Abstract
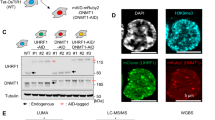
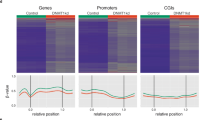
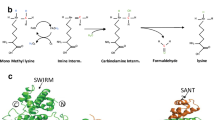
Understanding the mechanisms that link changes in DNA methylation with histone modifications is particularly relevant in the case of tumor suppressor genes that undergo transcriptional silencing in cancer cells in association with promoter CpG island hypermethylation. In this study, we show that two histone lysine methylation marks associated with active transcription, dimethylation of H3K79 (H3K79me2) and trimethylation of H3K4 (H3K4me3), are present in all the unmethylated promoters analysed, and both of them are lost when these promoters become hypermethylated. Most importantly, pharmacological and genetic interventions that cause DNA demethylation and partial recovery of gene transcription, result in the restoration of H3K4me3, but not of H3K79me2. We also show that DOT1L, the major H3K79 histone methyltransferase, is no longer recruited to the promoters that are demethylated after 5-aza-deoxycytidine treatment or genetic deletion of DNA methyltransferases. Knock-down and transfection experiments for DOT1L show that this enzyme has a direct role in maintaining the euchromatic and active status of these genes when unmethylated. These findings suggest that DNA demethylating interventions alone are not able to restore a complete euchromatic status and a full transcriptional reactivation of the epigenetically silenced tumor suppressor genes, and reinforce the necessity of targeting multiple elements of the epigenetics machinery for a successful treatment of malignancies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Akiyama Y, Watkins N, Suzuki H, Jair KW, van Engeland M, Esteller M et al. (2003). GATA-4 and GATA-5 transcription factor genes and potential downstream antitumor target genes are epigenetically silenced in colorectal and gastric cancer. Mol Cell Biol 23: 8429–8439.
Ballestar E, Paz MF, Valle L, Wei S, Fraga MF, Espada J et al. (2003). Methyl-CpG binding proteins identify novel sites of epigenetic inactivation in human cancer. EMBO J 22: 6335–6345.
Berger SL . (2007). The complex language of chromatin regulation during transcription. Nature 447: 407–412.
Cameron EE, Bachman KE, Myohanen S, Herman JG, Baylin SB . (1999). Synergy of demethylation and histone deacetylase inhibition in the re-expression of genes silenced in cancer. Nat Genet 21: 103–107.
Costello JF, Fruhwald MC, Smiraglia DJ, Rush LJ, Robertson GP, Gao X et al. (2000). Aberrant CpG-island methylation has non-random and tumor-type specific patterns. Nat Genet 24: 132–138.
Esteller M . (2008). Epigenetics in cancer. N Engl J Med 358: 1148–1159.
Esteller M, Corn PG, Baylin SB, Herman JG . (2001). A gene hypermethylation profile of human cancer. Cancer Res 61: 3225–3229.
Fahrner JA, Eguchi S, Herman JG, Baylin SB . (2002). Dependence of histone modifications and gene expression on DNA hypermethylation in cancer. Cancer Res 62: 7213–7218.
Feng Q, Wang H, Ng HH, Erdjument-Bromage H, Tempst P, Struhl K et al. (2002). Methylation of H3.lysine 79 is mediated by a new family of HMTases without a SET domain. Curr Biol 12: 1052–1058.
Garcia-Manero G . (2008). Demethylating agents in myeloid malignancies. Curr Opin Oncol 20: 705–710.
Jones PA, Baylin SB . (2007). The epigenomics of cancer. Cell 128: 683.
Kondo Y, Shen L, Issa JP . (2003). Critical role of histone methylation in tumor suppressor gene silencing in colorectal cancer. Mol Cell Biol 23: 206–215.
Kouzarides T . (2007). Chromatin modifications and their function. Cell 128: 693–705.
Lopez-Serra L, Esteller M . (2008). Proteins that bind methylated DNA and human cancer: reading the wrong words. Br J Cancer 98: 1881–1885.
McGarvey KM, Fahrner JA, Greene E, Martens J, Jenuwein T, Baylin SB . (2006). Silenced tumor suppressor genes reactivated by DNA demethylation do not return to a fully euchromatic chromatin state. Cancer Res 66: 3541–3549.
Milne TA, Martin ME, Brock HW, Slany RK, Hess JL . (2005). Leukemogenic MLL fusion proteins bind across a broad region of the Hoxa9 locus, promoting transcription and multiple histone modifications. Cancer Res 65: 11367–11374.
Min J, Feng Q, Li Z, Zhang Y, Xu RM . (2003). Structure of the catalytic domain of human DOT1L, a non-SET domain nucleosomal histone methyltransferase. Cell 112: 711–723.
Ng HH, Feng Q, Wang H, Erdjument-Bromage H, Tempst P, Zhang Y et al. (2002). Lysine methylation within the globular domain of histone H3 by Dot1 is important for telomeric silencing and Sir protein association. Genes Dev 16: 1518–1527.
Nguyen CT, Weisenberger DJ, Velicescu M, Gonzales FA, Lin JC, Liang G et al. (2002). Histone H3-lysine 9 methylation is associated with aberrant gene silencing in cancer cells and is rapidly reversed by 5-aza-2′-deoxicytidine. Cancer Res 62: 6456–6461.
Okada Y, Feng Q, Lin Y, Jiang Q, Li Y, Coffield VM et al. (2005). hDOT1L links histone methylation to leukemogenesis. Cell 121: 167–178.
Oki Y, Aoki E, Issa JP . (2007). Decitabine—bedside to bench. Crit Rev Oncol Hematol 61: 140–152.
Paz MF, Fraga MF, Avila S, Guo M, Pollan M, Herman JG et al. (2003). A systematic profile of DNA methylation in human cancer cell lines. Cancer Res 63: 1114–1121.
Rhee I, Bachman KE, Park BH, Jair KW, Yen RW, Schuebel KE et al. (2002). DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature 416: 552–556.
Simon JA, Lange CA . (2008). Roles of the EZH2 histone methyltransferase in cancer epigenetics. Mutat Res 647: 21–29.
Suzuki H, Gabrielson E, Chen W, Anbazhagan R, van Engeland M, Weijenberg MP et al. (2002). A genomic screen for genes upregulated by demethylation and histone deacetylase inhibition in human colorectal cancer. Nat Genet 31: 141–149.
Turner BM . (2002). Cellular memory and the histone code. Cell 111: 285–291.
van Leeuwen F, Gafken PR, Gottschling DE . (2002). Dot1p modulates silencing in yeast by methylation of the nucleosome core. Cell 109: 745–756.
Wang Z, Zang C, Rosenfeld JA, Schones DE, Barski A, Cuddapah S et al. (2008). Combinatorial patterns of histone acetylations and methylations in the human genome. Nat Genet 40: 897–903.
Wysocka J, Swigut T, Milne TA, Dou Y, Zhang X, Burlingame AL et al. (2005). WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell 121: 859–872.
Yoo CB, Jones PA . (2006). Epigenetic therapy of cancer: past, present and future. Nat Rev Drug Discov 5: 37–50.
Acknowledgements
We thank Dr Yi Zhang for providing us with the hDOT1L expression vector for the transfection experiments. Supported by the European Union Grants FP7 CANCERDIP (HEALTH-F2-2007-200620) and SMARTER (LSHG-CT-2006-037415), the Health (FIS PI08 1345) and Education and Science (I+D+I MCYT08-03 and Consolider MEC09-05) Departments of the Spanish Government, the Health Department of the Catalan Government and the Spanish Association Against Cancer (AECC). FVJ is funded by FCT (Portuguese Foundation for Science and Technology) SFRH/BD/11757/2003. ME is an ICREA (Institució Catalana de Recerca i Estudis Avançats) Research Professor.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Supplementary information
Rights and permissions
About this article
Cite this article
Jacinto, F., Ballestar, E. & Esteller, M. Impaired recruitment of the histone methyltransferase DOT1L contributes to the incomplete reactivation of tumor suppressor genes upon DNA demethylation. Oncogene 28, 4212–4224 (2009). https://doi.org/10.1038/onc.2009.267
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.267
Keywords
This article is cited by
-
Writing of H3K4Me3 overcomes epigenetic silencing in a sustained but context-dependent manner
Nature Communications (2016)
-
Bat3 facilitates H3K79 dimethylation by DOT1L and promotes DNA damage-induced 53BP1 foci at G1/G2 cell-cycle phases
The EMBO Journal (2012)
-
Dot1 binding induces chromatin rearrangements by histone methylation-dependent and -independent mechanisms
Epigenetics & Chromatin (2011)