Abstract
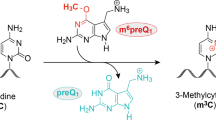
A previous bioinformatics-based search for riboswitches yielded several candidate motifs in eubacteria. One of these motifs commonly resides in the 5′ untranslated regions of genes involved in the biosynthesis of queuosine (Q), a hypermodified nucleoside occupying the anticodon wobble position of certain transfer RNAs. Here we show that this structured RNA is part of a riboswitch selective for 7-aminomethyl-7-deazaguanine (preQ1), an intermediate in queuosine biosynthesis. Compared with other natural metabolite-binding RNAs, the preQ1 aptamer appears to have a simple structure, consisting of a single stem-loop and a short tail sequence that together are formed from as few as 34 nucleotides. Despite its small size, this aptamer is highly selective for its cognate ligand in vitro and has an affinity for preQ1 in the low nanomolar range. Relatively compact RNA structures can therefore serve effectively as metabolite receptors to regulate gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Ptashne, M. & Gann, A. Genes and Signals (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2002).
Mandal, M. & Breaker, R.R. Gene regulation by riboswitches. Nat. Rev. Mol. Cell Biol. 5, 451–463 (2004).
Soukup, J.K. & Soukup, G.A. Riboswitches exert genetic control through metabolite-induced conformational change. Curr. Opin. Struct. Biol. 14, 344–349 (2004).
Winkler, W.C. Riboswitches and the role of noncoding RNAs in bacterial metabolic control. Curr. Opin. Chem. Biol. 9, 594–602 (2005).
Winkler, W.C. & Breaker, R.R. Regulation of bacterial gene expression by riboswitches. Annu. Rev. Microbiol. 59, 487–517 (2005).
Breaker, R.R. in The RNA World 3rd edn. (eds. Gesteland, R.F., Cech, T.R. & Atkins, J.F.) 89–107 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2006).
Barrick, J.E. et al. New RNA motifs suggest an expanded scope for riboswitches in bacterial genetic control. Proc. Natl. Acad. Sci. USA 101, 6421–6426 (2004).
Corbino, K.A. et al. Evidence for a second class of S-adenosylmethionine riboswitches and other regulatory RNA motifs in alpha-proteobacteria. Genome Biol. 6, R70 (2005).
Reader, J.S., Metzgar, D., Schimmel, P. & de Crécy-Lagard, V. Identification of four genes necessary for biosynthesis of the modified nucleoside queuosine. J. Biol. Chem. 279, 6280–6285 (2004).
Harada, F. & Nishimura, S. Possible anticodon sequences of tRNAHis, tRNAAsn, and tRNAAsp from Escherichia coli B. Universal presence of nucleoside Q in the first position of the anticodons of these transfer ribonucleic acids. Biochemistry 11, 301–308 (1972).
Van Lanen, S.G. et al. From cyclohydrolase to oxidoreductase: discovery of nitrile reductase activity in a common fold. Proc. Natl. Acad. Sci. USA 102, 4264–4269 (2005).
Gaur, R. & Varshney, U. Genetic analysis identifies a function for the queC (ybaX) gene product at an initial step in the queuosine biosynthetic pathway in Escherichia coli. J. Bacteriol. 187, 6893–6901 (2005).
Okada, N. et al. Novel mechanism of post-transcriptional modification of tRNA. Insertion of bases of Q precursors into tRNA by a specific tRNA transglycosylase reaction. J. Biol. Chem. 254, 3067–3073 (1979).
Reuter, K., Slany, R., Ullrich, F. & Kersten, H. Structure and organization of Escherichia coli genes involved in biosynthesis of the deazaguanine derivative queuine, a nutrient factor for eukaryotes. J. Bacteriol. 173, 2256–2264 (1991).
Salazar, J.C., Ambrogelly, A., Crain, P.F., McCloskey, J.A. & Soll, D. A truncated aminoacyl-tRNA synthetase modifies RNA. Proc. Natl. Acad. Sci. USA 101, 7536–7541 (2004).
Blaise, M. et al. A minimalist glutamyl-tRNA synthetase dedicated to aminoacylation of the tRNAAsp QUC anticodon. Nucleic Acids Res. 32, 2768–2775 (2004).
Iwata-Reuyl, D. Biosynthesis of the 7-deazaguanosine hypermodified nucleosides of transfer RNA. Bioorg. Chem. 31, 24–43 (2003).
Durand, J.M. et al. vacC, a virulence-associated chromosomal locus of Shigella flexneri, is homologous to tgt, a gene encoding tRNA-guanine transglycosylase (Tgt) of Escherichia coli K-12. J. Bacteriol. 176, 4627–4634 (1994).
Noguchi, S., Nishimura, Y., Hirota, Y. & Nishimura, S. Isolation and characterization of an Escherichia coli mutant lacking tRNA-guanine transglycosylase. Function and biosynthesis of queuosine in tRNA. J. Biol. Chem. 257, 6544–6550 (1982).
Frey, B., Janel, G., Michelsen, U. & Kersten, H. Mutations in the Escherichia coli fnr and tgt genes: control of molybdate reductase activity and the cytochrome d complex by fnr. J. Bacteriol. 171, 1524–1530 (1989).
Bienz, M. & Kubli, E. Wild type tRNATyr reads the TMV RNA stop codon, but Q base-modified tRNATyr does not. Nature 294, 188–190 (1981).
Meier, F., Suter, B., Grosjean, H., Keith, G. & Kubli, E. Queuosine modification of the wobble base in tRNAHis influences in vivo decoding properties. EMBO J. 4, 823–827 (1985).
Urbonavičius, J., Qian, Q., Durand, J.M., Hagervall, T.G. & Björk, G.R. Improvement of reading frame maintenance is a common function for several tRNA modifications. EMBO J. 20, 4863–4873 (2001).
Kuchino, Y., Kasai, H., Nihei, K. & Nishimura, S. Biosynthesis of the modified nucleoside Q in transfer RNA. Nucleic Acids Res. 3, 393–398 (1976).
Okada, N. et al. Structure determination of a nucleoside Q precursor isolated from E. coli tRNA: 7-(aminomethyl)-7-deazaguanosine. Nucleic Acids Res. 5, 2289–2296 (1978).
Slany, R.K., Bosl, M., Crain, P.F. & Kersten, H. A new function of S-adenosylmethionine: the ribosyl moiety of AdoMet is the precursor of the cyclopentenediol moiety of the tRNA wobble base queuine. Biochemistry 32, 7811–7817 (1993).
Slany, R.K., Bosl, M. & Kersten, H. Transfer and isomerization of the ribose moiety of AdoMet during the biosynthesis of queuosine tRNAs, a new unique reaction catalyzed by the QueA protein from Escherichia coli. Biochimie 76, 389–393 (1994).
Frey, B., McCloskey, J., Kersten, W. & Kersten, H. New function of vitamin B12: cobamide-dependent reduction of epoxyqueuosine to queuosine in tRNAs of Escherichia coli and Salmonella typhimurium. J. Bacteriol. 170, 2078–2082 (1988).
Gusarov, I. & Nudler, E. The mechanism of intrinsic transcription termination. Mol. Cell 3, 495–504 (1999).
Yarnell, W.S. & Roberts, J.W. Mechanism of intrinsic transcription termination and antitermination. Science 284, 611–615 (1999).
Soukup, G.A. & Breaker, R.R. Relationship between internucleotide linkage geometry and the stability of RNA. RNA 5, 1308–1325 (1999).
Wickiser, J.K., Winkler, W.C., Breaker, R.R. & Crothers, D.M. The speed of RNA transcription and metabolite binding kinetics operate an FMN riboswitch. Mol. Cell 18, 49–60 (2005).
Wickiser, J.K., Cheah, M.T., Breaker, R.R. & Crothers, D.M. The kinetics of ligand binding by an adenine-sensing riboswitch. Biochemistry 44, 13404–13414 (2005).
Lemay, J.F., Penedo, J.C., Tremblay, R., Lilley, D.M. & Lafontaine, D.A. Folding of the adenine riboswitch. Chem. Biol. 13, 857–868 (2006).
Gilbert, S.D., Stoddard, C.D., Wise, S.J. & Batey, R.T. Thermodynamic and kinetic characterization of ligand binding to the purine riboswitch aptamer domain. J. Mol. Biol. 359, 754–768 (2006).
Mandal, M. & Breaker, R.R. Adenine riboswitches and gene activation by disruption of a transcription terminator. Nat. Struct. Mol. Biol. 11, 29–35 (2004).
Batey, R.T., Gilbert, S.D. & Montange, R.K. Structure of a natural guanine-responsive riboswitch complexed with the metabolite hypoxanthine. Nature 432, 411–415 (2004).
Serganov, A. et al. Structural basis for discriminative regulation of gene expression by adenine- and guanine-sensing mRNAs. Chem. Biol. 11, 1729–1741 (2004).
Noeske, J. et al. An intermolecular base triple as the basis of ligand specificity and affinity in the guanine- and adenine-sensing riboswitch RNAs. Proc. Natl. Acad. Sci. USA 102, 1372–1377 (2005).
Fuchs, R.T., Grundy, F.J. & Henkin, T.M. The S(MK) box is a new SAM-binding RNA for translational regulation of SAM synthetase. Nat. Struct. Mol. Biol. 13, 226–233 (2006).
Tatusov, R.L. et al. The COG database: new developments in phylogenetic classification of proteins from complete genomes. Nucleic Acids Res. 29, 22–28 (2001).
Dewey, V. & Kidder, G.W. Partial purification and properties of a nucleoside hydrolase from Crithidia. Arch. Biochem. Biophys. 157, 380–387 (1973).
Miller, R.L., Sabourin, C.L., Krenitsky, T.A., Berens, R.L. & Marr, J.J. Nucleoside hydrolases from Trypanosoma cruzi. J. Biol. Chem. 259, 5073–5077 (1984).
Gündüz, U. & Katze, J.R. Salvage of the nucleic acid base queuine from queuine-containing tRNA by animal cells. Biochem. Biophys. Res. Commun. 109, 159–167 (1982).
Gündüz, U. & Katze, J.R. Queuine salvage in mammalian cells. Evidence that queuine is generated from queuosine 5′-phosphate. J. Biol. Chem. 259, 1110–1113 (1984).
Nissen, P., Ippolito, J.A., Ban, N., Moore, P.B. & Steitz, T.A. RNA tertiary interactions in the large ribosomal subunit: the A-minor motif. Proc. Natl. Acad. Sci. USA 98, 4899–4903 (2001).
Akimoto, H., Imamiya, E., Hitaka, T., Nomura, H. & Nishimura, S. Synthesis of queuine, the base of naturally occurring hypermodified nucleoside (queuosine), and its analogues. J. Chem. Soc. [Perkin 1] 1988, 1637–1644 (1988).
Migawa, M.T., Hinkley, J.M., Hoops, G.C. & Townsend, L.B. A two step synthesis of the nucleoside Q precursor 2-amino-5-cyanopyrrolo[2,3-d]pyrimidine-4-one (preQ0). Synth. Commun. 26, 3317–3322 (1996).
Akimoto, H. et al. Queuine analogues. Their synthesis and inhibition of growth of mouse L5178Y cells in vitro. J. Med. Chem. 29, 1749–1753 (1986).
West, R.A. 4-Hydroxypyrrolo[2,3-d]pyrimidine: Mannich reaction. J. Org. Chem. 26, 4959–4961 (1961).
Wellcome Foundation, Ltd. Great Britain patent number 981458 (1965).
Eddy, S.R. & Durbin, R. RNA sequence analysis using covariance models. Nucleic Acids Res. 22, 2079–2088 (1994).
Weinberg, Z. & Ruzzo, W.L. Sequence-based heuristics for faster annotation of non-coding RNA families. Bioinformatics 22, 35–39 (2006).
Tatusov, R.L. et al. The COG database: an updated version includes eukaryotes. BMC Bioinformatics 4, 41 (2003).
Gerstein, M., Sonnhammer, E.L. & Chothia, C. Volume changes in protein evolution. J. Mol. Biol. 236, 1067–1078 (1994).
Guérout-Fleury, A.M., Frandsen, N. & Stragier, P. Plasmids for ectopic integration in Bacillus subtilis. Gene 180, 57–61 (1996).
Roth, A., Nahvi, A., Lee, M., Jona, I. & Breaker, R.R. Characteristics of the glmS ribozyme suggest only structural roles for divalent metal ions. RNA 12, 607–619 (2006).
Mandal, M., Boese, B., Barrick, J.E., Winkler, W.C. & Breaker, R.R. Riboswitches control fundamental biochemical pathways in Bacillus subtilis and other bacteria. Cell 113, 577–586 (2003).
Sudarsan, N., Wickiser, J.K., Nakamura, S., Ebert, M.S. & Breaker, R.R. An mRNA structure in bacteria that controls gene expression by binding lysine. Genes Dev. 17, 2688–2697 (2003).
Acknowledgements
We thank J.A. Collins for assistance in generating B. subtilis transformants and members of the Breaker laboratory for helpful discussions. This work was supported by grants from the US National Institutes of Health (R33 DK07027 and GM 068819) and the US National Science Foundation (EIA 0323510) to R.R.B. RNA research in the Breaker laboratory is also supported by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
A. Roth, W.C.W., E.E.R., I.J., A. Ritwik, J.N.K. and R.W. carried out in vitro biochemical analyses of queC RNA constructs. E.E.R. conducted assays of reporter gene expression in vivo. B.W.K.L., J.L. and D.I.-R. synthesized preQ1 and related compounds. J.E.B. performed bioinformatics searches and prepared sequence alignments. A. Roth and R.R.B. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.R.B. is a cofounder of BioRelix, a company interested in the use of riboswitches as drug targets. Although this publication is not directly related to the development of antimicrobials and thus does not currently constitute a competing financial interest, the discovery of novel riboswitches may ultimately lead to development of new classes of antibiotics.
Supplementary information
Supplementary Fig. 1
PreQ1 aptamer sequences. (PDF 128 kb)
Supplementary Fig. 2
PreQ1 riboswitch locations and associated genes. (PDF 164 kb)
Rights and permissions
About this article
Cite this article
Roth, A., Winkler, W., Regulski, E. et al. A riboswitch selective for the queuosine precursor preQ1 contains an unusually small aptamer domain. Nat Struct Mol Biol 14, 308–317 (2007). https://doi.org/10.1038/nsmb1224
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1224
This article is cited by
-
A small RNA that cooperatively senses two stacked metabolites in one pocket for gene control
Nature Communications (2022)
-
Engineered pegRNAs improve prime editing efficiency
Nature Biotechnology (2022)
-
A chemical probe based on the PreQ1 metabolite enables transcriptome-wide mapping of binding sites
Nature Communications (2021)
-
Development of a new oligonucleotide block location-based feature extraction (BLBFE) method for the classification of riboswitches
Molecular Genetics and Genomics (2020)
-
Synthetic ligands for PreQ1 riboswitches provide structural and mechanistic insights into targeting RNA tertiary structure
Nature Communications (2019)