Abstract
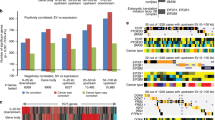
Chromosomal abnormalities are good guideposts when hunting for cancer-related genes. We analyzed copy number alterations of 163 primary gastric cancers using array-based comparative genomic hybridization and simultaneously performed a genome-wide integrated analysis of copy number and gene expression using microarray data for 58 tumors. We showed that chromosome 6p21 amplification frequently occurred secondary to ERBB2 amplification, was associated with poorer prognosis and caused overexpression of half of the genes mapped. A comprehensive small interfering RNA knockdown of 58 genes overexpressed in tumors identified 32 genes that reduced gastric cancer cell growth. Enforced expression of 16 of these genes promoted cell growth in vitro, and six genes showing more than two-fold activity conferred tumor-forming ability in vivo. Among these six candidates, GLO1, encoding a detoxifying enzyme glyoxalase I (GLO1), exhibited the strongest tumor-forming activity. Coexpression of other genes with GLO1 enhanced growth-stimulating activity. A GLO1 inhibitor, S-p-bromobenzyl glutathione cyclopentyl diester, inhibited the growth of two-thirds of 24 gastric cancer cell lines examined. The efficacy was found to be associated with the mRNA expression ratio of GLO1 to GLO2, encoding glyoxalase II (GLO2), another constituent of the glyoxalase system. GLO1 downregulation affected cell growth through inactivating central carbon metabolism and reduced the transcriptional activities of nuclear factor kappa B and activator protein-1. Our study demonstrates that GLO1 is a novel metabolic oncogene of the 6p21 amplicon, which promotes tumor growth and aberrant transcriptional signals via regulating cellular metabolic activities for energy production and could be a potential therapeutic target in gastric cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lochhead P, El-Omar EM . Gastric cancer. Br Med Bull 2008; 85: 87–100.
Smith MG, Hold GL, Tahara E, El-Omar EM . Cellular and molecular aspects of gastric cancer. World J Gastroenterol 2006; 12: 2979–2990.
Tamura G . Alterations of tumor suppressor and tumor-related genes in the development and progression of gastric cancer. World J Gastroenterol 2006; 12: 192–198.
Gorringe KL, Boussioutas A Melbourne Gastric Cancer Group, Peter Mac Microarray Facility, Bowtell DDL . Novel regions of chromosomal amplification at 6p21, 5p13, and 12q14 in gastric cancer identified by array comparative genomic hybridization. Genes Chromosomes Cancer 2005; 42: 247–259.
Takada H, Imoto I, Tsuda H, Nakanishi Y, Ichikura T, Mochizuki H et al. ADAM23, a possible tumor suppressor gene, is frequently silenced in gastric cancers by homozygous deletion or aberrant promoter hypermethylation. Oncogene 2005; 24: 8051–8060.
Vauhkonen H, Vauhkonen M, Sajantila A, Sipponen P, Knuutila S . DNA copy number aberrations in intestinal-type gastric cancer revealed by array-based comparative genomic hybridization. Cancer Genet Cytogenet 2006; 167: 150–154.
Yang S, Jeung H-C, Jeong HJ, Choi YH, Kim JE, Jung J-J et al. Identification of genes with correlated patterns of variations in DNA copy number and gene expression level in gastric cancer. Genomics 2007; 89: 451–459.
Myllykangas S, Junnila S, Kokkola A, Autio R, Scheinin I, Kiviluoto T et al. Integrated gene copy number and expression microarray analysis of gastric cancer highlights potential target genes. Int J Cancer 2008; 123: 817–825.
Tsukamoto Y, Uchida T, Kaman S, Noguchi T, Nguyen LT, Tanigawa M et al. Genome-wide analysis of DNA copy number alternations and gene expression in gastric cancer. J Pathol 2008; 216: 471–482.
Santos GC, Zielenska M, Prasad M, Squire JA . Chromosome 6p amplification and cancer progression. J Clin Pathol 2007; 60: 1–7.
Tanami H, Tsuda H, Okabe S, Iwai T, Sugihara K, Imoto I et al. Involvement of cyclin D3 in liver metastasis of colorectal cancer, revealed by genome-wide copy-number analysis. Lab Invest 2005; 85: 1118–1129.
Kasugai Y, Tagawa H, Kameoka Y, Morishima Y, Nakamura S, Seto M . Identification of CCND3 and BYSL as candidate targets for the 6p21 amplification in diffuse large B-cell lymphoma. Clin Cancer Res 2005; 11: 8265–8272.
Lu X-Y, Lu Y, Zhao Y-J, Jaeweon K, Kang J, Xiao-Nan L et al. Cell cycle regulator gene CDC5L, a potential target for 6p12-p21 amplicon in osteosarcoma. Mol Cancer Res 2008; 6: 937–946.
Chiang DY, Villanueva A, Hoshida Y, Peix J, Newell P, Minguez B et al. Focal gains of VEGFA and molecular classification of hepatocellular carcinoma. Cancer Res 2008; 68: 6779–6788.
Thornalley PJ . The glyoxalase system: new developments towards functional characterization of a metabolic pathway fundamental to biological life. Biochem J 1990; 269: 1–11.
Xue M, Rabbani N, Momiji H, Imbasi P, Anwar MM, Kitteringham N et al. Transcriptional control of glyoxalase 1 by Nrf2 provides a stress-responsive defence against dicarbonyl glycation. Biochem J 2012; 443: 213–222.
Hansen F, de Souza DF, Silveira Sda L, Hoefel AL, Fontoura JB, Tramontina AC et al. Methylglyoxal alters glucose metabolism and increases AGEs content in C6 glioma cells. Metab Brain Dis 2012; 27: 531–539.
Hirayama A, Kami K, Sugimoto M, Sugawara M, Toki N, Onozuka H et al. Quantitative metabolome profiling of colon and stomach cancer microenvironment by capillary electrophoresis time-of-flight mass spectrometry. Cancer Res 2009; 69: 4918–4925.
DeBerardinis RJ, Mancuso A, Daikhin E, Nissim I, Yudkoff M, Wehrli S et al. Beyond aerobic glycolysis: Transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc Natl Acad Sci USA 2007; 104: 19345–19350.
Ranganathan S, Walsh ES, Tew KD . Glyoxalase I in detoxification: studies using a glyoxalase I transfectant cell line. Biochem J 1995; 309: 127–131.
Sakamoto H, Mashima T, Sato S, Hashimoto Y, Yamori T, Tsuruo T . Selective activation of apoptosis program by S-p-bromobenzylglutathione cyclopentyl diester in glyoxalase I-overexpressing human lung cancer cells. Clin Cancer Res 2001; 7: 2513–2518.
Santarius T, Bignell GR, Greenman CD, Widaa S, Chen L, Mahoney CL et al. GLOI—a novel amplified gene in human cancer. Genes Chromosomes Cancer 2010; 49: 711–725.
de Hemptinne V, Rondas D, Toepoel M, Vancompernolle K . Phosphorylation on Thr-106 and NO-modification of glyoxalase I suppress the TNF-induced transcriptional activity of NF-κB. Mol Cell Biochem 2009; 325: 169–178.
Maeda S, Omata M . Inflammation and cancer: Role of nuclear factor-kappaB activation. Cancer Sci 2008; 99: 836–842.
Aggarwal BB, Gehlot P . Inflammation and cancer: How friendly is the relationship for cancer patients? Curr Opin Pharmacol 2009; 9: 351–369.
Kim DJ, Park K-S, Kim J-H, Yang S-H, Yoon JY, Han B-G et al. Helicobacter pylori proinflammatory protein up-regulates NF-κB as a cell-translocating Ser/Thr kinase. Proc Natl Acad Sci USA 2010; 107: 21418–21423.
Obama K, Kato T, Hasegawa S, Satoh S, Nakamura Y, Furukawa Y . Overexpression of peptidyl-prolyl isomerase-like 1 is associated with the growth of colon cancer cells. Clin Cancer Res 2006; 12: 70–76.
Chen J, Kobayashi M, Darmanin S, Qiao Y, Gully C, Zhao R et al. Hypoxia-mediated up-regulation of Pim-1 contributes to solid tumor formation. Am J Pathol 2009; 175: 400–411.
Kuiper RP, Schepens M, Thijssen J, van Asseldonk M, van den Berg E, Bridge J et al. Upregulation of the transcription factor TFEB in t(6;11)(p21;q13)-positive renal cell carcinomas due to promoter substitution. Hum Mol Genet 2003; 12: 1661–1669.
Kim SS, Shago M, Kaustov L, Boutros PC, Clendening JW, Sheng Y et al. CUL7 is a novel antiapoptotic oncogene. Cancer Res 2007; 67: 9616–9622.
Lewis BC, Prescott JE, Campbell SE, Shim H, Orlowski RZ, Dang CV . Tumor induction by the c-Myc target genes rcl and lactate dehydrogenase A. Cancer Res 2000; 60: 6178–6783.
Varela-Rey M, Fernández-Ramos D, Martínez-López N, Embade N, Gómez-Santos L, Beraza N et al. Impaired liver regeneration in mice lacking glycine N-methyltransferase. Hepatology 2009; 50: 443–452.
Ghiorghi YK, Zeller KI, Dang CV, Kaminski PA . The c-MYC target gene Rcl (C6orf108) encodes a novel enzyme, deoxynucleoside 5’-monophosphate N-glycosidase. J Biol Chem 2007; 282: 8150–8156.
Thornalley PJ . Protein and nucleotide damage by glyoxal and methylglyoxal in physiological systems–role in aging and disease. Drug Metabol Drug Interact 2008; 23: 125–150.
Vander Heiden MG, Cantley LC, Thompson CB . Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 2009; 324: 1029–1033.
Levine AJ, Puzio-Kuter AM . The control of the metabolic switch in cancers by oncogenes and tumor suppressor genes. Science 2010; 330: 1340–1344.
Bair III WB, Cabell CM, Uchida K, Bause AS, Wondrak GT . GLO1 overexpression in human malignant melanoma. Melanoma Res 2010; 20: 85–96.
Antognelli C, Mezzasoma L, Fettucciari K, Mearini E, Talesa VN . Role of glyoxalase I in the proliferation and apoptosis control of human LNCaP and PC3 prostate cancer cells. Prostate 2013; 73: 121–132.
Antognelli C, Mezzasoma L, Fettucciari K, Talesa VN . A novel mechanism of methylglyoxal cytotoxity in prostate cancer cells. Int J Biochem Cell Biol 2013; 45: 836–844.
Ranganathan S, Ciaccio PJ, Walsh ES, Tew KD . Genomic sequence of human glyoxalase-I: analysis of promoter activity and its regulation. Gene 1999; 240: 149–155.
Mitsuishi Y, Taguchi K, Kawatani Y, Shibata T, Nukiwa T, Aburatani H et al. Nrf2 redirects glucose and glutamine into anabolic pathways in metabolic reprogramming. Cancer Cell 2012; 22: 66–79.
Figueroa ME, Abdel-Wahab O, Lu C, Ward PS, Patel J, Shih A et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 2010; 18: 553–567.
Nakamura Y, Migita T, Hosoda F, Okada N, Gotoh M, Arai Y et al. Krüppel-like factor 12 plays a significant role in poorly differentiated gastric cancer progression. Int J Cancer 2009; 125: 1859–1867.
Yanagihara K, Tanaka H, Takigahira M, Ino Y, Yamaguchi Y, Toge T et al. Establishment of two cell lines from human gastric scirrhous carcinoma that possess the potential to metastasize spontaneously in nude mice. Cancer Sci 2004; 95: 575–582.
Yagi T, Morimoto A, Eguchi M, Hibi S, Sako M, Ishii E et al. Identification of a gene expression signature associated with pediatric AML prognosis. Blood 2003; 102: 1849–1856.
Oray B, Norton SJ . Glyoxalase I from mouse liver. Methods Enzymol 1982; 90: 542–546.
Acknowledgements
We thank Setsuo Hirohashi for generous support and encouragement; Yukihiro Nakanishi for advice on histological classification of gastric tumors; Tokuki Sakiyama and Go Maeno for helping with the analysis of the aCGH data; Jun Yasuda for advice on the RNA interference techniques; Yu Nakamura, Michiyo Fukushima, Satomi Uryu and Yasuko Kuwabara for providing considerable contributions to the sample preparation of BAC DNA and patients’ DNA and array hybridization; and Sayaka Kadoguchi and Kenjiro Kami for advice on metabolome analysis. This research was supported in part by a Grant-in-Aid for the Comprehensive 10-Year-Strategy for Cancer Control and from the Ministry of Health, Labor and Welfare, Japan, a grant from the New Energy and Industrial Technology Development Organization (NEDO), Japan and a grant from the National Institute of Biomedical Innovation (NiBio), Japan. MM and NK were recipients of a Research Resident Fellowship from the Foundation for Promotion of Cancer Research in Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Hosoda, F., Arai, Y., Okada, N. et al. Integrated genomic and functional analyses reveal glyoxalase I as a novel metabolic oncogene in human gastric cancer. Oncogene 34, 1196–1206 (2015). https://doi.org/10.1038/onc.2014.57
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2014.57
This article is cited by
-
Development of pathway-oriented screening to identify compounds to control 2-methylglyoxal metabolism in tumor cells
Communications Chemistry (2023)
-
Ovarian dysfunction following prenatal exposure to an insecticide, chlordecone, associates with altered epigenetic features
Epigenetics & Chromatin (2019)
-
MMSET I acts as an oncoprotein and regulates GLO1 expression in t(4;14) multiple myeloma cells
Leukemia (2019)
-
Nucleolar-nucleoplasmic shuttling of TARG1 and its control by DNA damage-induced poly-ADP-ribosylation and by nucleolar transcription
Scientific Reports (2018)
-
Hormetic potential of methylglyoxal, a side-product of glycolysis, in switching tumours from growth to death
Scientific Reports (2017)