Abstract
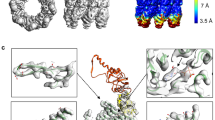
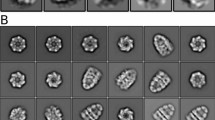
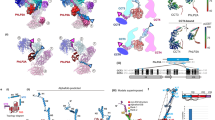
The chaperonin GroEL assists protein folding through ATP-dependent, cooperative movements that alternately create folding chambers in its two rings. The substitution E461K at the interface between these two rings causes temperature-sensitive, defective protein folding in Escherichia coli. To understand the molecular defect, we have examined the mutant chaperonin by cryo-EM. The normal out-of-register alignment of contacts between subunits of opposing wild-type rings is changed in E461K to an in-register one. This is associated with loss of cooperativity in ATP binding and hydrolysis. Consistent with the loss of negative cooperativity between rings, the cochaperonin GroES binds simultaneously to both E461K rings. These GroES-bound structures were unstable at higher temperature, dissociating into complexes of single E461K rings associated with GroES. Lacking the allosteric signal from the opposite ring, these complexes cannot release their GroES and become trapped, dead-end states.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sigler, P.B. et al. Structure and function in GroEL-mediated protein folding. Annu. Rev. Biochem. 67, 581–608 (1998).
Brinker, A. et al. Dual function of protein confinement in chaperonin-assisted protein folding. Cell 107, 223–233 (2001).
Saibil, H.R. & Ranson, N.A. The chaperonin folding machine. Trends Biochem. Sci. 27, 627–632 (2002).
Todd, M.J., Lorimer, G.H. & Thirumalai, D. Chaperonin-facilitated protein folding: optimization of rate and yield by an iterative annealing mechanism. Proc. Natl. Acad. Sci. USA 93, 4030–4035 (1996).
Rye, H.S. et al. GroEL-GroES cycling: ATP and nonnative polypeptide direct alternation of folding-active rings. Cell 97, 325–338 (1999).
Gray, T.E. & Fersht, A.R. Cooperativity in ATP hydrolysis by GroEL is increased by GroES. FEBS Lett. 292, 254–258 (1991).
Yifrach, O. & Horovitz, A. Nested cooperativity in the ATPase activity of the oligomeric chaperonin GroEL. Biochemistry 34, 5303–5308 (1995).
Burston, S.G., Ranson, N.A. & Clarke, A.R. The origins and consequences of asymmetry in the chaperonin reaction cycle. J. Mol. Biol. 249, 138–152 (1995).
Roseman, A.M., Chen, S., White, H., Braig, K. & Saibil, H.R. The chaperonin ATPase cycle: mechanism of allosteric switching and movements of substrate-binding domains in GroEL. Cell 87, 241–251 (1996).
Rye, H.S. et al. Distinct actions of cis and trans ATP within the double ring of the chaperonin GroEL. Nature 388, 792–798 (1997).
Ranson, N.A. et al. ATP-bound states of GroEL captured by cryo-electron microscopy. Cell 107, 869–879 (2001).
Ditzel, L. et al. Crystal structure of the thermosome, the archaeal chaperonin and homolog of CCT. Cell 93, 125–138 (1998).
Horwich, A.L., Low, K.B., Fenton, W.A., Hirshfield, I.N. & Furtak, K. Folding in vivo of bacterial cytoplasmic proteins: role of GroEL. Cell 74, 909–917 (1993).
Fenton, W.A., Kashi, Y., Furtak, K. & Horwich, A.L. Residues in chaperonin GroEL required for polypeptide binding and release. Nature 371, 614–661 (1994).
Sot, B., Galan, A., Valpuesta, J.M., Bertrand, S. & Muga, A. Salt bridges at the inter-ring interface regulate the thermostat of GroEL. J. Biol. Chem. 277, 34024–34029 (2002).
Yifrach, O. & Horovitz, A. Two lines of allosteric communication in the oligomeric chaperonin GroEL are revealed by the single mutation Arg196→Ala. J. Mol. Biol. 243, 397–401 (1994).
Weissman, J.S. et al. Mechanism of GroEL action: productive release of polypeptide from a sequestered position under GroES. Cell 83, 577–587 (1995).
Ma, J., Sigler, P.B., Xu, Z. & Karplus, M. A dynamic model for the allosteric mechanism of GroEL. J. Mol. Biol. 302, 303–313 (2000).
Kafri, G. & Horovitz, A. Transient kinetic analysis of ATP-induced allosteric transitions in the eukaryotic chaperonin containing TCP-1. J. Mol. Biol. 326, 981–987 (2003).
Braig, K. et al. The crystal structure of the bacterial chaperonin GroEL at 2.8 Å. Nature 371, 578–586 (1994).
Horovitz, A., Bochkareva, E.S., Kovalenko, O. & Girshovich, A.S. Mutation Ala2→Ser destabilizes intersubunit interactions in the molecular chaperone GroEL. J. Mol. Biol. 231, 58–64 (1993).
Ranson, N.A., Dunster, N.J., Burston, S.G. & Clarke, A.R. Chaperonins can catalyse the reversal of early aggregation steps when a protein misfolds. J. Mol. Biol. 250, 581–586 (1995).
Crowther, R.A., Henderson, R. & Smith, J.M. MRC image processing programs. J. Struct. Biol. 116, 9–16 (1996).
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Braig, K., Adams, P.D. & Brunger, A.T. Conformational variability in the refined structure of the chaperonin GroEL at 2.8 Å resolution. Nat. Struct. Biol. 2, 1083–1094 (1995).
Brunger, A.T. & Karplus, M. Polar hydrogen positions in proteins—empirical energy placement and neutron diffraction comparison. Proteins 4, 148–156 (1988).
Brooks B.R. et al. CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J. Comp. Chem. 4, 187–217 (1983).
MacKerell, A.D. Jr. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Acknowledgements
We thank R. Westlake, D. Houldershaw and S. Terrill for computing support, W. Fenton and M. Karplus for advice and discussion, M. Jaffer and W. Williams for help with particle counting, and The Wellcome Trust and the Howard Hughes Medical Institute for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Sewell, B., Best, R., Chen, S. et al. A mutant chaperonin with rearranged inter-ring electrostatic contacts and temperature-sensitive dissociation. Nat Struct Mol Biol 11, 1128–1133 (2004). https://doi.org/10.1038/nsmb844
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb844
This article is cited by
-
Structural basis for active single and double ring complexes in human mitochondrial Hsp60-Hsp10 chaperonin
Nature Communications (2020)
-
Towards single biomolecule handling and characterization by MEMS
Analytical and Bioanalytical Chemistry (2008)
-
Essential function of the built-in lid in the allosteric regulation of eukaryotic and archaeal chaperonins
Nature Structural & Molecular Biology (2007)
-
Markov propagation of allosteric effects in biomolecular systems: application to GroEL–GroES
Molecular Systems Biology (2006)