Abstract
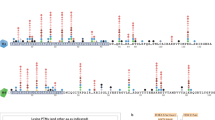
Histones comprise the major protein component of chromatin, the scaffold in which the eukaryotic genome is packaged, and are subject to many types of post-translational modifications (PTMs), especially on their flexible tails. These modifications may constitute a 'histone code' and could be used to manage epigenetic information that helps extend the genetic message beyond DNA sequences. This proposed code, read in part by histone PTM–binding 'effector' modules and their associated complexes, is predicted to define unique functional states of chromatin and/or regulate various chromatin-templated processes. A wealth of structural and functional data show how chromatin effector modules target their cognate covalent histone modifications. Here we summarize key features in molecular recognition of histone PTMs by a diverse family of 'reader pockets', highlighting specific readout mechanisms for individual marks, common themes and insights into the downstream functional consequences of the interactions. Changes in these interactions may have far-reaching implications for human biology and disease, notably cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Allis, C.D., Jenuwein, T., Reinberg, D. & Caparros, M.L (eds.). Epigenetics (Cold Spring Harbor Laboratory Press, Woodbury, New York, 2006).
Berger, S.L. The complex language of chromatin regulation during transcription. Nature 447, 407–412 (2007).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Pokholok, D.K. et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122, 517–527 (2005).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Mikkelsen, T.S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007).
Jenuwein, T. & Allis, C.D. Translating the histone code. Science 293, 1074–1080 (2001).
Strahl, B.D. & Allis, C.D. The language of covalent histone modifications. Nature 403, 41–45 (2000).
Ura, K., Kurumizaka, H., Dimitrov, S., Almouzni, G. & Wolffe, A.P. Histone acetylation: influence on transcription, nucleosome mobility and positioning, and linker histone-dependent transcriptional repression. EMBO J. 16, 2096–2107 (1997).
Shogren-Knaak, M. et al. Histone H4–K16 acetylation controls chromatin structure and protein interactions. Science 311, 844–847 (2006).
Ahn, S.H. et al. Sterile 20 kinase phosphorylates histone H2B at serine 10 during hydrogen peroxide-induced apoptosis in S. cerevisiae. Cell 120, 25–36 (2005).
Cosgrove, M.S., Boeke, J.D. & Wolberger, C. Regulated nucleosome mobility and the histone code. Nat. Struct. Mol. Biol. 11, 1037–1043 (2004).
Wolffe, A.P. & Hayes, J.J. Chromatin disruption and modification. Nucleic Acids Res. 27, 711–720 (1999).
Seet, B.T., Dikic, I., Zhou, M.M. & Pawson, T. Reading protein modifications with interaction domains. Nat. Rev. Mol. Cell Biol. 7, 473–483 (2006).
Ruthenburg, A.J., Allis, C.D. & Wysocka, J. Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol. Cell 25, 15–30 (2007).
Cheung, P., Allis, C.D. & Sassone-Corsi, P. Signaling to chromatin through histone modifications. Cell 103, 263–271 (2000).
Allfrey, V.G., Faulkner, R. & Mirsky, A.E. Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc. Natl. Acad. Sci. USA 51, 786–794 (1964).
Taunton, J., Hassig, C.A. & Schreiber, S.L. A mammalian histone deacetylase related to the yeast transcriptional regulator Rpd3p. Science 272, 408–411 (1996).
Brownell, J.E. et al. Tetrahymena histone acetyltransferase A: a homolog to yeast Gcn5p linking histone acetylation to gene activation. Cell 84, 843–851 (1996).
Dhalluin, C. et al. Structure and ligand of a histone acetyltransferase bromodomain. Nature 399, 491–496 (1999).
Zeng, L. & Zhou, M.M. Bromodomain: an acetyl-lysine binding domain. FEBS Lett. 513, 124–128 (2002).
Owen, D.J. et al. The structural basis for the recognition of acetylated histone H4 by the bromodomain of histone acetyltransferase gcn5p. EMBO J. 19, 6141–6149 (2000).
Kuo, M.H. et al. Transcription-linked acetylation by Gcn5p of histones H3 and H4 at specific lysines. Nature 383, 269–272 (1996).
Jacobson, R.H., Ladurner, A.G., King, D.S. & Tjian, R. Structure and function of a human TAFII250 double bromodomain module. Science 288, 1422–1425 (2000).
VanDemark, A.P. et al. Autoregulation of the rsc4 tandem bromodomain by gcn5 acetylation. Mol. Cell 27, 817–828 (2007).
Kasten, M. et al. Tandem bromodomains in the chromatin remodeler RSC recognize acetylated histone H3 Lys14. EMBO J. 23, 1348–1359 (2004).
Cheng, X., Collins, R.E. & Zhang, X. Structural and sequence motifs of protein (histone) methylation enzymes. Annu. Rev. Biophys. Biomol. Struct. 34, 267–294 (2005).
Shi, Y. & Whetstine, J.R. Dynamic regulation of histone lysine methylation by demethylases. Mol. Cell 25, 1–14 (2007).
Swigut, T. & Wysocka, J. H3K27 demethylase at last: what it means for memory and plasticity of gene expression in developmental processes. Cell 131, 29–32 (2007).
Martin, C. & Zhang, Y. The diverse functions of histone lysine methylation. Nat. Rev. Mol. Cell Biol. 6, 838–849 (2005).
Guenther, M.G., Levine, S.S., Boyer, L.A., Jaenisch, R. & Yuung, R.A. A chromatin landmark and transcription initiation at most promoters in human cells. Cell 130, 77–78 (2007).
Bernstein, B.E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Hughes, R.M., Wiggins, K.R., Khorasanizadeh, S. & Waters, M.L. Recognition of trimethyllysine by a chromodomain is not driven by the hydrophobic effect. Proc. Natl. Acad. Sci. USA 104, 11184–11188 (2007).
Ma, J.C. & Dougherty, D.A. Cation-π Interaction. Chem. Rev. 97, 1303–1324 (1997).
Li, H. et al. Structural basis for lower lysine methylation state-specific readout by MBT repeats and an engineered PHD finger. Mol. Cell (in the press).
Botuyan, M.V. et al. Structural basis for the methylation state-specific recognition of histone H4–K20 by 53BP1 and Crb2 in DNA repair. Cell 127, 1361–1373 (2006).
Burley, S.K. & Petsko, G.A. Weakly polar interactions in proteins. Adv. Protein Chem. 39, 125–189 (1988).
Maurer-Stroh, S. et al. The Tudor domain 'Royal Family': Tudor, plant Agenet, Chromo, PWWP and MBT domains. Trends Biochem. Sci. 28, 69–74 (2003).
Pena, P.V. et al. Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 442, 100–103 (2006).
Li, H. et al. Molecular basis for site-specific read-out of histone H3K4me3 by the BPTF PHD finger of NURF. Nature 442, 91–95 (2006).
Paro, R. & Hogness, D.S. The Polycomb protein shares a homologous domain with a heterochromatin-associated protein of Drosophila. Proc. Natl. Acad. Sci. USA 88, 263–267 (1991).
Lachner, M., O'Carroll, D., Rea, S., Mechtler, K. & Jenuwein, T. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410, 116–120 (2001).
Bannister, A.J. et al. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410, 120–124 (2001).
Jacobs, S.A. et al. Specificity of the HP1 chromo domain for the methylated N-terminus of histone H3. EMBO J. 20, 5232–5241 (2001).
Jacobs, S.A. & Khorasanizadeh, S. Structure of HP1 chromodomain bound to a lysine 9-methylated histone H3 tail. Science 295, 2080–2083 (2002).
Nielsen, P.R. et al. Structure of the HP1 chromodomain bound to histone H3 methylated at lysine 9. Nature 416, 103–107 (2002).
Fischle, W. et al. Molecular basis for the discrimination of repressive methyl-lysine marks in histone H3 by Polycomb and HP1 chromodomains. Genes Dev. 17, 1870–1881 (2003).
Min, J., Zhang, Y. & Xu, R.M. Structural basis for specific binding of Polycomb chromodomain to histone H3 methylated at Lys 27. Genes Dev. 17, 1823–1828 (2003).
Nielsen, A.L. et al. Heterochromatin formation in mammalian cells: interaction between histones and HP1 proteins. Mol. Cell 7, 729–739 (2001).
Zhao, T., Heyduk, T., Allis, C.D. & Eissenberg, J.C. Heterochromatin protein 1 binds to nucleosomes and DNA in vitro. J. Biol. Chem. 275, 28332–28338 (2000).
Lusser, A., Urwin, D.L. & Kadonaga, J.T. Distinct activities of CHD1 and ACF in ATP-dependent chromatin assembly. Nat. Struct. Mol. Biol. 12, 160–166 (2005).
Sims, R.J. III, Belotserkovskaya, R. & Reinberg, D. Elongation by RNA polymerase II: the short and long of it. Genes Dev. 18, 2437–2468 (2004).
Konev, A.Y. et al. CHD1 motor protein is required for deposition of histone variant H3.3 into chromatin in vivo. Science 317, 1087–1090 (2007).
Taverna, S.D. et al. Long-distance combinatorial linkage between methylation and acetylation on histone H3 N termini. Proc. Natl. Acad. Sci. USA 104, 2086–2091 (2007).
Hake, S.B. et al. Expression patterns and post-translational modifications associated with mammalian histone H3 variants. J. Biol. Chem. 281, 559–568 (2006).
Pray-Grant, M.G., Daniel, J.A., Schieltz, D., Yates, J.R. III & Grant, P.A. Chd1 chromodomain links histone H3 methylation with SAGA- and SLIK-dependent acetylation. Nature 433, 434–438 (2005).
Flanagan, J.F. et al. Double chromodomains cooperate to recognize the methylated histone H3 tail. Nature 438, 1181–1185 (2005).
Sims, R.J. III et al. Human but not yeast CHD1 binds directly and selectively to histone H3 methylated at lysine 4 via its tandem chromodomains. J. Biol. Chem. 280, 41789–41792 (2005).
Okuda, M., Horikoshi, M. & Nishimura, Y. Structural polymorphism of chromodomains in Chd1. J. Mol. Biol. 365, 1047–1062 (2007).
Klose, R.J. et al. The transcriptional repressor JHDM3A demethylates trimethyl histone H3 lysine 9 and lysine 36. Nature 442, 312–316 (2006).
Whetstine, J.R. et al. Reversal of histone lysine trimethylation by the JMJD2 family of histone demethylases. Cell 125, 467–481 (2006).
Klose, R.J., Kallin, E.M. & Zhang, Y. JmjC-domain-containing proteins and histone demethylation. Nat. Rev. Genet. 7, 715–727 (2006).
Huang, Y., Fang, J., Bedford, M.T., Zhang, Y. & Xu, R.M. Recognition of histone H3 lysine-4 methylation by the double tudor domain of JMJD2A. Science 312, 748–751 (2006).
Kim, J. et al. Tudor, MBT and chromo domains gauge the degree of lysine methylation. EMBO Rep. 7, 397–403 (2006).
Sanders, S.L. et al. Methylation of histone H4 lysine 20 controls recruitment of Crb2 to sites of DNA damage. Cell 119, 603–614 (2004).
Huyen, Y. et al. Methylated lysine 79 of histone H3 targets 53BP1 to DNA double-strand breaks. Nature 432, 406–411 (2004).
Huang, J. et al. p53 is regulated by the lysine demethylase LSD1. Nature 449, 105–108 (2007).
Koga, H. et al. A human homolog of Drosophila lethal(3)malignant brain tumor (l(3)mbt) protein associates with condensed mitotic chromosomes. Oncogene 18, 3799–3809 (1999).
Pirrotta, V. Chromatin-silencing mechanisms in Drosophila maintain patterns of gene expression. Trends Genet. 13, 314–318 (1997).
Boccuni, P., MacGrogan, D., Scandura, J.M. & Nimer, S.D. The human L(3)MBT polycomb group protein is a transcriptional repressor and interacts physically and functionally with TEL (ETV6). J. Biol. Chem. 278, 15412–15420 (2003).
Trojer, P. et al. L3MBTL1, a histone-methylation-dependent chromatin lock. Cell 129, 915–928 (2007).
Wang, W.K. et al. Malignant brain tumor repeats: a three-leaved propeller architecture with ligand/peptide binding pockets. Structure 11, 775–789 (2003).
Santos-Rosa, H. et al. Active genes are tri-methylated at K4 of histone H3. Nature 419, 407–411 (2002).
Taverna, S.D. et al. Yng1 PHD finger binding to H3 trimethylated at K4 promotes NuA3 HAT activity at K14 of H3 and transcription at a subset of targeted ORFs. Mol. Cell 24, 785–796 (2006).
Vermeulen, M. et al. Selective anchoring of TFIID to nucleosomes by trimethylation of histone H3 lysine 4. Cell 131, 58–69 (2007).
Tsukiyama, T. & Wu, C. Purification and properties of an ATP-dependent nucleosome remodeling factor. Cell 83, 1011–1020 (1995).
Mizuguchi, G., Tsukiyama, T., Wisniewski, J. & Wu, C. Role of nucleosome remodeling factor NURF in transcriptional activation of chromatin. Mol. Cell 1, 141–150 (1997).
Wysocka, J. et al. A PHD finger of NURF couples histone H3 lysine 4 trimethylation with chromatin remodelling. Nature 442, 86–90 (2006).
Bienz, M. The PHD finger, a nuclear protein-interaction domain. Trends Biochem. Sci. 31, 35–40 (2006).
Doyon, Y. et al. ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol. Cell 21, 51–64 (2006).
Millar, C.B. & Grunstein, M. Genome-wide patterns of histone modifications in yeast. Nat. Rev. Mol. Cell Biol. 7, 657–666 (2006).
Martin, D.G. et al. The Yng1p PHD finger is a methyl-histone binding module that recognizes lysine 4 methylated histone H3. Mol. Cell. Biol. 26, 7871–7879 (2006).
Shi, X. et al. ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442, 96–99 (2006).
Ruthenburg, A.J., Li, H., Taverna, S.D., Patel, D.J. & Allis, C.D. Multivalent readout of histone modifications by linked effector modules on a nucleosome scaffold. Nat. Rev. Mol. Cell Biol. (in the press).
Edmondson, D.G., Smith, M.M. & Roth, S.Y. Repression domain of the yeast global repressor Tup1 interacts directly with histones H3 and H4. Genes Dev. 10, 1247–1259 (1996).
Ooi, S.K. et al. DNMT3L connects unmethylated lysine 4 of histone H3 to de novo methylation of DNA. Nature 448, 714–717 (2007).
Lan, F. et al. Recognition of unmethylated histone H3 lysine 4 links BHC80 to LSD1-mediated gene repression. Nature 448, 718–722 (2007).
Couture, J.F., Collazo, E. & Trievel, R.C. Molecular recognition of histone H3 by the WD40 protein WDR5. Nat. Struct. Mol. Biol. 13, 698–703 (2006).
Ruthenburg, A.J. et al. Histone H3 recognition and presentation by the WDR5 module of the MLL1 complex. Nat. Struct. Mol. Biol. 13, 704–712 (2006).
Schuetz, A. et al. Structural basis for molecular recognition and presentation of histone H3 by WDR5. EMBO J. 25, 4245–4252 (2006).
Han, Z. et al. Structural basis for the specific recognition of methylated histone H3 lysine 4 by the WD-40 protein WDR5. Mol. Cell 22, 137–144 (2006).
Hakimi, M.A. et al. A core-BRAF35 complex containing histone deacetylase mediates repression of neuronal-specific genes. Proc. Natl. Acad. Sci. USA 99, 7420–7425 (2002).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Marmorstein, L.Y. et al. A human BRCA2 complex containing a structural DNA binding component influences cell cycle progression. Cell 104, 247–257 (2001).
Gruenbaum, Y., Cedar, H. & Razin, A. Substrate and sequence specificity of a eukaryotic DNA methylase. Nature 295, 620–622 (1982).
Bestor, T.H. & Ingram, V.M. Two DNA methyltransferases from murine erythroleukemia cells: purification, sequence specificity, and mode of interaction with DNA. Proc. Natl. Acad. Sci. USA 80, 5559–5563 (1983).
Okano, M., Xie, S. & Li, E. Cloning and characterization of a family of novel mammalian DNA (cytosine-5) methyltransferases. Nat. Genet. 19, 219–220 (1998).
Bourc'his, D., Xu, G.L., Lin, C.S., Bollman, B. & Bestor, T.H. Dnmt3L and the establishment of maternal genomic imprints. Science 294, 2536–2539 (2001).
Argentaro, A. et al. Structural consequences of disease-causing mutations in the ATRX-DNMT3-DNMT3L (ADD) domain of the chromatin-associated protein ATRX. Proc. Natl. Acad. Sci. USA 104, 11939–11944 (2007).
Yokoyama, A. et al. Leukemia proto-oncoprotein MLL forms a SET1-like histone methyltransferase complex with menin to regulate Hox gene expression. Mol. Cell. Biol. 24, 5639–5649 (2004).
Wysocka, J. et al. WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell 121, 859–872 (2005).
Dou, Y. et al. Regulation of MLL1 H3K4 methyltransferase activity by its core components. Nat. Struct. Mol. Biol. 13, 713–719 (2006).
Verreault, A., Kaufman, P.D., Kobayashi, R. & Stillman, B. Nucleosomal DNA regulates the core-histone-binding subunit of the human Hat1 acetyltransferase. Curr. Biol. 8, 96–108 (1998).
Muslin, A.J., Tanner, J.W., Allen, P.M. & Shaw, A.S. Interaction of 14–3-3 with signaling proteins is mediated by the recognition of phosphoserine. Cell 84, 889–897 (1996).
Yaffe, M.B. et al. The structural basis for 14–3-3:phosphopeptide binding specificity. Cell 91, 961–971 (1997).
Dougherty, M.K. & Morrison, D.K. Unlocking the code of 14–3-3. J. Cell Sci. 117, 1875–1884 (2004).
Thomson, S. et al. The nucleosomal response associated with immediate-early gene induction is mediated via alternative MAP kinase cascades: MSK1 as a potential histone H3/HMG-14 kinase. EMBO J. 18, 4779–4793 (1999).
Soloaga, A. et al. MSK2 and MSK1 mediate the mitogen- and stress-induced phosphorylation of histone H3 and HMG-14. EMBO J. 22, 2788–2797 (2003).
Nowak, S.J. & Corces, V.G. Phosphorylation of histone H3: a balancing act between chromosome condensation and transcriptional activation. Trends Genet. 20, 214–220 (2004).
Duncan, E.A., Anest, V., Cogswell, P. & Baldwin, A.S. The kinases MSK1 and MSK2 are required for epidermal growth factor-induced, but not tumor necrosis factor-induced, histone H3 Ser10 phosphorylation. J. Biol. Chem. 281, 12521–12525 (2006).
Cheung, P. et al. Synergistic coupling of histone H3 phosphorylation and acetylation in response to epidermal growth factor stimulation. Mol. Cell 5, 905–915 (2000).
Macdonald, N. et al. Molecular basis for the recognition of phosphorylated and phosphoacetylated histone h3 by 14–3-3. Mol. Cell 20, 199–211 (2005).
Downs, J.A., Nussenzweig, M.C. & Nussenzweig, A. Chromatin dynamics and the preservation of genetic information. Nature 447, 951–958 (2007).
Rogakou, E.P., Pilch, D.R., Orr, A.H., Ivanova, V.S. & Bonner, W.M. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J. Biol. Chem. 273, 5858–5868 (1998).
Burma, S., Chen, B.P., Murphy, M., Kurimasa, A. & Chen, D.J. ATM phosphorylates histone H2AX in response to DNA double-strand breaks. J. Biol. Chem. 276, 42462–42467 (2001).
Peng, A. & Chen, P.L. NFBD1, like 53BP1, is an early and redundant transducer mediating Chk2 phosphorylation in response to DNA damage. J. Biol. Chem. 278, 8873–8876 (2003).
Lee, M.S., Edwards, R.A., Thede, G.L. & Glover, J.N. Structure of the BRCT repeat domain of MDC1 and its specificity for the free COOH-terminal end of the gamma-H2AX histone tail. J. Biol. Chem. 280, 32053–32056 (2005).
Stucki, M. et al. MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks. Cell 123, 1213–1226 (2005).
Clapperton, J.A. et al. Structure and mechanism of BRCA1 BRCT domain recognition of phosphorylated BACH1 with implications for cancer. Nat. Struct. Mol. Biol. 11, 512–518 (2004).
Wu, G. et al. Structure of a β-TrCP1-Skp1-β-catenin complex: destruction motif binding and lysine specificity of the SCF(β-TrCP1) ubiquitin ligase. Mol. Cell 11, 1445–1456 (2003).
Xiong, J.P. et al. Crystal structure of the extracellular segment of integrin alpha Vbeta3. Science 294, 339–345 (2001).
Wall, M.A. et al. The structure of the G protein heterotrimer Gi alpha 1 beta 1 gamma 2. Cell 83, 1047–1058 (1995).
Hirota, T., Lipp, J.J., Toh, B.H. & Peters, J.M. Histone H3 serine 10 phosphorylation by Aurora B causes HP1 dissociation from heterochromatin. Nature 438, 1176–1180 (2005).
Fischle, W. et al. Regulation of HP1-chromatin binding by histone H3 methylation and phosphorylation. Nature 438, 1116–1122 (2005).
Fischle, W., Wang, Y. & Allis, C.D. Binary switches and modification cassettes in histone biology and beyond. Nature 425, 475–479 (2003).
Sampath, S.C. et al. Methylation of histone mimic within the histone methyltransferase G9a regulates protein complex assembly. Mol. Cell 27, 596–608 (2007).
Wang, G.G., Allis, C.D. & Chi, P. Chromatin remodeling and cancer, part I: covalent histone modifications. Trends Mol. Med. 13, 363–372 (2007).
Hsu, J.Y. et al. Mitotic phosphorylation of histone H3 is governed by Ipl1/aurora kinase and Glc7/PP1 phosphatase in budding yeast and nematodes. Cell 102, 279–291 (2000).
Chen, C.C., Smith, D.L., Bruegger, B.B., Halpern, R.M. & Smith, R.A. Occurrence and distribution of acid-labile histone phosphates in regenerating rat liver. Biochemistry 13, 3785–3789 (1974).
Carrozza, M.J. et al. Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123, 581–592 (2005).
Joshi, A.A. & Struhl, K. Eaf3 chromodomain interaction with methylated H3–K36 links histone deacetylation to Pol II elongation. Mol. Cell 20, 971–978 (2005).
Li, B. et al. Combined action of PHD and chromo domains directs the Rpd3S HDAC to transcribed chromatin. Science 316, 1050–1054 (2007).
Allis, C.D., Bowen, J.K., Abraham, G.N., Glover, C.V. & Gorovsky, M.A. Proteolytic processing of histone H3 in chromatin: a physiologically regulated event in Tetrahymena micronuclei. Cell 20, 55–64 (1980).
Garcia, B.A., Pesavento, J.J., Mizzen, C.A. & Kelleher, N.L. Pervasive combinatorial modification of histone H3 in human cells. Nat. Methods 4, 487–489 (2007).
Strahl, B.D. et al. Methylation of histone H4 at arginine 3 occurs in vivo and is mediated by the nuclear receptor coactivator PRMT1. Curr. Biol. 11, 996–1000 (2001).
Wang, H. et al. Methylation of histone H4 at arginine 3 facilitating transcriptional activation by nuclear hormone receptor. Science 293, 853–857 (2001).
An, W., Kim, J. & Roeder, R.G. Ordered cooperative functions of PRMT1, p300, and CARM1 in transcriptional activation by p53. Cell 117, 735–748 (2004).
Cuthbert, G.L. et al. Histone deimination antagonizes arginine methylation. Cell 118, 545–553 (2004).
Wang, Y. et al. Human PAD4 regulates histone arginine methylation levels via demethylimination. Science 306, 279–283 (2004).
Selenko, P. et al. SMN tudor domain structure and its interaction with the Sm proteins. Nat. Struct. Biol. 8, 27–31 (2001).
Sprangers, R., Groves, M.R., Sinning, I. & Sattler, M. High-resolution X-ray and NMR structures of the SMN Tudor domain: conformational variation in the binding site for symmetrically dimethylated arginine residues. J. Mol. Biol. 327, 507–520 (2003).
Kirmizis, A. et al. Arginine methylation at histone H3R2 controls deposition of H3K4 trimethylation. Nature 449, 928–932 (2007).
Guccione, E. et al. Methylation of histone H3R2 by PRMT6 and H3K4 by an MLL complex are mutually exclusive. Nature 449; 933–937 (2007)
Ahn, S.H., Diaz, R.L., Grunstein, M. & Allis, C.D. Histone H2B deacetylation at lysine 11 is required for yeast apoptosis induced by phosphorylation of H2B at serine 10. Mol. Cell 24, 211–220 (2006).
Latham, J.A. & Dent, S.Y.R. Cross-regulation of histone modifications. Nat. Struct. Mol. Biol. 14, 1017–1024 (2007).
Feinberg, A.P. Phenotypic plasticity and the epigenetics of human disease. Nature 447, 433–440 (2007).
Spadaccini, R., Perrin, H., Bottomley, M.J., Ansieau, S. & Sattler, M. Structure and functional analysis of the MYND domain. J. Mol. Biol. 358, 498–508 (2006).
Liu, Y. et al. Structural basis for recognition of SMRT/N-CoR by the MYND domain and its contribution to AML1/ETO's activity. Cancer Cell 11, 483–497 (2007).
Chang, B., Chen, Y., Zhao, Y. & Bruick, R.K. JMJD6 is a histone arginine demethylase. Science 318, 444–447 (2007).
Acknowledgements
We apologize to all of the researchers whose important contributions could not be acknowledged because of space constraints. We thank members of the Patel and Allis laboratories as well as the anonymous reviewers for critically reading the manuscript, and A. VanDemark (University of Pittsburgh) and C.P. Hill (University of Utah) for providing coordinates for Rsc4p in Figure 2e. D.J.P. is supported by funds from the Abby Rockefeller Mauze Trust and the Dewitt Wallace and Maloris Foundations, C.D.A. and S.D.T are supported by US National Institutes of Health grants GM53512 and GM63959 and by funds from The Rockefeller University, and A.J.R. is supported by a postdoctoral fellowship from the Irvington Foundation.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Rights and permissions
About this article
Cite this article
Taverna, S., Li, H., Ruthenburg, A. et al. How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat Struct Mol Biol 14, 1025–1040 (2007). https://doi.org/10.1038/nsmb1338
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1338
This article is cited by
-
Caf1 regulates the histone methyltransferase activity of Ash1 by sensing unmodified histone H3
Epigenetics & Chromatin (2023)
-
Limited choice of natural amino acids as mimetics restricts design of protein lysine methylation studies
Nature Communications (2023)
-
Systematic characterization of chromodomain proteins reveals an H3K9me1/2 reader regulating aging in C. elegans
Nature Communications (2023)
-
Epigenetic mechanisms to propagate histone acetylation by p300/CBP
Nature Communications (2023)
-
De novo methylation of histone H3K23 by the methyltransferases EHMT1/GLP and EHMT2/G9a
Epigenetics & Chromatin (2022)