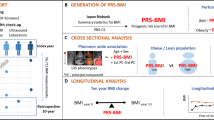
Abstract
Background:
Central obesity is a rising epidemic, and often occurs in parallel with dyslipidemia. Furthermore, enhancement of ectopic fat deposition has been observed in both human studies and animal models of altered lipidemic control. Though APOA1/C3/A4/A5 genetic polymorphisms are associated with dyslipidemia, their effect on central obesity is less known.
Method:
The anthropometric and metabolic parameters were taken from obese (body mass index (BMI) ⩾25 kg m−2) and non-obese healthy (BMI <25) Taiwanese patients at the initiation weight-loss intervention and 6 months later. The effects of APOA1/C3/A4/A5 genetic polymorphisms were analyzed cross-sectionally and longitudinally. Gender contributions were specifically examined.
Patients:
Three hundred and ninety-eight participants (obese n=262; non-obese healthy n=136) were recruited in total, and 130 obese patients underwent weight-loss treatments.
Results:
APOA5 rs662799 minor allele carriage was associated with unfavorable metabolic profiles in obese but not non-obese individuals at baseline. Further analysis identified gender–genotype interactions in waist-hip ratio (WHR), and that one rs662799 minor allele increased 0.032 WHR unit in obese males as analyzed by linear regression adjusted for age, BMI and plasma triglyceride (TG) (95% confidence interval (CI)=0.014–0.050, P=0.001). The rs662799-associated WHR elevation resulted in increased frequency of central obesity (WHR ⩾1.0) in rs662799 carrying obese males as analyzed by binary logistic regression adjusted for age, BMI and plasma TG (odds ratio=6.52, 95% CI=1.87–22.73, P=0.003). In contrast, APOA5 rs662799 and central obesity were no longer correlated 6 months into weight-loss treatments, owing to significant WHR reductions in male rs662799 minor allele carriers (P=0.001). Meanwhile, hypertriglyceridemia was more prevalent in both male and female obese rs662799 minor allele carriers at baseline (males, P=0.034, females, P=0.007).
Conclusion:
This study highlights the gender-specific and weight-sensitive effects of APOA5 rs662799 on central obesity in Taiwanese individuals, and that these effects are dyslipidemia-independent and weight-loss responsive.
Similar content being viewed by others
Introduction
Obesity has reached a globally epidemic level, and is forecasted to affect 800 million individuals by 2015.1, 2 Prolonged obesity increases risks of developing metabolic syndrome (MetS), type 2 diabetes and cardiovascular diseases.2 In particular, central obesity is considered more pathogenic than peripheral obesity, and often occurs in parallel with dyslipidemia and insulin resistance.3, 4 Central obesity can be defined by elevated waist-hip ratio (WHR) or waist circumference (WC), which has been designated to be the prerequisite factor for the MetS criteria by the International Diabetes Federation.5 Therefore, active weight-loss through weight-loss surgery or planned diet regime in centrally obese patients can improve their glycemic and lipidemic controls.6, 7 Interestingly, alterations in lipidemic control and ectopic fat deposition have been observed concurrently in patients during viral infections and associated treatments.8, 9 Moreover, mice models deficient in controlling lipid or glucose metabolism have developed lipodystrophic or obesogenic phenotypes.10, 11
Several genetic polymorphisms situated in the APOA1/C3/A4/A5 gene cluster are associated with dyslipidemia, insulin resistance and MetS. APOA1/C3/A4/A5 transcribes for apolipoproteins (apos) A1, C3, A4 and A5,12 which participate in plasma triglyceride (TG) and cholesterol metabolism. ApoA1 is the structural protein of high-density lipoprotein (HDL), which is involved in reverse cholesterol transport.13 Genetic polymorphisms in APOA1 are associated with fasting and postprandial plasma lipids, and responses to medication for dyslipidemia.14, 15 The promoter region-situated rs670 on APOA1 is correlated with HDL-cholesterol (HDL-C) levels, MetS and type 2 diabetes.16, 17, 18 ApoC3 inhibits plasma lipoprotein lipase activity, and is found on TG-rich lipoproteins and HDL.19 The single-nucleotide polymorphisms (SNPs) within APOC3 coding or promoter regions are associated with the altered TG metabolism.20, 21 Two SNPs situated in the insulin-responsive element of APOC3, rs2854116 and rs2854117 are known to be associated with postprandial plasma TG and fatty liver.20, 22 The function of apoA4 is less known, but in vitro studies suggest its participation in lipoprotein lipase activity and lecithin-cholesterol acyltransferase activity.23, 24 APOA4 rs675, a missense SNP that alters threonine347 to serine, is associated with response to fenofibrate treatment.25 ApoA5 regulates TG metabolism and is proposed to interact with lipoprotein lipase activity.26, 27 Thus, mutations and SNPs on APOA5, especially rs662799, rs3135506 and the Chinese-predominant rs2075291, are associated with dyslipidemia and MetS.12, 28, 29
The pathogenesis of metabolic conditions including central obesity, dyslipidemia and impaired glucose control are gender dependent.30 Central obesity is gender dependent, in that men are more susceptible to abdominal fat deposition than premenopausal women.31, 32 The complications of central obesity are also gender dependent, as it predisposes the highest risk in hypertriglycedemia in men, but poses more risk of IR in females.33 Gender-specific effects are also observed in genetic predispositions to central obesity, as genome-wide screening identified adipogenesis-associated SNPs favoring central obesity in women.34, 35 Reflectively, the risks for dyslipidemia predisposed by SNPs are gender specific.16, 27, 36, 37 APOA1 rs670-associated increase in poly-unsaturated fatty acid-induced HDL-C was observed only in females.16 APOC3 rs2854117 is correlated strongly with hypertriglyceridemia in men, whereas it is correlated with increased insulin resistance in women.36 Rs662799 and rs3135506 on APOA5 are associated with hypertriglyceridemia in men with greater effects,27 but associated with lower HDL-C levels only in women.37
Though the effect of APOA1/C3/A4/A5 gene cluster polymorphisms on dyslipidemia has been studied in great detail, the contributions these polymorphisms in central obesity are less known, especially in Asian populations. Furthermore, the effect of weight-loss in obese patients carrying APOA1/C3/A4/A5 SNPs in improving associated metabolic factors is not yet established. This study analyzes the contributions of APOA1/C3/A4/A5 SNPs to obesity-associated anthropometric and metabolic parameters in Taiwanese obese (BMI ⩾25) patients and non-obese healthy (BMI <25) participants at baseline, and again in the obese patients 6 months after they have initiated weight-loss intervention. As gender has a vital role in the pathogenesis and pathophysiology of central obesity, the gender-specific contributions are examined.
Subjects and methods
Patients and treatments
Taiwanese non-obese healthy (BMI <25) volunteers and obese Taiwanese patients (BMI⩾25) visiting the Weight Management Clinic and at National Cheng Kung University Hospital (NCKUH), Tainan, Taiwan from 2007 to 2010 were recruited to participate in this study. Exclusion criteria included current smokers, high alcohol consumers (more than two drinks daily), diagnosis of inflammatory disease or cancer, and use of hormone replacement therapy. In total, 262 obese patients (112 males and 150 females) and 136 non-obese participants (50 males and 86 females) were enrolled. Of the recruited obese patients, 55 males and 75 females underwent weight-loss intervention and returned for regular visits over a 6-month period (study endpoint). The weight-loss intervention methods were balanced low-calorie diet (500 Kcal deficits per day) alone (18 males and 32 females),38 diet with 120 mg Orlistat (Roche, St Louis, MO, USA) t.i.d (6 males and 4 females),38 diet with Sibutramine (Abbot, Abbot Park, IL, USA) 10 or 15 mg daily (19 males and 25 females)38 and bariatric surgery (mini-gastric bypass/sleeve, 12 males and 14 females). Patient compliance of the diet program was supervised by trained professional dieticians reviewing patient’s nutritional diary during regular clinic visits of every 3 weeks. The study received approval from the local institutional review board, and all patients gave informed written consent.
Anthropometric and laboratory measurements
At 0 month and after 6 month of weight-loss intervention, patients underwent physical examination by trained personnel who took anthropometric measurements, including body height, bodyweight, WC and hip circumference (HC). Body mass index (BMI) was calculated as bodyweight (kg)/(body height (m)).2 Whole-venous blood was collected after overnight fasting, and at 2 h post 75 g oral glucose challenge. WHR was calculated as WC (cm)/HC (cm). Plasma levels of total TG, total cholesterol (TC), HDL-C, low-density lipoprotein cholesterol, and total glucose were measured with an automated instrument biochemically (Roche Modulator DP, Roche). Fasting serum insulin was measured by radioimmunoassay (Coat-A-Count Insulin in vitro Diagnostic Test Kit, Siemens, Los Angeles, CA, USA). Homeostasis model assessment for insulin resistance was calculated as (fasting plasma glucose (mmol l−1) × fasting serum insulin (μU ml−1))/22.5. The MetS definition criteria used in this study comply with that proposed by the International Diabetes Federation with appropriate WC adjustments for Asians.39 In short, MetS is diagnosed in the presence of elevated WC (WC ⩾80 cm in females and ⩾90 cm in males) plus two of the following: (i) hypertriglycedemia (hyper-TG; TG ⩾1.7 mmol/l); (ii) low HDL-C (HDL-C <1.3 mmol/l in females and <1.0 mmol/l in males); (iii) impaired fasting glucose (fasting glucose ⩾5.6 mmol/l); and (iv) hypertension (systolic blood pressure ⩾130 mm Hg or diastolic blood pressure ⩾90 mm Hg). Impairment in glucose tolerance was defined plasma glucose 2 h after oral glucose challenge (PC-2 h) ⩾7.8 mmol l−1 without concurrent impaired fasting glucose. The cutoff values used for elevated WHR were ⩾1.0 in males and ⩾0.9 in females.
DNA extraction, SNP selection and genotyping
DNA was extracted from buffy coats by a standardized procedure using a commercial kit (GeneElute blood genomic DNA kit, Sigma-Aldrich, St Louis, MO, USA). Concentrations were determined using NanoDrop 2000 (Thermo Scientific, Wilmington, DE, USA). Seven SNPs on APOA1/C3/A4/A5 gene cluster were selected: APOA1 rs670, APOC3 rs2854116 and rs2854117, APOA4 rs675 and APOA5 rs662799, rs3135506 and rs2075291. The phenotypes of the selected SNPs is summarized in Supplementary Table 2. Next, the SNP genotypes were determined using commercial primer and probes from Applied Biosystems (ABI, Foster City, CA, USA) (APOA4 rs675; APOA5 rs662799, rs3135506 and rs2075291) and TIB MOBIOL (Berlin, Germany) (APOA1 rs670; APOC3 rs2854116 and rs2854117). Finally, fluorescence data were collected by a Step-One-Plus Sequence Detection System (ABI, Foster City, CA, USA) or LightCycler 480 (Roche).
Statistical analysis
Haploview40 was used for the analysis of SNP linkage disequilibrium, and tests with LOD values ⩾2 and D′ values >0.80 were considered as presence of significant linkage. The Hardy–Weinberg Equilibrium of SNPs was determined using Fisher’s exact test, and the distribution of SNP genotype among participant groups were tested by the χ2-test. Differences in continuous metabolic parameters of participant groups dichotomized by SNP genotypes were tested by one-way analysis of variance. The correlations between SNP minor alleles and continuous parameters were analyzed by Pearson’s correlation, and those with a statistical significance were further analyzed by linear regression. Correlations of SNP genotypes with anthropometric and metabolic conditions were demonstrated by the χ2-test, and those with a statistical significance were further analyzed by binary logistic regression. Potential confounding from factors such as age and BMI were adjusted by multivariate analysis. Genotype–gender interactions were tested using the general linear model univariate analysis. The significance of improvements in metabolic parameters from baseline to study endpoint was analyzed by the paired t-test, and the differences of improvements in parameters between groups were tested by Student’s t-test. Statistical analysis was performed using SPSS 13. In all cases, P-values <0.05 were considered statistically significant.
Results
Demographic data of enrolled obese and non-obese patients
Two hundred and sixty-two obese (BMI ⩾25) patients and 136 non-obese (BMI <25) healthy volunteers were recruited from the Weight Management Clinic and routine health check-up in NCKUH from 2007 to 2010. Table 1 summarizes the characteristics of the recruited patients, indicating that the obese patients had significant higher anthropometric parameters as bodyweight, BMI, body fat, WC, HC and WHR, and less favorable metabolic parameters as higher TG, TC, low-density lipoprotein cholesterol, plasma glucose, fasting insulin, homeostasis model assessment for insulin resistance and lower HDL-C than non-obese healthy volunteers (Table 1A, all P-values <0.001). Analyzed based on the International Diabetes Federation guidelines, our results indicated that recruited obese patients had higher proportions of central obesity (as elevated WC or elevated WHR), dyslipidemia, impaired glycemic control, hypertension and MetS than non-obese individuals (Table 1B, all P-values <0.001). The gender distribution of obese and non-obese participants were comparable (Table 1B, P=0.282). Of the seven genotyped SNPs, APOA4 rs675 and APOA5 rs3135506 were homozygotes of the major allele in all recruited patients. The genotype distributions of APOA1 rs670, APOC3 rs2854116, APOC3 rs2854117, APOA5 rs662799 and APOA5 rs2075291 were comparable in obese and non-obese individuals (Supplementary Table 2). Supplementary Figure 1 shows the linkage disequilibrium plot of APOA1 rs670, APOC3 rs2854116, APOC3 rs2854117, APOA5 rs662799 and APOA5 rs2075291. Strong linkage disequilibrium (LOD >2) was observed among APOA1 rs670, APOC3 rs2854116 and APOC3 rs2854117, and between APOA5 rs667299 and rs2075291 (Supplementary Figure 1). Of note, the two selected SNPs on APOA5 displayed no (rs662799 vs rs670/rs2854117, rs2075291 vs rs2854116/rs2854117, LOD<2, D′<0.80, white) or insignificant (rs662799 vs rs2854116, rs2075291 vs rs670, LOD>2, D′ <0.80, pink) linkage with SNPs on APOA1 and APOC3 (Supplementary Figure 1). The five SNPs were analyzed against anthropometric and metabolic parameters. APOA1 rs670, APOC3 rs2854116, APOC3 rs2854117 and APOA5 rs2075291 showed no or minimal association with tested parameters in obese or non-obese participants. In contrast, APOA5 rs662799 showed significant associations with metabolic profiles in obese patients (Table 2).
APOA5 rs662799 correlated with unfavorable metabolic profiles in obese patients
APOA5 rs662799 showed significant associations with TG and TC in obese patients (Table 2): the mean values for TG and TC were significantly different among APOA5 rs662799 genotype carriers (P<0.001 and P=0.044, respectively), but only the association with TG showed linearity (P<0.001). Moreover, significantly higher proportions of obese APOA5 rs662799 C allele carriers (T/C+C/C) were centrally obese than non-C allele carriers (T/T) in the sense of elevated WHR (32.38 vs 19.67%, P=0.033) (Table 3). The C allele carriers were more likely to have hyper-TG, low HDL-C and MetS than non-C carriers (hyper-TG: 37.14 vs 16.39%, P<0.001; low HDL-C: 62.86 vs 45.08%, P=0.011; and MetS: 52.38 vs 35.25%, P=0.015) (Table 3). Though APOA5 rs662799 was associated with hypertriglyceridemia in non-obese participants, the number of patients with the metabolic condition was minimal (Supplementary Table 3).
Gender-dependent effect on associations of APOA5 rs662799 minor allele carriage with central obesity and MetS in obese patients
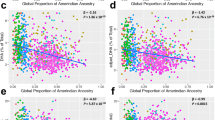
Upon further analysis, we found that gender contributed to APOA5 rs662799’s association with metabolic parameters in obese patients (Figure 1). Gender–genotype interactions were observed in WHR (P=0.004, Figure 1a) and TG (P<0.001, Figure 1b) but not BMI (Figure 1c) and HDL-C (Figure 1d) of obese patients. The mean values of WHR (P<0.001, Figure 1a) and TG (P<0.001, Figure 1b) were significantly different among male APOA5 rs662799 genotype carriers. This was reflected by the linear association of APOA5 rs662799 C allele with WHR and TG in obese males (WHR P<0.001, and TG P<0.001, Table 4). One APOA5 rs662799 C allele conferred to increase of 0.032 unit of WHR (95% CI=0.014–0.050, P=0.001) and 0.905 mmol l−1 TG (95% CI=0.445–1.365, P<0.001) in obese males after appropriate adjustments (Table 4). The correlations of APOA5 rs662799 C allele with WHR and TG were independent, as the statistical significance sustained after adjustments for TG and WHR, respectively (Table 4). In contrast, APOA5 rs662799 C allele did not show significant linear associations with WHR or TG in obese females (Table 4).
Test of gender- APOA5 rs662799 genotype interaction in WHR, plasma TG, BMI and plasma HDL-C in obese patients. The mean±s.d. values of analyzed parameter (WHR (a), TG (b), BMI (c) and HDL-c (d)) are shown under the designated APOA5 rs662799 genotype in a gender-specific manner. The cutoff values for elevated WHR (⩾1.0 unit for males and ⩾0.9 unit for females), hyper-TG (TG ⩾1.7 mmol l−1), low HDL-C (HDL-C <1.0 mmol−l for males and <1.3 mmol−l for females) are shown as dashed line in figures. The number of individuals with the anthropometric and metabolic conditions is shown under the designated groups. The differences in mean±s.d. values were tested by one-way analysis of variance (ANOVA). The genotype–gender interactions were tested by general linear model univariate analysis.
Gender-stratification of obese patients revealed that the overall association of APOA5 rs662799 C allele carriage with increased frequencies of central obesity and MetS was contributed mainly by obese males (Table 5). Obese male C allele carriers had increased odds of elevated WHR by 6.52-fold (95% CI=1.87–22.73, P=0.003) as compared with non-C allele carriers after adjustments for age, BMI and TG (Table 5). Though the frequency of MetS increased in obese males APOA5 rs662799 C allele carriers as compared with non-C allele carriers, the effect was diminished after additional adjustment for WHR (Table 5). In addition, the correlation of APOA5 rs662799 C allele with WHR and central obesity in obese males remained strong and significant after further adjustment for APOA5 rs2075291 (Supplementary Table 4).
On the other hand, the carriage of APOA5 rs662799 C allele in obese females did not statistically increase the occurrence of central obesity, MetS or low-HDLC after appropriate adjustments (Table 5). APOA5 rs662799 C allele carriage increased odds of hyper-TG in both obese males and obese females, however, after adjustment for age, BMI plus WHR only that in obese females remained statistically significant (odds ratio=5.63, 95% CI=1.89–16.78, P=0.002, Table 5). Similarly, additional adjustment for APOA5 rs2075291 did not diminish the correlation of APOA5 rs662799 C allele carriage with hyper-TG in obese females (Supplementary Table 4).
The unfavorable metabolic profiles contributed by APOA5 rs662799 in obese males at baseline were no longer present after significant weight-loss
One-hundred and thirty of the 262 obese patients initiated weight-loss intervention, and completed the return visit 6 months later, and their clinical data were analyzed longitudinally. The baseline BMI of APOA5 rs662799 C allele carriers and non-C allele carriers were comparable in obese male (P=0.530, Figure 2a) and obese female patients (P=0.310, Supplementary Figure 2A). Moreover, 6 months after weight-loss intervention, APOA5 rs662799 C and non-C allele carriers in both genders achieved significant improvement in BMI (P<0.001 and P=0.015 in males, Figure 2a; P<0.001 and P<0.001 in females, Supplementary Figure 2A). The number of patients treated with diet alone, diet with Orlistat (Roche), diet with Sibutramine (Abbot) or bariatric surgery were comparable (P=0.632) as shown in Figure 2a.
Obese males carrying APOA5 rs662799 C allele have significant improvements in BMI, WHR and TG at 6 months after weight-loss intervention. The improvements in BMI (a), WHR (b) and TG (c) are shown. The values shown under specified timepoints or above genotype subgroups were mean±s.d. Paired t-test was used for comparing 0 month and 6 months values in specified subgroups, while the Student’s t-test was used for comparing values between different groups.
Consistent with the baseline findings of the current study, baseline WHR of obese male APOA5 rs662799 C allele carriers was higher than that of non-C allele carriers (0.98±0.05 vs 0.93±0.05 unit, P=0.005, Figure 2b). Interestingly, significant improvements in WHR was achieved in APOA5 rs662799 C allele carriers (0.98±0.05 to 0.94±0.05, P=0.001, Figure 2b). On the contrary, the changes in WHR were not evident in APOA5 rs662799 non-C allele males (0.93±0.05 to 0.92±0.07, P=0.267). Of note, significant improvements of male APOA5 rs662799 C allele and non-C allele carriers were observed in both WC (P<0.001 and P=0.004) and HC (P<0.001 and P=0.002). The significant improvement in WHR at 6 months after weight-loss intervention resulted in the comparable WHR (P=0.240, Figure 2b) between male APOA5 rs662799 C allele and non-C allele carriers at study endpoint. In contrast, APOA5 rs662799 C allele carriage was not associated with difference in baseline or post-intervention WHR in females, and no significant improvement was observed in APOA5 rs662799 C allele carriers (Supplementary Figure 2B). Additionally, TG improvements were observed in both males (P<0.001, Figure 2c) and females (P=0.009 Supplementary Figure 2C).
Discussion
This study has analyzed the effects of APOA1/C3/A4/A5 dyslipidemia-associated SNPs on central obesity using cross-sectional and longitudinal methods. In particular, APOA5 rs662799 has been found to exhibit gender-specific associations with metabolic parameters in obese individuals. Obese males carrying APOA5 rs662799 minor variant had increased WHR and TG, and thus have increased odds in central obesity, hypertriglycedemia and MetS. Importantly, after comparable reductions in BMI, weight-loss intervention significantly improved the gender-specific impairments predisposed by APOA5 rs662799 polymorphism.
APOA5 rs662799’s effect on dyslipidemia has been widely studied, and its association with plasma lipid has exhibited gender-specificities.27, 37, 41 The present study confirmed APOA5 rs662799’s significant impact on TG and hypertriglycedemia in obese patients of both genders. One of the first relevant publications found that the effect of APOA5 rs662799 on increased plasma TG was significant in Caucasian males but not their female counterparts.27 Moreover, in both Turkish and German participants, APOA5 rs662799 minor variant was associated with isolated hypertriglycedemia in males, whereas in females accompanying lower HDL-C levels were observed.37, 41 This observation is echoed by our data, which showed APOA5 rs662799 minor allele linearly-correlated with TG only in obese males, but associated with low HDL-C accompanied hypertriglycedemia in females (Table 5).
Of particular interest, our results indicated APOA5 rs662799 minor variant associated with WHR and central obesity as reflected by greater WHR in obese males. The effect of APOA5 rs662799 on central obesity was evident only in obese males, but was not observed in obese females and non-obese participants. Interestingly, men’s higher tendency of adipose accumulation in the central region did not conceal the contribution of APOA5 rs662799 on central obesity. The findings were consistent after adjustments for TG, age and BMI, showing an independent nature of the association. According to a recent meta-analysis study, APOA5 rs662799 have displayed a persistent and cross-ethnic association with MetS.12 Correspondingly, our results indicated that APOA5 rs662799 was associated with MetS risk in obese males, particularly through the correlation with WHR and TG. This observation further emphasizes the importance of APOA5 rs662799, as central obesity is recognized as the precedent factor promoting the other components of MetS.
Two separate studies have found the association of WC with APOA5 rs3135506 in Hispanics,42 and BMI with APOA5 rs2075291 in Chinese.43 However, APOA5 rs3135506 genotype was found to be ubiquitous among our study group, while APOA5 rs2075291 was associated with increased TG but not BMI. Genotype–dietary fat interactions on APOA5 rs662799’s effect on central obesity has been reported, as the variant carriers consuming Mediterranean low-fat diet have increased WC than non-carriers.44 The pathogenic mechanism of APOA5 rs662799 in WHR is outside the scope of our study. Nevertheless, recent studies reported treatment of apoA5 in in vitro-differentiated human adipocytes have specific regulation on the amount of intracellular lipid content,45 and that APOA5-transgenic mice have increased inguinal fat pads.46 Of specific importance, a decrease in hepatic apoA5 was shown to accompany improvements in liver steatosis in individuals who underwent bariatric surgery.47 Furthermore, several reports illustrated the effect of apoA5 on intracellular lipid droplet in hepatoma cell lines,48 and that APOA5-transgenic mice have increased hepatic TG.49 As central obesity is associated with the exacerbation of liver steatosis in obese individuals,50 the results of ours and others stress the importance of weight control for APOA5 rs662799 variant carriers in maintaining a healthy profile.
The response of APOA5 rs662799-associated metabolic disorders to weight-loss intervention has not been thoroughly analyzed previously. Only two studies have focused on the post hocs of weight reduction in APOA5 rs662799 minor variant carriers.51, 52 Examination of obese males undergoing restricted diet intervention found APOA5 rs662799 minor variant carriers had comparable BMI and lipid parameters, yet significant greater reduction in BMI was achieved than their counterparts at 3 months after intervention.51 The other study found that female carriers of APOA5 rs662799 minor variants had a significantly higher baseline TG; 9 weeks after life-style intervention, the variant carriers achieved a greater reduction of TG than their control counterparts.52 During a longer regular visit period (6 months) achieved in this study, our results indicated weight loss can ameliorate the unfavorable metabolic profiles in risk carriers of APOA5 rs662799 polymorphisms. Moreover, although WHR of the risk variant carriers were higher than those of non-risk counterparts at baseline, the difference became insignificant after a comparable reduction in BMI after a 6-month period. Analysis of normal weight individuals further strengthened this finding, as APOA5 rs662799 showed no significance in correlation with metabolic parameters. In sum, this study demonstrates the importance of weight management in individuals carrying APOA5 rs662799 polymorphisms, as intervention can reduce the unfavorable metabolic profile predisposed by these SNPs.
In conclusion, this study conducted a systematic and through longitudinal analysis on the effects of the well-known SNPs on APOA1/C3/A4/A5 in gender-stratified obese patients undergoing weight-loss intervention. We showed that APOA5 rs662799 polymorphism (C allele carriers: T/C and C/C) was unfavorable in obese patients; however, this predicament was improved prominently by weight management. Despite our relatively small sample size, appropriate statistical adjustments demonstrated the consistency of our main findings. Moreover, baseline observations and preliminary analysis in normal weight volunteers supported the intervention outcome of the patients, despite the varied intervention methods among the patients. This pioneering study supports the generally accepted view of weight maintenance in maintaining a healthy metabolic profile. Results of this study provide further insight into the benefits of weight-loss intervention in Asian obese patients with central obesity and dyslipidemia.
References
Houtkooper RH, Auwerx J . Obesity: new life for antidiabetic drugs. Nature 2010; 466: 443–444.
Withrow D, Alter DA . The economic burden of obesity worldwide: a systematic review of the direct costs of obesity. Obes Rev 2010; 12: 131–141.
Despres JP, Lemieux I . Abdominal obesity and metabolic syndrome. Nature 2006; 444: 881–887.
Eckel RH, Kahn SE, Ferrannini E, Goldfine AB, Nathan DM, Schwartz MW et al. Obesity and type 2 diabetes: what can be unified and what needs to be individualized? Diabetes Care 2011; 34: 1424–1430.
Eckel RH, Alberti KG, Grundy SM, Zimmet PZ . The metabolic syndrome. Lancet 2010; 375: 181–183.
Knowler WC, Barrett-Connor E, Fowler SE, Hamman RF, Lachin JM, Walker EA et al. Reduction in the incidence of type 2 diabetes with lifestyle intervention or metformin. N Engl J Med 2002; 346: 393–403.
Sjostrom L, Lindroos AK, Peltonen M, Torgerson J, Bouchard C, Carlsson B et al. Lifestyle, diabetes, and cardiovascular risk factors 10 years after bariatric surgery. N Engl J Med 2004; 351: 2683–2693.
Carr A, Samaras K, Thorisdottir A, Kaufmann GR, Chisholm DJ, Cooper DA . Diagnosis, prediction, and natural course of HIV-1 protease-inhibitor-associated lipodystrophy, hyperlipidaemia, and diabetes mellitus: a cohort study. Lancet 1999; 353: 2093–2099.
Serfaty L, Andreani T, Giral P, Carbonell N, Chazouilleres O, Poupon R . Hepatitis C virus induced hypobetalipoproteinemia: a possible mechanism for steatosis in chronic hepatitis C. J Hepatol 2001; 34: 428–434.
Tsai YS, Tsai PJ, Jiang MJ, Chou TY, Pendse A, Kim HS et al. Decreased PPAR gamma expression compromises perigonadal-specific fat deposition and insulin sensitivity. Mol Endocrinol 2009; 23: 1787–1798.
Weinstock PH, Levak-Frank S, Hudgins LC, Radner H, Friedman JM, Zechner R et al. Lipoprotein lipase controls fatty acid entry into adipose tissue, but fat mass is preserved by endogenous synthesis in mice deficient in adipose tissue lipoprotein lipase. Proc Natl Acad Sci USA 1997; 94: 10261–10266.
Povel CM, Boer JM, Reiling E, Feskens EJ . Genetic variants and the metabolic syndrome: a systematic review. Obes Rev 2011; 12: 952–967.
Silva RA, Huang R, Morris J, Fang J, Gracheva EO, Ren G et al. Structure of apolipoprotein A-I in spherical high density lipoproteins of different sizes. Proc Natl Acad Sci USA 2008; 105: 12176–12181.
Marin C, Lopez-Miranda J, Gomez P, Paz E, Perez-Martinez P, Fuentes F et al. Effects of the human apolipoprotein A-I promoter G-A mutation on postprandial lipoprotein metabolism. Am J Clin Nutr 2002; 76: 319–325.
Smith JA, Arnett DK, Kelly RJ, Ordovas JM, Sun YV, Hopkins PN et al. The genetic architecture of fasting plasma triglyceride response to fenofibrate treatment. Eur J Hum Genet 2008; 16: 603–613.
Ordovas JM, Corella D, Cupples LA, Demissie S, Kelleher A, Coltell O et al. Polyunsaturated fatty acids modulate the effects of the APOA1 G-A polymorphism on HDL-cholesterol concentrations in a sex-specific manner: the Framingham Study. Am J Clin Nutr 2002; 75: 38–46.
Morcillo S, Cardona F, Rojo-Martinez G, Esteva I, Ruiz-de-Adana MS, Tinahones F et al. Association between MspI polymorphism of the APO AI gene and Type 2 diabetes mellitus. Diabet Med 2005; 22: 782–788.
Phillips CM, Goumidi L, Bertrais S, Field MR, McManus R, Hercberg S et al. Gene-nutrient interactions and gender may modulate the association between ApoA1 and ApoB gene polymorphisms and metabolic syndrome risk. Atherosclerosis 2011; 214: 408–414.
Ooi EM, Barrett PH, Chan DC, Watts GF . Apolipoprotein C-III: understanding an emerging cardiovascular risk factor. Clin Sci 2008; 114: 611–624.
Russo GT, Meigs JB, Cupples LA, Demissie S, Otvos JD, Wilson PW et al. Association of the Sst-I polymorphism at the APOC3 gene locus with variations in lipid levels, lipoprotein subclass profiles and coronary heart disease risk: the Framingham offspring study. Atherosclerosis 2001; 158: 173–181.
Lopez-Miranda J, Williams C, Lairon D . Dietary, physiological, genetic and pathological influences on postprandial lipid metabolism. Br J Nutr 2007; 98: 458–473.
Petersen KF, Dufour S, Hariri A, Nelson-Williams C, Foo JN, Zhang XM et al. Apolipoprotein C3 gene variants in nonalcoholic fatty liver disease. N Engl J Med 2010; 362: 1082–1089.
Goldberg IJ, Scheraldi CA, Yacoub LK, Saxena U, Bisgaier CL . Lipoprotein ApoC-II activation of lipoprotein lipase. Modulation by apolipoprotein A-IV. J Biol Chem 1990; 265: 4266–4272.
Steinmetz A, Utermann G . Activation of lecithin: cholesterol acyltransferase by human apolipoprotein A-IV. J Biol Chem 1985; 260: 2258–2264.
Feitosa MF, An P, Ordovas JM, Ketkar S, Hopkins PN, Straka RJ et al. Association of gene variants with lipid levels in response to fenofibrate is influenced by metabolic syndrome status. Atherosclerosis 2011; 215: 435–439.
Merkel M, Loeffler B, Kluger M, Fabig N, Geppert G, Pennacchio LA et al. Apolipoprotein AV accelerates plasma hydrolysis of triglyceride-rich lipoproteins by interaction with proteoglycan-bound lipoprotein lipase. J Biol Chem 2005; 280: 21553–21560.
Pennacchio LA, Olivier M, Hubacek JA, Cohen JC, Cox DR, Fruchart JC et al. An apolipoprotein influencing triglycerides in humans and mice revealed by comparative sequencing. Science 2001; 294: 169–173.
Moreno R, Perez-Jimenez F, Marin C, Moreno JA, Gomez P, Bellido C et al. A single nucleotide polymorphism of the apolipoprotein A-V gene -1131T>C modulates postprandial lipoprotein metabolism. Atherosclerosis 2006; 189: 163–168.
Kao JT, Wen HC, Chien KL, Hsu HC, Lin SW . A novel genetic variant in the apolipoprotein A5 gene is associated with hypertriglyceridemia. Hum Mol Genet 2003; 12: 2533–2539.
Iyer A, Kauter K, Brown L . Gender differences in metabolic syndrome: a key research issue? Endocr Metab Immune Disord Drug Targets 2011; 11: 182–188.
Blouin K, Boivin A, Tchernof A . Androgens and body fat distribution. J Steroid Biochem Mol Biol 2008; 108: 272–280.
Williams CM . Lipid metabolism in women. Proc Nutr Soc 2004; 63: 153–160.
Yeh WT, Chang HY, Yeh CJ, Tsai KS, Chen HJ, Pan WH . Do centrally obese Chinese with normal BMI have increased risk of metabolic disorders? Int J Obes 2005; 29: 818–825.
Heid IM, Jackson AU, Randall JC, Winkler TW, Qi L, Steinthorsdottir V et al. Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution. Nat Genet 2010; 42: 949–960.
Lindgren CM, Heid IM, Randall JC, Lamina C, Steinthorsdottir V, Qi L et al. Genome-wide association scan meta-analysis identifies three Loci influencing adiposity and fat distribution. PLoS Genet 2009; 5: e1000508.
Coban N, Onat A, Guclu-Geyik F, Komurcu-Bayrak E, Sansoy V, Hergenc G et al. Gender- and obesity-specific effect of apolipoprotein C3 gene (APOC3) -482C>T polymorphism on triglyceride concentration in Turkish adults. Clin Chem Lab Med 2012; 50: 285–292.
Komurcu-Bayrak E, Onat A, Poda M, Humphries SE, Palmen J, Guclu F et al. Gender-modulated impact of apolipoprotein A5 gene (APOA5) -1131T>C and c.56C>G polymorphisms on lipids, dyslipidemia and metabolic syndrome in Turkish adults. Clin Chem Lab Med 2008; 46: 778–784.
Wu CH, Kuo HC, Chang CS, Yu C . What extent of weight loss can benefit the health-related quality of life in motivated obese Chinese? Asia Pac J Clin Nutr 2009; 18: 423–432.
Chen HJ, Pan WH . Probable blind spot in the International Diabetes Federation definition of metabolic syndrome. Obesity 2007; 15: 1096–1100.
Barrett JC, Fry B, Maller J, Daly MJ . Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 2005; 21: 263–265.
Evans D, Buchwald A, Beil FU . The single nucleotide polymorphism -1131T>C in the apolipoprotein A5 (APOA5) gene is associated with elevated triglycerides in patients with hyperlipidemia. J Mol Med 2003; 81: 645–654.
Smith CE, Tucker KL, Lai CQ, Parnell LD, Lee YC, Ordovas JM . Apolipoprotein A5 and lipoprotein lipase interact to modulate anthropometric measures in Hispanics of Caribbean origin. Obesity 2010; 18: 327–332.
Wu CK, Chang YC, Hua SC, Wu HY, Lee WJ, Chiang FT et al. A triglyceride-raising APOA5 genetic variant is negatively associated with obesity and BMI in the Chinese population. Obesity 2010; 18: 1964–1968.
Sanchez-Moreno C, Ordovas JM, Smith CE, Baraza JC, Lee YC, Garaulet M . APOA5 gene variation interacts with dietary fat intake to modulate obesity and circulating triglycerides in a Mediterranean population. J Nutr 2011; 141: 380–385.
Zheng XY, Zhao SP, Yu BL, Wu CL, Liu L . Apolipoprotein A5 internalized by human adipocytes modulates cellular triglyceride content. Biol Chem 2012; 393: 161–167.
Pamir N, McMillen TS, Li YI, Lai CM, Wong H, LeBoeuf RC . Overexpression of apolipoprotein A5 in mice is not protective against body weight gain and aberrant glucose homeostasis. Metabolism 2009; 58: 560–567.
Ress C, Moschen AR, Sausgruber N, Tschoner A, Graziadei I, Weiss H et al. The role of apolipoprotein A5 in non-alcoholic fatty liver disease. Gut 2011; 60: 985–991.
Shu X, Ryan RO, Forte TM . Intracellular lipid droplet targeting by apolipoprotein A-V requires the carboxyl-terminal segment. J Lipid Res 2008; 49: 1670–1676.
Shu X, Nelbach L, Ryan RO, Forte TM . Apolipoprotein A-V associates with intrahepatic lipid droplets and influences triglyceride accumulation. Biochim Biophys Acta 2010; 1801: 605–608.
Cheung O, Kapoor A, Puri P, Sistrun S, Luketic VA, Sargeant CC et al. The impact of fat distribution on the severity of nonalcoholic fatty liver disease and metabolic syndrome. Hepatology 2007; 46: 1091–1100.
Aberle J, Evans D, Beil FU, Seedorf U . A polymorphism in the apolipoprotein A5 gene is associated with weight loss after short-term diet. Clin Genet 2005; 68: 152–154.
Suchanek P, Lorenzova A, Poledne R, Hubacek JA . Changes of plasma lipids during weight reduction in females depends on APOA5 variants. Ann Nutr Metab 2008; 53: 104–108.
Acknowledgements
We are grateful for Miss Ting-Yu Hou’s technical assistance and Miss Yu-Chen Shih for her administrative assistance. This study is supported by the National Science Council, Taiwan (grant no. NSC 100-2321-B-006 -015 -MY3, NSC-100-2320-B-006 -007 -MY3) and NCKUH (NCKUH-9801002).
Author Contributions:
M-C Hsu designed research, performed experiments, analyzed data and wrote the paper; C-S Chang and K-T Lee participated in research design, collection of clinical data and patient samples; Y-S Tsai and P-H Kuo participated in research design and collection of patient samples; H-Y Sun participated in research design and discussion; K-C Young participated in research design, discussion and paper-editing; C-H Wu contributed to research design, discussion and paper-editing, collection of clinical data and patient samples.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Nutrition & Diabetes website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Hsu, MC., Chang, CS., Lee, KT. et al. Central obesity in males affected by a dyslipidemia-associated genetic polymorphism on APOA1/C3/A4/A5 gene cluster. Nutr & Diabetes 3, e61 (2013). https://doi.org/10.1038/nutd.2013.2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nutd.2013.2
Keywords
This article is cited by
-
Gene-environment interactions due to quantile-specific heritability of triglyceride and VLDL concentrations
Scientific Reports (2020)
-
Phosphorylation of glycogen synthase kinase-3β in metabolically abnormal obesity affects immune stimulation-induced cytokine production
International Journal of Obesity (2015)
-
Triglyceride-raising APOA5 genetic variants are associated with obesity and non-HDL-C in Chinese children and adolescents
Lipids in Health and Disease (2014)