Abstract
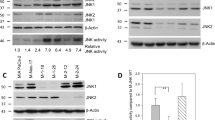
The c-Jun N-terminal kinases (JNKs) are members of the mitogen-activated protein kinase family and have been implicated in tumorigenesis. One isoform in particular, JNK2α, has been shown to be frequently activated in primary brain tumors, to enhance several tumorigenic phenotypes and to increase tumor formation in mice. As JNK is frequently activated in non-small cell lung carcinoma (NSCLC), we investigated the role of the JNK2α isoform in NSCLC formation by examining its expression in primary tumors and by modulating its expression in cultured cell lines. We discovered that 60% of the tested primary NSCLC tumors had three-fold higher JNK2 protein and two- to three-fold higher JNK2α mRNA expression than normal lung control tissue. To determine the importance of JNK2α in NSCLC progression, we reduced JNK2α expression in multiple NSCLC cell lines using short hairpin RNA. Cell lines deficient in JNK2α had decreased cellular growth and anchorage-independent growth, and the tumors were four-fold smaller in mass. To elucidate the mechanism by which JNK2α induces NSCLC growth, we analyzed the JNK substrate, signal transducer and activator of transcription 3 (STAT3). Our data demonstrates for the first time that JNK2α can regulate the transcriptional activity of STAT3 by phosphorylating the Ser727 residue, thereby regulating the expression of oncogenic genes, such as c-Myc. Furthermore, reintroduction of JNK2α2 or STAT3 restored the tumorigenicity of the NSCLC cells, demonstrating that JNK2α is important for NSCLC progression. Our studies reveal a novel mechanism in which phosphorylation of STAT3 is mediated by a constitutively active JNK2 isoform, JNK2α.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Achcar Rde O, Cagle PT, Jagirdar J . (2007). Expression of activated and latent signal transducer and activator of transcription 3 in 303 non-small cell lung carcinomas and 44 malignant mesotheliomas: possible role for chemotherapeutic intervention. Arch Pathol Lab Med 131: 1350–1360.
Alvarez JV, Greulich H, Sellers WR, Meyerson M, Frank DA . (2006). Signal transducer and activator of transcription 3 is required for the oncogenic effects of non-small-cell lung cancer-associated mutations of the epidermal growth factor receptor. Cancer Res 66: 3162–3168.
Arbour N, Naniche D, Homann D, Davis RJ, Flavell RA, Oldstone MB . (2002). c-Jun NH(2)-terminal kinase (JNK)1 and JNK2 signaling pathways have divergent roles in CD8(+) T cell-mediated antiviral immunity. J Exp Med 195: 801–810.
Berner JM, Sorlie T, Mertens F, Henriksen J, Saeter G, Mandahl N et al. (1999). Chromosome band 9p21 is frequently altered in malignant peripheral nerve sheath tumors: studies of CDKN2A and other genes of the pRB pathway. Genes Chromosomes Cancer 26: 151–160.
Bhattacharya S, Ray RM, Johnson LR . (2005). STAT3-mediated transcription of Bcl-2, Mcl-1 and c-IAP2 prevents apoptosis in polyamine-depleted cells. Biochem J 392: 335–344.
Bode AM, Dong Z . (2007). The functional contrariety of JNK. Mol Carcinog 46: 591–598.
Bogoyevitch MA . (2006). The isoform-specific functions of the c-Jun N-terminal Kinases (JNKs): differences revealed by gene targeting. Bioessays 28: 923–934.
Bogoyevitch MA, Arthur PG . (2008). Inhibitors of c-Jun N-terminal kinases: JuNK no more? Biochim Biophys Acta 1784: 76–93.
Broers JL, Viallet J, Jensen SM, Pass H, Travis WD, Minna JD et al. (1993). Expression of c-myc in progenitor cells of the bronchopulmonary epithelium and in a large number of non-small cell lung cancers. Am J Respir Cell Mol Biol 9: 33–43.
Bromberg JF, Wrzeszczynska MH, Devgan G, Zhao Y, Pestell RG, Albanese C et al. (1999). Stat3 as an oncogene. Cell 98: 295–303.
Butterfield L, Storey B, Maas L, Heasley LE . (1997). c-Jun NH2-terminal kinase regulation of the apoptotic response of small cell lung cancer cells to ultraviolet radiation. J Biol Chem 272: 10110–10116.
Chen N, Nomura M, She QB, Ma WY, Bode AM, Wang L et al. (2001). Suppression of skin tumorigenesis in c-Jun NH(2)-terminal kinase-2-deficient mice. Cancer Res 61: 3908–3912.
Cheng A, Uetani N, Simoncic PD, Chaubey VP, Lee-Loy A, McGlade CJ et al. (2002). Attenuation of leptin action and regulation of obesity by protein tyrosine phosphatase 1B. Dev Cell 2: 497–503.
Chung J, Uchida E, Grammer TC, Blenis J . (1997). STAT3 serine phosphorylation by ERK-dependent and -independent pathways negatively modulates its tyrosine phosphorylation. Mol Cell Biol 17: 6508–6516.
Cui J, Holgado-Madruga M, Su W, Tsuiki H, Wedegaertner P, Wong AJ . (2005). Identification of a specific domain responsible for JNK2alpha2 autophosphorylation. J Biol Chem 280: 9913–9920.
Ford J, Ahmed S, Allison S, Jiang M, Milner J . (2008). JNK2-dependent regulation of SIRT1 protein stability. Cell Cycle 7: 3091–3097.
Gao SP, Bromberg JF . (2006). Touched and moved by STAT3. Sci STKE 2006: pe30.
Gazzeri S, Brambilla E, Caron de Fromentel C, Gouyer V, Moro D, Perron P et al. (1994). p53 genetic abnormalities and myc activation in human lung carcinoma. Int J Cancer 58: 24–32.
Gough DJ, Corlett A, Schlessinger K, Wegrzyn J, Larner AC, Levy DE . (2009). Mitochondrial STAT3 supports Ras-dependent oncogenic transformation. Science 324: 1713–1716.
Haura EB, Zheng Z, Song L, Cantor A, Bepler G . (2005). Activated epidermal growth factor receptor-Stat-3 signaling promotes tumor survival in vivo in non-small cell lung cancer. Clin Cancer Res 11: 8288–8294.
Khatlani TS, Wislez M, Sun M, Srinivas H, Iwanaga K, Ma L et al. (2007). c-Jun N-terminal kinase is activated in non-small-cell lung cancer and promotes neoplastic transformation in human bronchial epithelial cells. Oncogene 26: 2658–2666.
Kortylewski M, Jove R, Yu H . (2005). Targeting STAT3 affects melanoma on multiple fronts. Cancer Metastasis Rev 24: 315–327.
Lee C, Dhillon J, Wang MY, Gao Y, Hu K, Park E et al. (2008). Targeting YB-1 in HER-2 overexpressing breast cancer cells induces apoptosis via the mTOR/STAT3 pathway and suppresses tumor growth in mice. Cancer Res 68: 8661–8666.
Lee SO, Lou W, Johnson CS, Trump DL, Gao AC . (2004). Interleukin-6 protects LNCaP cells from apoptosis induced by androgen deprivation through the Stat3 pathway. Prostate 60: 178–186.
Lim CP, Cao X . (1999). Serine phosphorylation and negative regulation of Stat3 by JNK. J Biol Chem 274: 31055–31061.
Lufei C, Koh TH, Uchida T, Cao X . (2007). Pin1 is required for the Ser727 phosphorylation-dependent Stat3 activity. Oncogene 26: 7656–7664.
Maeda S, Karin M . (2003). Oncogene at last--c-Jun promotes liver cancer in mice. Cancer Cell 3: 102–104.
Nitta R, Okada Y, Hirokawa N . (2008). Structural model for strain-dependent microtubule activation of Mg-ADP release from kinesin. Nat Struct Mol Biol 15: 1067–1075.
Nitta RT, Jameson SA, Kudlow BA, Conlan LA, Kennedy BK . (2006). Stabilization of the retinoblastoma protein by A-type nuclear lamins is required for INK4A-mediated cell cycle arrest. Mol Cell Biol 26: 5360–5372.
Nitta RT, Smith CL, Kennedy BK . (2007). Evidence that proteasome-dependent degradation of the retinoblastoma protein in cells lacking A-type lamins occurs independently of gankyrin and MDM2. PLoS One 2: e963.
Oleinik NV, Krupenko NI, Krupenko SA . (2007). Cooperation between JNK1 and JNK2 in activation of p53 apoptotic pathway. Oncogene 26: 7222–7230.
Onishi A, Chen Q, Humtsoe JO, Kramer RH . (2008). STAT3 signaling is induced by intercellular adhesion in squamous cell carcinoma cells. Exp Cell Res 314: 377–386.
Potapova O, Gorospe M, Bost F, Dean NM, Gaarde WA, Mercola D et al. (2000). c-Jun N-terminal kinase is essential for growth of human T98G glioblastoma cells. J Biol Chem 275: 24767–24775.
Qin HR, Kim HJ, Kim JY, Hurt EM, Klarmann GJ, Kawasaki BT et al. (2008). Activation of signal transducer and activator of transcription 3 through a phosphomimetic serine 727 promotes prostate tumorigenesis independent of tyrosine 705 phosphorylation. Cancer Res 68: 7736–7741.
Rapp UR, Korn C, Ceteci F, Karreman C, Luetkenhaus K, Serafin V et al. (2009). MYC is a metastasis gene for non-small-cell lung cancer. PLoS One 4: e6029.
Rodrigues GA, Park M, Schlessinger J . (1997). Activation of the JNK pathway is essential for transformation by the Met oncogene. Embo J 16: 2634–2645.
Shair KH, Bendt KM, Edwards RH, Bedford EC, Nielsen JN, Raab-Traub N . (2007). EBV latent membrane protein 1 activates Akt, NFkappaB, and Stat3 in B cell lymphomas. PLoS Pathog 3: e166.
Tang GS, Cai JM, Ni J, Xiang YS, Cui JG, Zhu D et al. (2006). [Effects of STAT3 antisense oligodeoxynucleotides on apoptosis and proliferation of mouse melanoma cell line B16]. Ai Zheng 25: 269–274.
Tsuiki H, Tnani M, Okamoto I, Kenyon LC, Emlet DR, Holgado-Madruga M et al. (2003). Constitutively active forms of c-Jun NH2-terminal kinase are expressed in primary glial tumors. Cancer Res 63: 250–255.
Weerasinghe P, Garcia GE, Zhu Q, Yuan P, Feng L, Mao L et al. (2007). Inhibition of Stat3 activation and tumor growth suppression of non-small cell lung cancer by G-quartet oligonucleotides. Int J Oncol 31: 129–136.
Wen Z, Zhong Z, Darnell Jr JE . (1995). Maximal activation of transcription by Stat1 and Stat3 requires both tyrosine and serine phosphorylation. Cell 82: 241–250.
Wu X, Tu X, Joeng KS, Hilton MJ, Williams DA, Long F . (2008). Rac1 activation controls nuclear localization of beta-catenin during canonical Wnt signaling. Cell 133: 340–353.
Xie W, Su K, Wang D, Paterson AJ, Kudlow JE . (1997). MDA468 growth inhibition by EGF is associated with the induction of the cyclin-dependent kinase inhibitor p21WAF1. Anticancer Res 17: 2627–2633.
Xu X, Heidenreich O, Kitajima I, McGuire K, Li Q, Su B et al. (1996). Constitutively activated JNK is associated with HTLV-1 mediated tumorigenesis. Oncogene 13: 135–142.
Yakut T, Egeli U, Gebitekin C . (2003). Investigation of c-myc and p53 gene alterations in the tumor and surgical borderline tissues of NSCLC and effects on clinicopathologic behavior: by the FISH technique. Lung 181: 245–258.
Yang YM, Bost F, Charbono W, Dean N, McKay R, Rhim JS et al. (2003). C-Jun NH(2)-terminal kinase mediates proliferation and tumor growth of human prostate carcinoma. Clin Cancer Res 9: 391–401.
Yeager T, Stadler W, Belair C, Puthenveettil J, Olopade O, Reznikoff C . (1995). Increased p16 levels correlate with pRb alterations in human urothelial cells. Cancer Res 55: 493–497.
Yeh HH, Giri R, Chang TY, Chou CY, Su WC, Liu HS . (2009). Ha-ras oncogene-induced Stat3 phosphorylation enhances oncogenicity of the cell. DNA Cell Biol 28: 131–139.
Zhang X, Blenis J, Li HC, Schindler C, Chen-Kiang S . (1995). Requirement of serine phosphorylation for formation of STAT-promoter complexes. Science 267: 1990–1994.
Zhang X, Zhang J, Wang L, Wei H, Tian Z . (2007). Therapeutic effects of STAT3 decoy oligodeoxynucleotide on human lung cancer in xenograft mice. BMC Cancer 7: 149.
Acknowledgements
We are grateful to Emily Piccione, Shawn Badal, Tiffany Lieu and Marina Holgado-Madruga for their useful discussions and critical review of the paper. This work was supported by the Mark Linder/American Brain Tumor Association Fellowship, NIH grants CA69495 and CA124832 and a research grant from the National Brain Tumor Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Rights and permissions
About this article
Cite this article
Nitta, R., Del Vecchio, C., Chu, A. et al. The role of the c-Jun N-terminal kinase 2-α-isoform in non-small cell lung carcinoma tumorigenesis. Oncogene 30, 234–244 (2011). https://doi.org/10.1038/onc.2010.414
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.414
Keywords
This article is cited by
-
Casein kinase 2 inhibition sensitizes medulloblastoma to temozolomide
Oncogene (2019)
-
JNKs function as CDK4-activating kinases by phosphorylating CDK4 and p21
Oncogene (2017)
-
γ-Glutamyl transferase 7 is a novel regulator of glioblastoma growth
BMC Cancer (2015)
-
Poly(ADP-ribose) polymerase family member 14 (PARP14) is a novel effector of the JNK2-dependent pro-survival signal in multiple myeloma
Oncogene (2013)
-
Cucurbitacin-I (JSI-124) activates the JNK/c-Jun signaling pathway independent of apoptosis and cell cycle arrest in B Leukemic Cells
BMC Cancer (2011)