Abstract
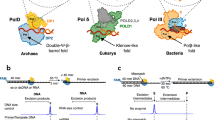
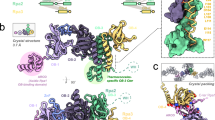
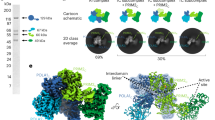
Genome replication generally requires primases, which synthesize an initial oligonucleotide primer, and DNA polymerases, which elongate the primer. Primase and DNA polymerase activities are combined, however, in newly identified replicases from archaeal plasmids, such as pRN1 from Sulfolobus islandicus. Here we present a structure-function analysis of the pRN1 primase-polymerase (prim-pol) domain. The crystal structure shows a central depression lined by conserved residues. Mutations on one side of the depression reduce DNA affinity. On the opposite side of the depression cluster three acidic residues and a histidine, which are required for primase and DNA polymerase activity. One acidic residue binds a manganese ion, suggestive of a metal-dependent catalytic mechanism. The structure does not show any similarity to DNA polymerases, but is distantly related to archaeal and eukaryotic primases, with corresponding active-site residues. We propose that archaeal and eukaryotic primases and the prim-pol domain have a common evolutionary ancestor, a bifunctional replicase for small DNA genomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kornberg, A. & Baker, T. DNA Replication (W.H. Freeman, New York, USA, 1991).
Frick, D.N. & Richardson, C.C. DNA primases. Annu. Rev. Biochem. 70, 39–80 (2001).
Augustin, M.A., Huber, R. & Kaiser, J.T. Crystal structure of a DNA-dependent RNA polymerase (DNA primase). Nat. Struct. Biol. 8, 57–61 (2001).
Keck, J.L., Roche, D.D., Lynch, A.S. & Berger, J.M. Structure of the RNA polymerase domain of E. coli primase. Science 287, 2482–2486 (2000).
Hubscher, U., Maga, G. & Spadari, S. Eukaryotic DNA polymerases. Annu. Rev. Biochem. 71, 133–163 (2002).
Steitz, T.A. DNA polymerases: structural diversity and common mechanisms. J. Biol. Chem. 274, 17395–81739 (1999).
Joyce, C.M. & Steitz, T.A. Function and structure relationships in DNA polymerases. Annu. Rev. Biochem. 63, 777–822 (1994).
Lipps, G., Rother, S., Hart, C. & Krauss, G. A novel type of replicative enzyme harbouring ATPase, primase and DNA polymerase activity. EMBO J. 22, 2516–2525 (2003).
De Guzman, R.N., Wu, Z.R., Stalling, C.C., Pappalardo, L., Borer, P.N. & Summers, M.F. Structure of the HIV-1 nucleocapsid protein bound to the SL3 Ψ-RNA recognition element. Science 279, 384–388 (1998).
Steitz, T.A. A mechanism for all polymerases. Nature 391, 231–223 (1998).
Sawaya, M.R., Pelletier, H., Kumar, A., Wilson, S.H. & Kraut, J. Crystal structure of rat DNA polymerase β: evidence for a common polymerase mechanism. Science 264, 1930–1937 (1994).
Davies, J.F., Almassy, R.J., Hostomska, Z., Ferre, R.A. & Hostomsky, Z. 2.3 A crystal structure of the catalytic domain of DNA polymerase beta. Cell 76, 1123–1133 (1994).
Doublie, S., Tabor, S., Long, A.M., Richardson, C.C. & Ellenberger, T. Crystal structure of a bacteriophage T7 DNA replication complex at 2.2 Å resolution. Nature 391, 251–258 (1998).
Sawaya, M.R., Prasad, R., Wilson, S.H., Kraut, J. & Pelletier, H. Crystal structures of human DNA polymerase β complexed with gapped and nicked DNA: evidence for an induced fit mechanism. Biochemistry 36, 11205–11215 (1997).
Kato, M., Ito, T., Wagner, G., Richardson, C.C. & Ellenberger, T. Modular architecture of the bacteriophage T7 primase couples RNA primer synthesis to DNA synthesis. Mol. Cell 11, 1349–1360 (2003).
Holm, L. & Sander, C. DALI: a network tool for protein structure comparison. Trends Biochem. Sci. 20, 478–480 (1995).
Koonin, E.V., Wolf, Y.I., Kondrashov, A.S., Aravind, L. Bacterial homologs of the small subunit of eukaryotic DNA primase. J. Mol. Microbiol. Biotechnol. 2, 509–512 (2000).
Forterre, P. The origin of DNA genomes and DNA replication proteins. Curr. Opin. Microbiol. 5, 525–532 (2002).
Liu, L. et al. The archaeal DNA primase: biochemical characterization of the p41-p46 complex from Pyrococcus furiosus. J. Biol. Chem. 276, 45484–45490 (2001).
Bocquier, A.A. et al. Archaeal primase: bridging the gap between RNA and DNA polymerases. Curr. Biol. 11, 452–456 (2001).
Meinhart, A., Silberzahn, T. & Cramer, P. The mRNA transcription/processing factor Ssu72 is a potential tyrosine phosphatase. J. Biol. Chem. 278, 15917–15921 (2003).
Budisa, N. et al. High-level biosynthetic substitution of methionine in proteins by its analogs 2-aminohexanoic acid, selenomethionine, telluromethionine and ethionine in Escherichia coli. Eur. J. Biochem. 230, 788–796 (1995).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1996).
Matthews, B.W. Solvent content of protein crystals. J. Mol. Biol. 33, 491–495 (1968).
Terwilliger, T.C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Carson, M. Ribbons. Methods Enzymol. 277, 493–505 (1997).
Acknowledgements
We thank A. Schmidt for excellent technical assistance, K. Zeth for data collection at ESRF, and C. Schulze-Briese and the staff of beamline X06SA of the Swiss Light Source for help. P.C. is supported by the Deutsche Forschungsgemeinschaft, the EMBO Young Investigator Programme and the Fonds der chemischen Industrie. G.L. is supported by the Deutsche Forschungsgemeinschaft. G.L. dedicates this work to G. Krauss, University of Bayreuth, on the occasion of his 60th birthday.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Lipps, G., Weinzierl, A., von Scheven, G. et al. Structure of a bifunctional DNA primase-polymerase. Nat Struct Mol Biol 11, 157–162 (2004). https://doi.org/10.1038/nsmb723
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb723
This article is cited by
-
Isolation and characterization of bacteriophage NTR1 infectious for Nocardia transvalensis and other Nocardia species
Virus Genes (2019)
-
Complete genomic sequence of bacteriophage P23: a novel Vibrio phage isolated from the Yellow Sea, China
Virus Genes (2019)
-
Primer synthesis by a eukaryotic-like archaeal primase is independent of its Fe-S cluster
Nature Communications (2017)
-
The Polyphyletic Origins of Primase–Helicase Bifunctional Proteins
Journal of Molecular Evolution (2017)
-
TruePrime is a novel method for whole-genome amplification from single cells based on TthPrimPol
Nature Communications (2016)