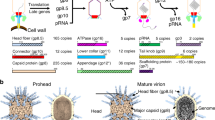
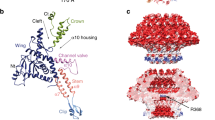
Abstract
Bacteriophage T4 and related viruses have a contractile tail that serves as an efficient mechanical device for infecting bacteria. A three-dimensional cryo-EM reconstruction of the mature T4 tail assembly at 15-Å resolution shows the hexagonal dome-shaped baseplate, the extended contractile sheath, the long tail fibers attached to the baseplate and the collar formed by six whiskers that interact with the long tail fibers. Comparison with the structure of the contracted tail shows that tail contraction is associated with a substantial rearrangement of the domains within the sheath protein and results in shortening of the sheath to about one-third of its original length. During contraction, the tail tube extends beneath the baseplate by about one-half of its total length and rotates by 345°, allowing it to cross the host's periplasmic space.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Leiman, P.G., Chipman, P.R., Kostyuchenko, V.A., Mesyanzhinov, V.V. & Rossmann, M.G. Three-dimensional rearrangement of proteins in the tail of bacteriophage T4 on infection of its host. Cell 118, 419–429 (2004).
Kanamaru, S. et al. Structure of the cell-puncturing device of bacteriophage T4. Nature 415, 553–557 (2002).
Letellier, L., Boulanger, P., de Frutos, M. & Jacquot, P. Channeling phage DNA through membranes: from in vivo to in vitro. Res. Microbiol. 154, 283–287 (2003).
Coombs, D.H. & Arisaka, F. in Molecular Biology of Bacteriophage T4 (ed. Karam, J.D.) 259–281 (American Society for Microbiology, Washington, DC, 1994).
Venyaminov, S.Y., Rodikova, L.P., Metlina, A.L. & Poglazov, B.F. Secondary structure change of bacteriophage T4 sheath protein during sheath contraction. J. Mol. Biol. 98, 657–664 (1975).
Moody, M.F. & Makowski, L. X-ray diffraction study of tail-tubes from bacteriophage T2L. J. Mol. Biol. 150, 217–244 (1981).
King, J. Assembly of the tail of bacteriophage T4. J. Mol. Biol. 32, 231–262 (1968).
Coombs, D.H. & Eiserling, F.A. Studies on the structure, protein composition and assembly of the neck of bacteriophage T4. J. Mol. Biol. 116, 375–405 (1977).
Cerritelli, M.E., Wall, J.S., Simon, M.N., Conway, J.F. & Steven, A.C. Stoichiometry and domainal organization of the long tail-fiber of bacteriophage T4: a hinged viral adhesin. J. Mol. Biol. 260, 767–780 (1996).
Makhov, A.M. et al. The short tail-fiber of bacteriophage T4: molecular structure and a mechanism for its conformational transition. Virology 194, 117–127 (1993).
Kikuchi, Y. & King, J. Genetic control of bacteriophage T4 baseplate morphogenesis. III. Formation of the central plug and overall assembly pathway. J. Mol. Biol. 99, 695–716 (1975).
Kostyuchenko, V.A. et al. The structure of bacteriophage T4 gene product 9: the trigger for tail contraction. Structure Fold Des. 7, 1213–1222 (1999).
Kellenberger, E., Stauffer, E., Haner, M., Lustig, A. & Karamata, D. Mechanism of the long tail-fiber deployment of bacteriophages T-even and its role in adsorption, infection and sedimentation. Biophys. Chem. 59, 41–59 (1996).
DeRosier, D.J. & Klug, A. Reconstruction of three dimensional structures from electron micrographs. Nature 217, 130–134 (1968).
Kostyuchenko, V.A. et al. Three-dimensional structure of bacteriophage T4 baseplate. Nat. Struct. Biol. 10, 688–693 (2003).
Smith, P.R., Aebi, U., Josephs, R. & Kessel, M. Studies of the structure of the T4 bacteriophage tail sheath. I. The recovery of three-dimensional structural information from the extended sheath. J. Mol. Biol. 106, 243–271 (1976).
Dewey, M.J., Wiberg, J.S. & Frankel, F.R. Genetic control of whisker antigen of bacteriophage T4D. J. Mol. Biol. 84, 625–634 (1974).
Efimov, V.P., Nepluev, I.V. & Mesyanzhinov, V.V. Bacteriophage T4 as a surface display vector. Virus Genes 10, 173–177 (1995).
Boudko, S.P. et al. Domain organization, folding and stability of bacteriophage T4 fibritin, a segmented coiled-coil protein. Eur. J. Biochem. 269, 833–841 (2002).
Strelkov, S.V., Tao, Y., Shneider, M.M., Mesyanzhinov, V.V. & Rossmann, M.G. Structure of bacteriophage T4 fibritin M: a troublesome packing arrangement. Acta Crystallogr. D Biol. Crystallogr. 54, 805–816 (1998).
Tao, Y., Strelkov, S.V., Mesyanzhinov, V.V. & Rossmann, M.G. Structure of bacteriophage T4 fibritin: a segmented coiled coil and the role of the C-terminal domain. Structure 5, 789–798 (1997).
Amos, L.A. & Klug, A. Three-dimensional image reconstructions of the contractile tail of T4 bacteriophage. J. Mol. Biol. 99, 51–64 (1975).
Arisaka, F., Takeda, S., Funane, K., Nishijima, N. & Ishii, S. Structural studies of the contractile tail sheath protein of bacteriophage T4. 2. Structural analyses of the tail sheath protein, gp18, by limited proteolysis, immunoblotting and immunoelectron microscopy. Biochemistry 29, 5057–5062 (1990).
Moody, M.F. Sheath of bacteriophage T4. 3. Contraction mechanism deduced from partially contracted sheaths. J. Mol. Biol. 80, 613–635 (1973).
Serysheva, I.I., Tourkin, A.I., Bartish, I.V. & Poglazov, B.F. GTPase activity of bacteriophage T4 sheath protein. J. Mol. Biol. 223, 23–25 (1992).
Conway, J.F. & Steven, A.C. Methods for reconstructing density maps of “single” particles from cryoelectron micrographs to subnanometer resolution. J. Struct. Biol. 128, 106–118 (1999).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Penczek, P., Radermacher, M. & Frank, J. Three-dimensional reconstruction of single particles embedded in ice. Ultramicroscopy 40, 33–53 (1992).
Wriggers, W., Milligan, R.A. & McCammon, J.A. Situs: a package for docking crystal structures into low-resolution maps from electron microscopy. J. Struct. Biol. 125, 185–195 (1999).
Acknowledgements
We thank S. Wilder and C. Towell for help in the preparation of the manuscript. The work was supported by a US National Science Foundation grant to M.G.R., a Howard Hughes Medical Institute grant and a Russian Fund for Basic Research grant to V.V.M., a Human Frontiers Science Program grant to M.G.R., F.A. and V.V.M., a Keck Foundation grant to M.G.R. for the purchase of the Philips CM300 field emission gun microscope and a reinvestment grant from Purdue University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kostyuchenko, V., Chipman, P., Leiman, P. et al. The tail structure of bacteriophage T4 and its mechanism of contraction. Nat Struct Mol Biol 12, 810–813 (2005). https://doi.org/10.1038/nsmb975
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb975
This article is cited by
-
Substantiation of propitious “Enzybiotic” from two novel bacteriophages isolated from a wastewater treatment plant in Qatar
Scientific Reports (2022)
-
Mechanics of tubular helical assemblies: ensemble response to axial compression and extension
Acta Mechanica Sinica (2021)
-
Biogenesis and structure of a type VI secretion baseplate
Nature Microbiology (2018)
-
Targeting mechanisms of tailed bacteriophages
Nature Reviews Microbiology (2018)
-
Recent Advancements in 3-D Structure Determination of Bacteriophages: from Negative Stain to CryoEM
Journal of the Indian Institute of Science (2018)