Abstract
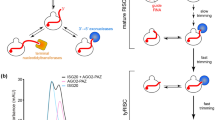
In C. elegans, DCR-1 is required for the maturation of both short interfering RNAs (siRNAs) and microRNAs (miRNAs), which are subsequently loaded into different Argonaute proteins to mediate silencing via distinct mechanisms. We used in vivo analyses to show that precursors of small RNAs contain structural features that direct the small RNAs into the RNA interference (RNAi) pathway or the miRNA-processing pathway. Nucleotide changes in the pre-let-7 miRNA precursor that make its stem fully complementary cause the resulting small RNA to be recognized as siRNA and induce binding to RDE-1, which leads to RNAi. Mismatches of 1 to 3 nucleotides at various positions in the stem of the precursor restore direction into the miRNA pathway, as the largest portion of such small RNA variants is associated with ALG-1. The Argonaute proteins to which the small RNAs are bound determine the silencing mode, and no functional overlap between RDE-1 and ALG-1 was detected.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Lee, R.C., Feinbaum, R.L. & Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843–854 (1993).
Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Kim, V.N. Small RNAs: classification, biogenesis, and function. Mol. Cells 19, 1–15 (2005).
Bernstein, E., Caudy, A.A., Hammond, S.M. & Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001).
Hutvagner, G. et al. A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 293, 834–838 (2001).
Zhang, H., Kolb, F.A., Brondani, V., Billy, E. & Filipowicz, W. Human Dicer preferentially cleaves dsRNAs at their termini without a requirement for ATP. EMBO J. 21, 5875–5885 (2002).
Zhang, H., Kolb, F.A., Jaskiewicz, L., Westhof, E. & Filipowicz, W. Single processing center models for human Dicer and bacterial RNase III. Cell 118, 57–68 (2004).
Schwarz, D.S. et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115, 199–208 (2003).
Lee, Y., Jeon, K., Lee, J.T., Kim, S. & Kim, V.N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 21, 4663–4670 (2002).
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003).
Yi, R., Qin, Y., Macara, I.G. & Cullen, B.R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 17, 3011–3016 (2003).
Lund, E., Guttinger, S., Calado, A., Dahlberg, J.E. & Kutay, U. Nuclear export of microRNA precursors. Science 303, 95–98 (2004).
Lee, Y.S. et al. Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell 117, 69–81 (2004).
Xie, Z. et al. Genetic and functional diversification of small RNA pathways in plants. PLoS Biol. 2, E104 (2004).
Vazquez, F. Arabidopsis endogenous small RNAs: highways and byways. Trends Plant Sci. 11, 460–468 (2006).
Brodersen, P. & Voinnet, O. The diversity of RNA silencing pathways in plants. Trends Genet. 22, 268–280 (2006).
Grishok, A. et al. Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106, 23–34 (2001).
Ketting, R.F. et al. Dicer functions in RNA interference and in synthesis of small RNA involved in developmental timing in C. elegans. Genes Dev. 15, 2654–2659 (2001).
Tabara, H. et al. The rde-1 gene, RNA interference, and transposon silencing in C. elegans. Cell 99, 123–132 (1999).
Sijen, T. et al. On the role of RNA amplification in dsRNA-triggered gene silencing. Cell 107, 465–476 (2001).
Yigit, E. et al. Analysis of the C. elegans Argonaute family reveals that distinct Argonautes act sequentially during RNAi. Cell 127, 747–757 (2006).
Valencia-Sanchez, M.A., Liu, J., Hannon, G.J. & Parker, R. Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev. 20, 515–524 (2006).
Pillai, R.S. et al. Inhibition of translational initiation by Let-7 MicroRNA in human cells. Science 309, 1573–1576 (2005).
Pillai, R.S., Bhattacharyya, S.N. & Filipowicz, W. Repression of protein synthesis by miRNAs: how many mechanisms? Trends Cell Biol. 17, 118–126 (2007).
Liu, Q. et al. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 301, 1921–1925 (2003).
Caudy, A.A. & Hannon, G.J. Induction and biochemical purification of RNA-induced silencing complex from Drosophila S2 cells. Methods Mol. Biol. 265, 59–72 (2004).
Rand, T.A., Ginalski, K., Grishin, N.V. & Wang, X. Biochemical identification of Argonaute 2 as the sole protein required for RNA-induced silencing complex activity. Proc. Natl. Acad. Sci. USA 101, 14385–14389 (2004).
Forstemann, K. et al. Normal microRNA maturation and germ-line stem cell maintenance requires Loquacious, a double-stranded RNA-binding domain protein. PLoS Biol. 3, e236 (2005).
Saito, K., Ishizuka, A., Siomi, H. & Siomi, M.C. Processing of pre-microRNAs by the Dicer-1-Loquacious complex in Drosophila cells. PLoS Biol. 3, e235 (2005).
Okamura, K., Ishizuka, A., Siomi, H. & Siomi, M.C. Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev. 18, 1655–1666 (2004).
Tabara, H., Yigit, E., Siomi, H. & Mello, C.C. The dsRNA binding protein RDE-4 interacts with RDE-1, DCR-1, and a DExH-box helicase to direct RNAi in C. elegans. Cell 109, 861–871 (2002).
Parker, G.S., Eckert, D.M. & Bass, B.L. RDE-4 preferentially binds long dsRNA and its dimerization is necessary for cleavage of dsRNA to siRNA. RNA 12, 807–818 (2006).
Sijen, T., Steiner, F.A., Thijssen, K.L. & Plasterk, R.H. Secondary siRNAs result from unprimed RNA synthesis and form a distinct class. Science 315, 244–247 (2007).
Kim, V.N. MicroRNA biogenesis: coordinated cropping and dicing. Nat. Rev. Mol. Cell Biol. 6, 376–385 (2005).
Tops, B.B., Plasterk, R.H. & Ketting, R.F. The Caenorhabditis elegans Argonautes ALG-1 and ALG-2: almost identical yet different. Cold Spring Harb. Symp. Quant. Biol. 71, 189–194 (2006).
Caudy, A.A. et al. A micrococcal nuclease homologue in RNAi effector complexes. Nature 425, 411–414 (2003).
Lu, R. et al. Animal virus replication and RNAi-mediated antiviral silencing in Caenorhabditis elegans. Nature 436, 1040–1043 (2005).
Wilkins, C. et al. RNA interference is an antiviral defence mechanism in Caenorhabditis elegans. Nature 436, 1044–1047 (2005).
Plasterk, R.H. RNA silencing: the genome's immune system. Science 296, 1263–1265 (2002).
Tomari, Y., Du, T. & Zamore, P.D. Sorting of Drosophila small silencing RNAs. Cell 130, 299–308 (2007).
Forstemann, K., Horwich, M.D., Wee, L., Tomari, Y. & Zamore, P.D. Drosophila microRNAs are sorted into functionally distinct argonaute complexes after production by dicer-1. Cell 130, 287–297 (2007).
Lee, R.C., Hammell, C.M. & Ambros, V. Interacting endogenous and exogenous RNAi pathways in Caenorhabditis elegans. RNA 12, 589–597 (2006).
Duchaine, T.F. et al. Functional proteomics reveals the biochemical niche of C. elegans DCR-1 in multiple small-RNA-mediated pathways. Cell 124, 343–354 (2006).
Pak, J. & Fire, A. Distinct populations of primary and secondary effectors during RNAi in C. elegans. Science 315, 241–244 (2007).
Ruby, J.G. et al. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 127, 1193–1207 (2006).
Vagin, V.V. et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324 (2006).
Houwing, S. et al. A role for Piwi and piRNAs in germ cell maintenance and transposon silencing in Zebrafish. Cell 129, 69–82 (2007).
Acknowledgements
We thank G. Hannon and F. Rivas (Cold Spring Harbor Laboratory) for providing reagents and help with the capture assays, B. Tops (Hubrecht Institute) and C. Mello (University of Massachusetts Medical School) for providing strains and W. Kloosterman for discussions and critical reading of the manuscript. This work was supported by a Vidi fellowship from the Dutch Scientific Organization (NWO) to T.S. and a European Union grant (HPRN-CT-2002-00257) and Netherlands Genomics Initiative grant (050-72-415) to F.A.S.
Author information
Authors and Affiliations
Contributions
F.A.S., S.W.H., K.L.O., K.L.T. and T.S. carried out experiments; T.S., F.A.S., R.F.K. and R.H.A.P. supervised the research; F.A.S. and T.S. wrote the paper.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 (PDF 1677 kb)
Rights and permissions
About this article
Cite this article
Steiner, F., Hoogstrate, S., Okihara, K. et al. Structural features of small RNA precursors determine Argonaute loading in Caenorhabditis elegans. Nat Struct Mol Biol 14, 927–933 (2007). https://doi.org/10.1038/nsmb1308
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1308
This article is cited by
-
Cell-type specific sequencing of microRNAs from complex animal tissues
Nature Methods (2018)
-
ARGONAUTE PIWI domain and microRNA duplex structure regulate small RNA sorting in Arabidopsis
Nature Communications (2014)
-
Regulation of microRNA biogenesis
Nature Reviews Molecular Cell Biology (2014)
-
Physiological stressors and invasive plant infections alter the small RNA transcriptome of the rice blast fungus, Magnaporthe oryzae
BMC Genomics (2013)
-
Differential effects of viral silencing suppressors on siRNA and miRNA loading support the existence of two distinct cellular pools of ARGONAUTE1
The EMBO Journal (2012)