Abstract
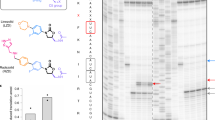
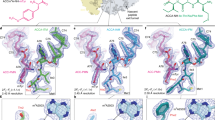
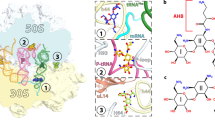
Aminoglycosides are widely used antibiotics that cause messenger RNA decoding errors, block mRNA and transfer RNA translocation, and inhibit ribosome recycling. Ribosome recycling follows the termination of protein synthesis and is aided by ribosome recycling factor (RRF) in bacteria. The molecular mechanism by which aminoglycosides inhibit ribosome recycling is unknown. Here we show in X-ray crystal structures of the Escherichia coli 70S ribosome that RRF binding causes RNA helix H69 of the large ribosomal subunit, which is crucial for subunit association, to swing away from the subunit interface. Aminoglycosides bind to H69 and completely restore the contacts between ribosomal subunits that are disrupted by RRF. These results provide a structural explanation for aminoglycoside inhibition of ribosome recycling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Magnet, S. & Blanchard, J.S. Molecular insights into aminoglycoside action and resistance. Chem. Rev. 105, 477–498 (2005).
Sutcliffe, J.A. Improving on nature: antibiotics that target the ribosome. Curr. Opin. Microbiol. 8, 534–542 (2005).
Davies, J., Gorini, L. & Davies, B.D. Misreading of RNA codewords induced by aminoglycoside antibiotics. Mol. Pharmacol. 1, 93–106 (1965).
Davies, J., Jones, D.S. & Khorana, H.G. A further study of misreading of codons induced by streptomycin and neomycin using ribopolynucleotides containing two nucleotides in alternating sequence as templates. J. Mol. Biol. 18, 48–57 (1966).
Cabañas, M.J., Vázquez, D. & Modolell, J. Inhibition of ribosomal translocation by aminoglycoside antibiotics. Biochem. Biophys. Res. Commun. 83, 991–997 (1978).
Misumi, M., Nishimura, T., Komai, T. & Tanaka, N. Interaction of kanamycin and related antibiotics with the large subunit of ribosomes and the inhibition of translocation. Biochem. Biophys. Res. Commun. 84, 358–365 (1978).
Hirokawa, G. et al. Post-termination complex disassembly by ribosome recycling factor, a functional tRNA mimic. EMBO J. 21, 2272–2281 (2002).
Wilson, D.N. & Nierhaus, K.H. The how and Y of cold shock. Nat. Struct. Mol. Biol. 11, 1026–1028 (2004).
Janosi, L. et al. Evidence for in vivo ribosome recycling, the fourth step in protein biosynthesis. EMBO J. 17, 1141–1151 (1998).
Zavialov, A.V., Hauryliuk, V.V. & Ehrenberg, M. Splitting of the posttermination ribosome into subunits by the concerted action of RRF and EF-G. Mol. Cell 18, 675–686 (2005).
Gao, N. et al. Mechanism for the disassembly of the posttermination complex inferred from cryo-EM studies. Mol. Cell 18, 663–674 (2005).
Peske, F., Rodnina, M.V. & Wintermeyer, W. Sequence of steps in ribosome recycling as defined by kinetic analysis. Mol. Cell 18, 403–412 (2005).
Hirokawa, G., Demeshkina, N., Iwakura, N., Kaji, H. & Kaji, A. The ribosome-recycling step: consensus or controversy? Trends Biochem. Sci. 31, 143–149 (2006).
Hirokawa, G. et al. The role of ribosome recycling factor in dissociation of 70S ribosomes into subunits. RNA 11, 1317–1328 (2005).
Teyssier, E. et al. Temperature-sensitive mutation in yeast mitochondrial ribosome-recycling factor (RRF). Nucleic Acids Res. 31, 4218–4226 (2003).
Rolland, N. et al. Plant ribosome recycling factor homologue is a chloroplastic protein and is bactericidal in Escherichia coli carrying temperature-sensitive ribosome recycling factor. Proc. Natl. Acad. Sci. USA 96, 5464–5469 (1999).
Janosi, L., Shimizu, I. & Kaji, A. Ribosome recycling factor (ribosome releasing factor) is essential for bacterial growth. Proc. Natl. Acad. Sci. USA 91, 4249–4253 (1994).
Campuzano, S., Vázquez, D. & Modolell, J. Functional interaction of neomycin B and related antibiotics with 30S and 50S ribosomal subunits. Biochem. Biophys. Res. Commun. 87, 960–966 (1979).
Carter, A.P. et al. Functional insights from the structure of the 30S ribosomal subunit and its interactions with antibiotics. Nature 407, 340–348 (2000).
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006).
Schuwirth, B.S. et al. Structures of the bacterial ribosome at 3.5 Å resolution. Science 310, 827–834 (2005).
Moazed, D. & Noller, H.F. Interaction of antibiotics with functional sites in 16S ribosomal RNA. Nature 327, 389–394 (1987).
Yusupov, M.M. et al. Crystal structure of the ribosome at 5.5 Å resolution. Science 292, 883–896 (2001).
Cannone, J.J. et al. The comparative RNA web (CRW) site: an online database of comparative sequence and structure information for ribosomal, intron, and other RNAs. BMC Bioinformatics 3, 2 (2002).
Lancaster, L., Kiel, M.C., Kaji, A. & Noller, H.F. Orientation of ribosome recycling factor in the ribosome from directed hydroxyl radical probing. Cell 111, 129–140 (2002).
Valle, M. et al. Locking and unlocking of ribosomal motions. Cell 114, 123–134 (2003).
Agrawal, R.K. et al. Visualization of ribosome-recycling factor on the Escherichia coli 70S ribosome: functional implications. Proc. Natl. Acad. Sci. USA 101, 8900–8905 (2004).
Connell, S.R. et al. Structural basis for interaction of the ribosome with the switch regions of GTP-bound elongation factors. Mol. Cell 25, 751–764 (2007).
Raj, V.S., Kaji, H. & Kaji, A. Interaction of RRF and EF-G from E. coli and T. thermophilus with ribosomes from both origins—insight into the mechanism of the ribosome recycling step. RNA 11, 275–284 (2005).
Wilson, D.N. et al. X-ray crystallography study on ribosome recycling: the mechanism of binding and action of RRF on the 50S ribosomal subunit. EMBO J. 24, 251–260 (2005).
Ali, I.K., Lancaster, L., Feinberg, J., Joseph, S. & Noller, H.F. Deletion of a conserved, central ribosomal intersubunit RNA bridge. Mol. Cell 23, 865–874 (2006).
Liiv, A., Karitkina, D., Maiväli, U. & Remme, J. Analysis of the function of E. coli 23S rRNA helix-loop 69 by mutagenesis. BMC Mol. Biol. 6, 18 (2005).
Rackham, O., Wang, K. & Chin, J.W. Functional epitopes at the ribosome subunit interface. Nat. Chem. Biol. 2, 254–258 (2006).
Frank, J. & Agrawal, R.K. A ratchet-like intersubunit reorganization of the ribosome during translocation. Nature 406, 318–322 (2000).
Horan, L.H. & Noller, H.F. Intersubunit movement is required for ribosomal translocation. Proc. Natl. Acad. Sci. USA 104, 4881–4885 (2007).
Campuzano, S., Vázquez, D. & Modolell, J. Dissociation of guanosine nucleotide-elongation factor G-ribosome complexes. Biochemistry 18, 1570–1574 (1979).
Senior, K. Duchenne muscular dystrophy improved by gentamicin. Mol. Med. Today 5, 461 (1999).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Yoshizawa, S., Fourmy, D. & Puglisi, J.D. Structural origins of gentamicin antibiotic action. EMBO J. 17, 6437–6448 (1998).
Cowtan, K. General quadratic functions in real and reciprocal space and their application to likelihood phasing. Acta Crystallogr. D Biol. Crystallogr. 56, 1612–1621 (2000).
Kleywegt, G.J. & Jones, T.A. Databases in protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 54, 1119–1131 (1998).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Toyoda, T. et al. Crystal structure combined with genetic analysis of the Thermus thermophilus ribosome recycling factor shows that a flexible hinge may act as a functional switch. RNA 6, 1432–1444 (2000).
Carson, M. Ribbons. Methods Enzymol. 277, 493–502 (1997).
Acknowledgements
We thank K. Frankel, S. Classen and G. Meigs for help with data measurement at the SIBYLS and 8.3.1 beamlines at the Advanced Light Source; K. Rajashankar, I. Kourinov and N. Sukumar for help with data measurement at Northeastern Collaborative Access Team beamline 24-IDC at the Advanced Photon Source; W. Wintermeyer, M. O'Connor and K. Fredrick for helpful discussions; A. Borovinskiy for help with figure preparation and overall support; and J. Doudna, H. Noller, P. Gunda and J. Dunkle for helpful comments on the manuscript. This work was supported by US National Institutes of Health (NIH) grants GM65050 (J.H.D.C.) and GM60429 (A.K.), the Nippon Paint Fund (H.K.), NIH National Cancer Institute grant CA92584 (for the SIBYLS and 8.3.1 beamlines), NIH National Center for Research Resources grant RR-15301 (for beamline 24-IDC) and US Department of Energy grants DE-AC03-76SF00098, KP110201, and LBNL LDRD 366851 (J.H.D.C.), DE-AC03 76SF00098 (for the SIBYLS and 8.3.1 beamlines) and DE-AC02-06CH11357 (for the Advanced Photon Source).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Methods and Supplementary Data (PDF 7354 kb)
Rights and permissions
About this article
Cite this article
Borovinskaya, M., Pai, R., Zhang, W. et al. Structural basis for aminoglycoside inhibition of bacterial ribosome recycling. Nat Struct Mol Biol 14, 727–732 (2007). https://doi.org/10.1038/nsmb1271
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1271
This article is cited by
-
Genome-wide identification of Kanamycin B binding RNA in Escherichia coli
BMC Genomics (2023)
-
Molecular basis of the pleiotropic effects by the antibiotic amikacin on the ribosome
Nature Communications (2023)
-
Geometric alignment of aminoacyl-tRNA relative to catalytic centers of the ribosome underpins accurate mRNA decoding
Nature Communications (2023)
-
Antibiotic thermorubin tethers ribosomal subunits and impedes A-site interactions to perturb protein synthesis in bacteria
Nature Communications (2023)
-
Classification of antimicrobial mechanism of action using dynamic bacterial morphology imaging
Scientific Reports (2022)