Abstract
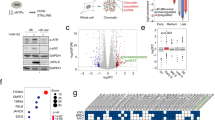
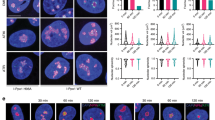
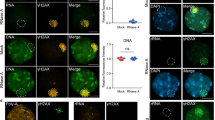
Ribosomal proteins (RPs) activate the p53 tumour-suppressor protein upon disruption of the nucleolus. However, the exact mechanisms for p53 transcriptional activation through RPs are not well understood. We show that the RPL11 is rapidly but transiently recruited at promoter sites of p53-regulated genes upon nucleolar stress induced by actinomycin D (ActD). Characterisation of molecular events at p53 promoter sites shows that L11 is required for the recruitment of p53 transcriptional co-activators p300/CBP and p53 K382 acetylation. We found that direct binding to Mdm2 E3 ligase and NEDDylation of L11 are critical regulators for L11 promoter recruitment. Our data suggest that binding of L11 to Mdm2 at the promoter results in relief from Mdm2-mediated transcriptional repression of p53. Analysis of chromatin and RNA polymerase II markers suggests that L11 is involved in the initiation step of transcriptional activation. Furthermore, analysis of 36 ActD-induced genes shows that L11 and NEDD8 are global regulators of the p53 activation response. The studies provide insights on how nucleolar stress through L11 and NEDD8 can activate the transcriptional activity of p53.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Ashcroft M, Taya Y, Vousden KH . (2000). Stress signals utilize multiple pathways to stabilize p53. Mol Cell Biol 20: 3224–3233.
Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z et al. (2007). High-resolution profiling of histone methylations in the human genome. Cell 129: 823–837.
Bernstein BE, Kamal M, Lindblad-Toh K, Bekiranov S, Bailey DK, Huebert DJ et al. (2005). Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120: 169–181.
Bhat KP, Itahana K, Jin A, Zhang Y . (2004). Essential role of ribosomal protein L11 in mediating growth inhibition-induced p53 activation. EMBO J 23: 2402–2412.
Boulon S, Westman BJ, Hutten S, Boisvert FM, Lamond AI . (2010). The nucleolus under stress. Mol Cell 40: 216–227.
Brooks CL, Gu W . (2003). Ubiquitination, phosphorylation and acetylation: the molecular basis for p53 regulation. Curr Opin Cell Biol 15: 164–171.
Choong ML, Yang H, Lee MA, Lane DP . (2009). Specific activation of the p53 pathway by low dose actinomycin D: a new route to p53 based cyclotherapy. Cell Cycle 8: 2810–2818.
Dai MS, Arnold H, Sun XX, Sears R, Lu H . (2007). Inhibition of c-Myc activity by ribosomal protein L11. EMBO J 26: 3332–3345.
Dai MS, Sun XX, Lu H . (2010). Ribosomal protein L11 associates with c-Myc at 5 S rRNA and tRNA genes and regulates their expression. J Biol Chem 285: 12587–12594.
Dikic I, Wakatsuki S, Walters KJ . (2009). Ubiquitin-binding domains–from structures to functions. Nat Rev Mol Cell Biol 10: 659–671.
Donati G, Bertoni S, Brighenti E, Vici M, Treré D, Volarevic S et al. (2011). The balance between rRNA and ribosomal protein synthesis up- and downregulates the tumour suppressor p53 in mammalian cells. Oncogene 30: 3274–3288.
el-Deiry WS, Tokino T, Waldman T, Oliner JD, Velculescu VE, Burrell M et al. (1995). Topological control of p21WAF1/CIP1 expression in normal and neoplastic tissues. Cancer Res 55: 2910–2919.
Espinosa JM . (2008). Mechanisms of regulatory diversity within the p53 transcriptional network. Oncogene 27: 4013–4023.
Espinosa JM, Verdun RE, Emerson BM . (2003). p53 functions through stress- and promoter-specific recruitment of transcription initiation components before and after DNA damage. Mol Cell 12: 1015–1027.
Fuchs SM, Laribee RN, Strahl BD . (2009). Protein modifications in transcription elongation. Biochim Biophys Acta 1789: 26–36.
Fumagalli S, Di Cara A, Neb-Gulati A, Natt F, Schwemberger S, Hall J et al. (2009). Absence of nucleolar disruption after impairment of 40S ribosome biogenesis reveals an rpL11-translation-dependent mechanism of p53 induction. Nat Cell Biol 11: 501–508.
Gu W, Roeder RG . (1997). Activation of p53 sequence-specific DNA binding by acetylation of the p53 C-terminal domain. Cell 90: 595–606.
Horn HF, Vousden KH . (2008). Cooperation between the ribosomal proteins L5 and L11 in the p53 pathway. Oncogene 27: 5774–5784.
Ito A, Lai CH, Zhao X, Saito S, Hamilton MH, Appella E et al. (2001). p300/CBP-mediated p53 acetylation is commonly induced by p53-activating agents and inhibited by MDM2. EMBO J 20: 1331–1340.
Kruse JP, Gu W . (2009). Modes of p53 regulation. Cell 137: 609–622.
Lennartsson A, Ekwall K . (2009). Histone modification patterns and epigenetic codes. Biochim Biophys Acta 1790: 863–868.
Lindstrom MS . (2009). Emerging functions of ribosomal proteins in gene-specific transcription and translation. Biochem Biophys Res Commun 379: 167–170.
Lindstrom MS, Jin A, Deisenroth C, White Wolf G, Zhang Y . (2007). Cancer-associated mutations in the MDM2 zinc finger domain disrupt ribosomal protein interaction and attenuate MDM2-induced p53 degradation. Mol Cell Biol 27: 1056–1068.
Liu L, Scolnick DM, Trievel RC, Zhang HB, Marmorstein R, Halazonetis TD et al. (1999). p53 sites acetylated in vitro by PCAF and p300 are acetylated in vivo in response to DNA damage. Mol Cell Biol 19: 1202–1209.
Lohrum MA, Ludwig RL, Kubbutat MH, Hanlon M, Vousden KH . (2003). Regulation of HDM2 activity by the ribosomal protein L11. Cancer Cell 3: 577–587.
Macias E, Jin A, Deisenroth C, Bhat K, Mao H, Lindstrom MS et al. (2010). An ARF-independent c-MYC-activated tumor suppression pathway mediated by ribosomal protein-Mdm2 interaction. Cancer Cell 18: 231–243.
Momand J, Zambetti GP, Olson DC, George D, Levine AJ . (1992). The mdm-2 oncogene product forms a complex with the p53 protein and inhibits p53-mediated transactivation. Cell 69: 1237–1245.
Montanaro L, Trere D, Derenzini M . (2008). Nucleolus, ribosomes, and cancer. Am J Pathol 173: 301–310.
Perry RP . (2007). Balanced production of ribosomal proteins. Gene 401: 1–3.
Perry RP, Kelley DE . (1970). Inhibition of RNA synthesis by actinomycin D: characteristic dose-response of different RNA species. J Cell Physiol 76: 127–139.
Phatnani HP, Greenleaf AL . (2006). Phosphorylation and functions of the RNA polymerase II CTD. Genes Dev 20: 2922–2936.
Prives C, Manley JL . (2001). Why is p53 acetylated? Cell 107: 815–818.
Rabut G, Peter M . (2008). Function and regulation of protein neddylation protein modifications: beyond the usual suspects’ review series. EMBO Rep 9: 969–976.
Rubbi CP, Milner J . (2003). Disruption of the nucleolus mediates stabilization of p53 in response to DNA damage and other stresses. EMBO J 22: 6068–6077.
Sakaguchi K, Herrera JE, Saito S, Miki T, Bustin M, Vassilev A et al. (1998). DNA damage activates p53 through a phosphorylation–acetylation cascade. Genes Dev 12: 2831–2841.
Saville MK, Sparks A, Xirodimas DP, Wardrop J, Stevenson LF, Bourdon JC et al. (2004). Regulation of p53 by the ubiquitin-conjugating enzymes UbcH5B/C in vivo. J Biol Chem 279: 42169–42181.
Sun XX, Wang YG, Xirodimas DP, Dai MS . (2010). Perturbation of 60 S ribosomal biogenesis results in ribosomal protein L5- and L11-dependent p53 activation. J Biol Chem 285: 25812–25821.
Sundqvist A, Liu G, Mirsaliotis A, Xirodimas DP . (2009). Regulation of nucleolar signalling to p53 through NEDDylation of L11. EMBO Rep 10: 1132–1139.
Tang Y, Zhao W, Chen Y, Zhao Y, Gu W . (2008). Acetylation is indispensable for p53 activation. Cell 133: 612–626.
Thut CJ, Goodrich JA, Tjian R . (1997). Repression of p53-mediated transcription by MDM2: a dual mechanism. Genes Dev 11: 1974–1986.
Wade M, Wang YV, Wahl GM . (2010). The p53 orchestra: Mdm2 and Mdmx set the tone. Trends Cell Biol 20: 299–309.
Wang M, Medeiros BC, Erba HP, Deangelo DJ, Giles FJ, Swords RT . (2011). Targeting protein neddylation: a novel therapeutic strategy for the treatment of cancer. Expert Opin Ther Targets 15: 253–264.
Wang YV, Wade M, Wong E, Li YC, Rodewald LW, Wahl GM . (2007). Quantitative analyses reveal the importance of regulated Hdmx degradation for p53 activation. Proc Natl Acad Sci USA 104: 12365–12370.
Watson IR, Irwin MS, Ohh M . (2011). NEDD8 pathways in cancer, sine quibus non. Cancer Cell 19: 168–176.
Welchman RL, Gordon C, Mayer RJ . (2005). Ubiquitin and ubiquitin-like proteins as multifunctional signals. Nat Rev Mol Cell Biol 6: 599–609.
Xirodimas DP . (2008). Novel substrates and functions for the ubiquitin-like molecule NEDD8. Biochem Soc Trans 36: 802–806.
Xirodimas DP, Sundqvist A, Nakamura A, Shen L, Botting C, Hay RT . (2008). Ribosomal proteins are targets for the NEDD8 pathway. EMBO Rep 9: 280–286.
Zhang Y, Lu H . (2009). Signaling to p53: ribosomal proteins find their way. Cancer Cell 16: 369–377.
Acknowledgements
Our research in the DPX laboratory is supported by the Association for International Cancer Research (AICR) and INSERM. DPX is an AICR Research Fellow.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Mahata, B., Sundqvist, A. & Xirodimas, D. Recruitment of RPL11 at promoter sites of p53-regulated genes upon nucleolar stress through NEDD8 and in an Mdm2-dependent manner. Oncogene 31, 3060–3071 (2012). https://doi.org/10.1038/onc.2011.482
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2011.482
Keywords
This article is cited by
-
Deciphering the role of neddylation in tumor microenvironment modulation: common outcome of multiple signaling pathways
Biomarker Research (2024)
-
Loss of RPS27a expression regulates the cell cycle, apoptosis, and proliferation via the RPL11-MDM2-p53 pathway in lung adenocarcinoma cells
Journal of Experimental & Clinical Cancer Research (2022)
-
Insights into the post-translational modification and its emerging role in shaping the tumor microenvironment
Signal Transduction and Targeted Therapy (2021)
-
NEDDylation promotes nuclear protein aggregation and protects the Ubiquitin Proteasome System upon proteotoxic stress
Nature Communications (2018)
-
NEDDylation antagonizes ubiquitination of proliferating cell nuclear antigen and regulates the recruitment of polymerase η in response to oxidative DNA damage
Protein & Cell (2017)