic factors. (d) Left: Kaplan–Meier analysis of PFS in stage I NSCLC cases according to ADAR1 genomic status. ADAR1 gene amplification is significantly associated with a shorter PFS. Number of events (progression) from 10 to 60 months in both groups. Right: forest plot of Cox multivariate regression, taking clinical features of the validation cohort into account. Parameters with an associated value of P<0.05 were considered to be independent prognostic factors. ADAR1 gene amplification was associated with PFS in the NSCLC cohort.
Our results indicate that the RNA-editing enzyme ADAR1 undergoes gene amplification as part of the natural history of NSCLC. The copy-number gain for ADAR1 is associated with overexpression of the transcript and protein in lung cancer cell lines and primary tumors. Under these circumstances, ADAR1 exerts growth-enhancing activity in vitro and in vivo, as we have shown by depletion and transfection experiments in cell culture and mouse models. From a functional standpoint, ADAR1 mediates the A-to-I editing levels of coding (NEIL1) and non-coding (miR-381) RNA transcripts. Interestingly, a high ADAR1 copy number is associated with poor outcome in early-stage lung cancer patients, which could be useful in the clinical management of these cases. Overall, our findings imply a role of an altered RNA editing pathway in lung tumorigenesis.
References
Siegel R, Naishadham D, Jemal A . Cancer statistics 2012 CA Cancer J Clin 2012; 62: 10–29.
Buettner R, Wolf J, Thomas RK . Lessons learned from lung cancer genomics: the emerging concept of individualized diagnostics and treatment. J Clin Oncol 2013; 31: 1858–1865.
Avesson L, Barry G . The emerging role of RNA and DNA editing in cancer. Biochim Biophys Acta 2014; 1845: 308–316.
Li JB, Levanon EY, Yoon J-K, Aach J, Xie B, LeProust E et al. Genome-wide identification of human RNA editing sites by parallel DNA capturing and sequencing. Science 2009; 324: 1210–1213.
Peng Z, Cheng Y, Tan BC, Kang L, Tian Z, Zhu Y et al. Comprehensive analysis of RNA-Seq data reveals extensive RNA editing in a human transcriptome. Nat Biotechnol 2012; 30: 253–260.
Sakurai M, Ueda H, Yano T, Okada S, Terajima H, Mitsuyama T et al. A biochemical landscape of A-to-I RNA editing in the human brain transcriptome. Genome Res 2014; 24: 522–534.
Bazak L, Haviv A, Barak M, Jacob-Hirsch J, Deng P, Zhang R et al. A-to-I RNA editing occurs at over a hundred million genomic sites, located in a majority of human genes. Genome Res 2014; 24: 365–376.
Alexandrov LB, Stratton MR . Mutational signatures: the patterns of somatic mutations hidden in cancer genomes. Curr Opin Genet Dev 2014; 24: 52–60.
Rommel PC, Bosque D, Gitlin AD, Croft GF, Heintz N, Casellas R et al. Fate mapping for activation-induced cytidine deaminase (AID) marks non-lymphoid cells during mouse development. PLoS One 2013; 8: e69208.
Chen L, Li Y, Lin CH, Chan THM, Chow RKK, Song Y et al. Recoding RNA editing of AZIN1 predisposes to hepatocellular carcinoma. Nat Med 2013; 19: 209–216.
Chan THM, Lin CH, Qi L, Fei J, Li Y, Yong KJ et al. A disrupted RNA editing balance mediated by ADARs (Adenosine DeAminases that act on RNA) in human hepatocellular carcinoma. Gut 2014; 63: 832–843.
Shoshan E, Mobley AK, Braeuer RR, Kamiya T, Huang L, Vasquez ME et al. Reduced adenosine-to-inosine miR-455-5p editing promotes melanoma growth and metastasis. Nat Cell Biol 2015; 17: 311–321.
Qin Y-R, Qiao J-J, Chan THM, Zhu Y-H, Li F-F, Liu H et al. Adenosine-to-inosine RNA editing mediated by ADARs in esophageal squamous cell carcinoma. Cancer Res 2014; 74: 840–851.
Jiang Q, Crews LA, Barrett CL, Chun H-J, Isquith JM, Zipeto MA et al. ADAR1 promotes malignant progenitor reprogramming in chronic myeloid leukemia. Proc Natl Acad Sci USA 2013; 110: 1041–1046.
Sato M, Vaughan MB, Girard L, Peyton M, Lee W, Shames DS et al. Multiple oncogenic changes (K-RASV12, p53 knockdown, mutant EGFRs, p16 bypass, telomerase) are not sufficient to confer a full malignant phenotype on human bronchial epithelial cells. Cancer Res 2006; 66: 2116–2128.
Desterro JM, Keegan LP, Lafarga M, Berciano MT, O'Connell M, Carmo-Fonseca M . Dynamic association of RNA-editing enzymes with the nucleolus. J Cell Sci 2003; 116: 1805–1818.
Yeo J, Goodman RA, Schirle NT, David SS, Beal PA . RNA editing changes the lesion specificity for the DNA repair enzyme NEIL1. Proc Natl Acad Sci USA 2010; 107: 20715–20719.
Daniel C, Venø MT, Ekdahl Y, Kjems J, Öhman M . A distant cis acting intronic element induces site-selective RNA editing. Nucleic Acids Res 2012; 40: 9876–9886.
Skalsky RL, Cullen BR . Reduced expression of brain-enriched microRNAs in glioblastomas permits targeted regulation of a cell death gene. PLoS One 2011; 6: e24248.
Liu M, Doublié S, Wallace SS . Neil3, the final frontier for the DNA glycosylases that recognize oxidative damage. Mutat Res 2013; 743-744: 4–11.
Formosa A, Markert EK, Lena AM, Italiano D, Finazzi-Agro E, Levine AJ et al. MicroRNAs, miR-154, miR-299-5p, miR-376a, miR-376c, miR-377, miR-381, miR-487b, miR-485-3p, miR-495 and miR-654-3p, mapped to the 14q32. 31 locus, regulate proliferation, apoptosis, migration and invasion in metastatic prostate cancer cells. Oncogene 2014; 33: 5173–5182.
Wang Z, Yang J, Xu G, Wang W, Liu C, Yang H et al. Targeting miR-381-NEFL axis sensitizes glioblastoma cells to temozolomide by regulating stemness factors and multidrug resistance factors. Oncotarget 2015; 6: 3147–3164.
Wilkerson MD, Yin X, Walter V, Zhao N, Cabanski CR, Hayward MC et al. Differential pathogenesis of lung adenocarcinoma subtypes involving sequence mutations, copy number, chromosomal instability, and methylation. PLoS One 2012; 7: e36530.
Acknowledgements
This work was supported by the European Research Council under the European Community's Seventh Framework Programme (FP7/2007-2013)/ERC grant agreement no. 268626—EPINORC project, the Grant agreement number HEALTH-F2-2010-258677—CURELUNG project, the Spanish Ministry of Economy and Competitiveness (MINECO Projects no. SAF2011-22803, PI13-01339 and SAF2014-55000- R), the Institute of Health Carlos III (ISCIII)—PI10/02992, Ministerio de Educación, Ciencia e Innovación Grant SAF2010-14935, the Cellex Foundation, the National Cancer Center Research and Development Fund (NCC Biobank: 23 A-1) and the Health and Science Departments of the Catalan Government (Generalitat de Catalunya) AGAUR—project no. 2009SGR1315 and 2014SGR633. We thank the staff of the Animal Core Facility of IDIBELL for mouse care and maintenance. LRdP and ME are ICREA Research Professors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Anadón, C., Guil, S., Simó-Riudalbas, L. et al. Gene amplification-associated overexpression of the RNA editing enzyme ADAR1 enhances human lung tumorigenesis. Oncogene 35, 4407–4413 (2016). https://doi.org/10.1038/onc.2015.469
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2015.469
This article is cited by
-
RNA-binding proteins and exoribonucleases modulating miRNA in cancer: the enemy within
Experimental & Molecular Medicine (2024)
-
ADAR1 polymorphisms contribute to increased susceptibility in pediatric acute lymphoblastic leukemia
Annals of Hematology (2023)
-
The A-to-I editing of KPC1 promotes intrahepatic cholangiocarcinoma by attenuating proteasomal processing of NF-κB1 p105 to p50
Journal of Experimental & Clinical Cancer Research (2022)
-
The edited UPF1 is correlated with elevated asparagine synthetase in pancreatic ductal adenocarcinomas
Molecular Biology Reports (2022)
-
RNA editing in the forefront of epitranscriptomics and human health
Journal of Translational Medicine (2019)