Abstract
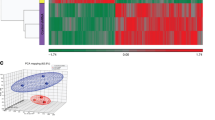
TAF15 (formerly TAFII68) is a member of the FET (FUS, EWS, TAF15) family of RNA- and DNA-binding proteins whose genes are frequently translocated in sarcomas. By performing global gene expression profiling, we found that TAF15 knockdown affects the expression of a large subset of genes, of which a significant percentage is involved in cell cycle and cell death. In agreement, TAF15 depletion had a growth-inhibitory effect and resulted in increased apoptosis. Among the TAF15-regulated genes, targets of microRNAs (miRNAs) generated from the onco-miR-17 locus were overrepresented, with CDKN1A/p21 being the top miRNAs-targeted gene. Interestingly, the levels of onco-miR-17 locus coded miRNAs (miR-17-5p and miR-20a) were decreased upon TAF15 depletion and shown to affect the post-transcriptional regulation of TAF15-dependent genes, such as CDKN1A/p21. Thus, our results demonstrate that TAF15 is required to regulate gene expression of cell cycle regulatory genes post-transcriptionally through a pathway involving miRNAs. The findings that high TAF15 levels are needed for rapid cellular proliferation and that endogenous TAF15 levels decrease during differentiation strongly suggest that TAF15 is a key regulator of maintaining a highly proliferative rate of cellular homeostasis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Bertolotti A, Lutz Y, Heard DJ, Chambon P, Tora L . hTAF(II)68, a novel RNA/ssDNA-binding protein with homology to the pro-oncoproteins TLS/FUS and EWS is associated with both TFIID and RNA polymerase II. EMBO J 1996; 15: 5022–5031.
Butler JE, Kadonaga JT . The RNA polymerase II core promoter: a key component in the regulation of gene expression. Genes Dev 2002; 16: 2583–2592.
Tora L . A unified nomenclature for TATA box binding protein (TBP)-associated factors (TAFs) involved in RNA polymerase II transcription. Genes Dev 2002; 16: 673–675.
Sankar S, Lessnick SL . Promiscuous partnerships in Ewing’s sarcoma. Cancer Genet 2011; 204: 351–365.
Bertolotti A, Bell B, Tora L . The N-terminal domain of human TAFII68 displays transactivation and oncogenic properties. Oncogene 1999; 18: 8000–8010.
Law WJ, Cann KL, Hicks GG . TLS, EWS and TAF15: a model for transcriptional integration of gene expression. Brief Funct Genomic Proteomic 2006; 5: 8–14.
Jobert L, Argentini M, Tora L . PRMT1 mediated methylation of TAF15 is required for its positive gene regulatory function. Exp Cell Res 2009; 315: 1273–1286.
Jobert L, Pinzon N, Van Herreweghe E, Jady BE, Guialis A, Kiss T et al. Human U1 snRNA forms a new chromatin-associated snRNP with TAF15. EMBO Rep 2009; 10: 494–500.
Pasquinelli AE . MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat Rev Genet 2012; 13: 271–282.
Gregory RI, Yan KP, Amuthan G, Chendrimada T, Doratotaj B, Cooch N et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 2004; 432: 235–240.
McKinsey EL, Parrish JK, Irwin AE, Niemeyer BF, Kern HB, Birks DK et al. A novel oncogenic mechanism in Ewing sarcoma involving IGF pathway targeting by EWS/Fli1-regulated microRNAs. Oncogene 2011; 30: 4910–4920.
De Vito C, Riggi N, Suva ML, Janiszewska M, Horlbeck J, Baumer K et al. Let-7a is a direct EWS-FLI-1 target implicated in Ewing’s sarcoma development. PLoS One 2011; 6: e23592.
de la Grange P, Dutertre M, Martin N, Auboeuf D . FAST DB: a website resource for the study of the expression regulation of human gene products. Nucleic Acids Res 2005; 33: 4276–4284.
Hsu SD, Lin FM, Wu WY, Liang C, Huang WC, Chan WL et al. miRTarBase: a database curates experimentally validated microRNA-target interactions. Nucleic Acids Res 2011; 39: D163–D169.
Mendell JT . miRiad roles for the miR-17-92 cluster in development and disease. Cell 2008; 133: 217–222.
Sherr CJ, Roberts JM . CDK inhibitors: positive and negative regulators of G1-phase progression. Genes Dev 1999; 13: 1501–1512.
Fontana L, Pelosi E, Greco P, Racanicchi S, Testa U, Liuzzi F et al. MicroRNAs 17-5p-20a-106a control monocytopoiesis through AML1 targeting and M-CSF receptor upregulation. Nat Cell Biol 2007; 9: 775–787.
Ivanovska I, Ball AS, Diaz RL, Magnus JF, Kibukawa M, Schelter JM et al. MicroRNAs in the miR-106b family regulate p21/CDKN1A and promote cell cycle progression. Mol Cell Biol 2008; 28: 2167–2174.
Petrocca F, Vecchione A, Croce CM . Emerging role of miR-106b-25/miR-17-92 clusters in the control of transforming growth factor beta signaling. Cancer Res 2008; 68: 8191–8194.
Fontana L, Fiori ME, Albini S, Cifaldi L, Giovinazzi S, Forloni M et al. Antagomir-17-5p abolishes the growth of therapy-resistant neuroblastoma through p21 and BIM. PLoS One 2008; 3: e2236.
Beveridge NJ, Tooney PA, Carroll AP, Tran N, Cairns MJ . Down-regulation of miR-17 family expression in response to retinoic acid induced neuronal differentiation. Cell signal 2009; 21: 1837–1845.
Thiele CJ, Reynolds CP, Israel MA . Decreased expression of N-myc precedes retinoic acid-induced morphological differentiation of human neuroblastoma. Nature 1985; 313: 404–406.
Rossi A, Granata F, Augusti-Tocco G, Canu N, Levi A, Possenti R . Expression in murine and human neuroblastoma cell lines of VGF, a tissue specific protein. Int J Dev Neurosci 1992; 10: 527–534.
Spitzer JI, Ugras S, Runge S, Decarolis P, Antonescu C, Tuschl T et al. mRNA and protein levels of FUS, EWSR1, and TAF15 are upregulated in liposarcoma. Genes, Chromosomes Cancer 2011; 50: 338–347.
Le Brigand K, Russell R, Moreilhon C, Rouillard JM, Jost B, Amiot F et al. An open-access long oligonucleotide microarray resource for analysis of the human and mouse transcriptomes. Nucleic Acids Res 2006; 34: e87.
Guil S, Caceres JF . The multifunctional RNA-binding protein hnRNP A1 is required for processing of miR-18a. Nature structural & molecular biology. Nat Struct Mol Biol 2007; 14: 591–596.
Hoell JI, Larsson E, Runge S, Nusbaum JD, Duggimpudi S, Farazi TA et al. RNA targets of wild-type and mutant FET family proteins. Nat Struct Mol Biol 2011; 18: 1428–1431.
Chaulk SG, Thede GL, Kent OA, Xu Z, Gesner EM, Veldhoen RA et al. Role of pri-miRNA tertiary structure in miR-17∼92 miRNA biogenesis. RNA biol 2011; 8: 1105–1114.
Blechingberg J, Holm IE, Nielsen AL . Characterization and expression analysis in the developing embryonic brain of the porcine FET family: FUS, EWS, and TAF15. Gene 2012; 493: 27–35.
Andersson MK, Stahlberg A, Arvidsson Y, Olofsson A, Semb H, Stenman G et al. The multifunctional FUS, EWS and TAF15 proto-oncoproteins show cell type-specific expression patterns and involvement in cell spreading and stress response. BMC Cell Biol 2008; 9: 37.
Greaves MF, Wiemels J . Origins of chromosome translocations in childhood leukaemia. Nat Rev Cancer 2003; 3: 639–649.
Pall GS, Hamilton AJ . Improved northern blot method for enhanced detection of small RNA. Nat Protoc 2008; 3: 1077–1084.
Lee Y, Kim VN . In vitro and in vivo assays for the activity of Drosha complex. Methods Enzymol 2007; 427: 89–106.
Acknowledgements
We are very grateful to JF Caceres, VN Kim and ME Fiori for providing the Pri-17–92, Flag-DGCR8 and pGL3-p21-3′UTR plasmids, respectively. We also thank N Charlet Berguerand and M Morlando for the advice in miRNA processing assay, D Devys, M Fournier, A Helmrich, P Bheda and P Laneve for critically reading the manuscript, R Contzler for the XCELLigence analysis and the IGBMC cell culture facility. This work was supported by funds from CNRS, INSERM and AICR (09–0258) grants. MB was supported by the Fondation Recherche Medicale and LJ by MRET and Association pour la Recherche sur le Cancer.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Ballarino, M., Jobert, L., Dembélé, D. et al. TAF15 is important for cellular proliferation and regulates the expression of a subset of cell cycle genes through miRNAs. Oncogene 32, 4646–4655 (2013). https://doi.org/10.1038/onc.2012.490
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2012.490
Keywords
This article is cited by
-
TAF15 promotes cell proliferation, migration and invasion of gastric cancer via activation of the RAF1/MEK/ERK signalling pathway
Scientific Reports (2023)
-
LncRNA GAS5 activates the HIF1A/VEGF pathway by binding to TAF15 to promote wound healing in diabetic foot ulcers
Laboratory Investigation (2021)
-
Potential biomarkers of childhood brain tumor identified by proteomics of cerebrospinal fluid from extraventricular drainage (EVD)
Scientific Reports (2021)
-
Interaction between the BAG1S isoform and HSP70 mediates the stability of anti-apoptotic proteins and the survival of osteosarcoma cells expressing oncogenic MYC
BMC Cancer (2019)
-
α-Amanitin Restrains Cancer Relapse from Drug-Tolerant Cell Subpopulations via TAF15
Scientific Reports (2016)