Abstract
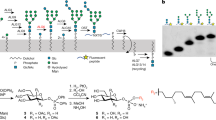
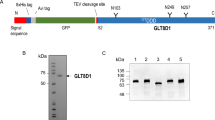
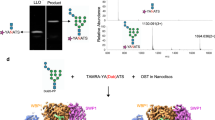
We present in vitro data that explain the recognition mechanism of misfolded glycoproteins by UDP-glucose glycoprotein–glucosyltransferase (UGGT). The glycoprotein exo-(1,3)-β-glucanase (β-Glc) bearing two glycans unfolds in a pH-dependent manner to become a misfolded substrate for UGGT. In the crystal structure of this glycoprotein, the local hydrophobicity surrounding each glycosylation site coincides with the differential recognition of N-linked glycans by UGGT. We introduced a single F280S point mutation, producing a β-Glc protein with full enzymatic activity that was both recognized as misfolded and monoglucosylated by UGGT. Contrary to current views, these data show that UGGT can modify N-linked glycans positioned at least 40 Å from localized regions of disorder and sense subtle conformational changes within structurally compact, enzymatically active glycoprotein substrates.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Helenius, A. & Aebi, M. Intracellular functions of N-linked glycans. Science 291, 2364–2369 (2001).
Pelletier, M.F., Bergeron, J.J.M. & Thomas, D.Y. Molecular chaperone systems in the endoplasmic reticulum. In Molecular Chaperones in the Cell (ed. Lund, P.) 180–200 (Oxford Univ. Press, Oxford, UK, 2000).
Herscovics, A. Importance of glycosidases in mammalian glycoprotein biosynthesis. Biochim. Biophys. Acta. 1473, 96–107 (1999).
Kornfeld, R. & Kornfeld, S. Assembly of asparagine-linked oligosaccharides. Annu. Rev. Biochem. 54, 631–664 (1985).
Grinna, L.S. & Robbins, P.W. Substrate specificities of rat liver microsomal glucosidases which process glycoproteins. J. Biol. Chem. 255, 2255–2258 (1980).
Pelletier, M.F. et al. The heterodimeric structure of glucosidase II is required for its activity, solubility and localization in vivo. Glycobiology 10, 815–827 (2000).
Schrag, J., Procopio, D., Cygler, M., Thomas, D.Y. & Bergeron, J.J.M. Lectin control of protein folding and sorting in the secretory pathway. Trends Biochem. Sci. 28, 49–57 (2003).
Zapun, A. et al. Conformation independent binding of monoglucosylated ribonuclease B to calnexin. Cell 88, 29–38 (1997).
Zapun, A. et al. Enhanced catalysis of ribonuclease B folding by the interaction of calnexin or calreticulin with ERp57. J. Biol. Chem. 273, 6009–6012 (1998).
Sousa, M.C., Ferrero-Garcia, M.A. & Parodi, A.J. Recognition of the oligosaccharide and protein moieties of glycoproteins by the UDP-Glc:glycoprotein glucosyltransferase. Biochemistry 31, 97–105 (1992).
Arnold, S.M. & Kaufman, R.J. The noncatalytic portion of human UDP-glucose:glycoprotein glucosytransferase I confers UDP-glucose binding and transferase function to the catalytic domain. J. Biol. Chem. 278, 43320–43328 (2003).
Sousa, M. & Parodi, A.J. The molecular basis for the recognition of misfolded glycoproteins by the UDP-Glc:glycoprotein glucosyltransferase. EMBO J. 14, 4196–4203 (1995).
Caramelo, J.J., Castro, O.A., Alonso, L.G., de Prat-Gay, G. & Parodi, A.J. UDP-Glc:glycoprotein glucosyltransferase recognizes structures and solvent accessible hydrophobic patches in molten globule-like folding intermediates. Proc. Natl. Acad. Sci. USA 100, 86–91 (2003).
Taylor, S.C., Thibault, P., Tessier, D.C., Bergeron, J.J.M. & Thomas, D.Y. Glycopeptide specificity of the secretory protein folding sensor UDP-glucose glycoprotein:glucosyltransferase. EMBO Rep. 4, 405–411 (2003).
Ritter, C. & Helenius, A. Recognition of local glycoprotein misfolding by the ER folding sensor UDP-glucose:glycoprotein glucosyltransferase. Nat. Struct. Biol. 7, 278–280 (2000).
Tessier, D.C. et al. Cloning and characterization of mammalian UDP-glucose glycoprotein: glucosyltransferase and the development of a specific substrate for this enzyme. Glycobiology 10, 403–412 (2000).
Sakon, J., Adney, W.S., Himmel, M.E., Thomas, S.R. & Karplus, P.A. Crystal structure of thermostable family 5 endocellulase E1 from Acidothermus cellulolyticus in complex with cellotetraose. Biochemistry 35, 10648–10660 (1996).
Basco, R.D. et al. Selective elongation of the oligosaccharide attached to the second potential glycosylation site of yeast exoglucanase: effects on the activity and properties of the enzyme. Biochem. J. 304, 917–922 (1994).
Ramirez, M., Hernandez, L.M. & Larriba, G. A similar protein portion for two exoglucanases secreted by Saccharomyces cerevisiae. Arch. Microbiol. 151, 391–398 (1989).
Bowler, B.E. et al. Destabilizing effects of replacing a surface lysine of cytochrome c with aromatic amino acids: implications for the denatured state. Biochemistry 32, 183–190 (1993).
Hammack, B. et al. The magnitude of changes in guanidine-HCl unfolding m-values in the protein, iso-1-cytochrome c, depends upon the substructure containing the mutation. Protein Sci. 7, 1789–1795 (1998).
Kimura, Y., Hess, D. & Sturm, A. The N-glycans of jack bean α-mannosidase. Structure, topology and function. Eur. J. Biochem. 264, 168–75 (1999).
Vernet, T., Dignard, D. & Thomas, D.Y. A family of yeast expression vectors. Gene 52, 225–233 (1987).
Wach, A., Brachat, A., Pohlmann, R. & Philippsen, P. New heterologous modules for classical or PCR-based gene disruptions in Saccharomyces cerevisiae. Yeast 10, 1793–1808 (1994).
Chen, D.C., Yang, B.C. & Kuo, T.T. One-step transformation of yeast in stationary phase. Curr. Genet. 21, 83–84 (1992).
Szewczyk, B. & Summers, D.F. Preparative elution of proteins blotted to Immobilon membranes. Anal. Biochem. 168, 48–53 (1988).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Cutfield, S.M. et al. The structure of the exo-β-(1,3)-glucanase from Candida albicans in native and bound forms: relationship between a pocket and groove in family 5 glycosyl hydrolases. J. Mol. Biol. 294, 771–783 (1999).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Acknowledgements
We thank M. Cygler, A. Matte and J. Schrag for their assistance and G. Larriba for antibodies. Operating grants from the Canadian Institutes of Health Research (to J.J.M.B. and D.Y.T) financially supported this work. A.D.F. was the recipient of a Canadian Institutes of Health Research postdoctoral fellowship, and is currently a fellow of the Human Frontier Science Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Taylor, S., Ferguson, A., Bergeron, J. et al. The ER protein folding sensor UDP-glucose glycoprotein–glucosyltransferase modifies substrates distant to local changes in glycoprotein conformation. Nat Struct Mol Biol 11, 128–134 (2004). https://doi.org/10.1038/nsmb715
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb715
This article is cited by
-
Identification of fungal lignocellulose-degrading biocatalysts secreted by Phanerochaete chrysosporium via activity-based protein profiling
Communications Biology (2022)
-
Post-translational protein modifications in schizophrenia
npj Schizophrenia (2020)
-
Multiple N-glycans cooperate in balancing misfolded BRI1 secretion and ER retention
Plant Molecular Biology (2020)
-
Structure Prediction of a Novel Exo-β-1,3-Glucanase: Insights into the Cold Adaptation of Psychrophilic Yeast Glaciozyma antarctica PI12
Interdisciplinary Sciences: Computational Life Sciences (2018)
-
Visualisation of a flexible modular structure of the ER folding-sensor enzyme UGGT
Scientific Reports (2017)