Abstract
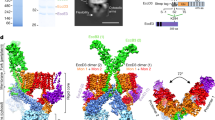
The type III secretion system (T3SS) ATPase is the conserved and essential inner-membrane component involved in the initial stages of selective secretion of specialized T3SS virulence effector proteins from the bacterial cytoplasm through to the infected host cell, a process crucial to subsequent pathogenicity. Here we present the 1.8-Å-resolution crystal structure of the catalytic domain of the prototypical T3SS ATPase EscN from enteropathogenic Escherichia coli (EPEC). Along with in vitro and in vivo mutational analysis, our data show that the T3SS ATPases share similarity with the F1 ATPases but have important structural and sequence differences that dictate their unique secretory role. We also show that T3SS ATPase activity is dependent on EscN oligomerization and describe the molecular features and possible functional implications of a hexameric ring model.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hueck, C.J. Type III protein secretion systems in bacterial pathogens of animals and plants. Microbiol. Mol. Biol. Rev. 62, 379–433 (1998).
Gophna, U., Ron, E.Z. & Graur, D. Bacterial type III secretion systems are ancient and evolved by multiple horizontal-transfer events. Gene 312, 151–163 (2003).
Bennett, J.C. & Hughes, C. From flagellum assembly to virulence: the extended family of type III export chaperones. Trends Microbiol. 8, 202–204 (2000).
Luo, Y. et al. Structural and biochemical characterization of the type III secretion chaperones CesT and SigE. Nat. Struct. Biol. 8, 1031–1036 (2001).
Thomas, N.A. et al. CesT is a multi-effector chaperone and recruitment factor required for the efficient type III secretion of both LEE- and non-LEE-encoded effectors of enteropathogenic Escherichia coli. Mol. Microbiol. 57, 1762–1779 (2005).
Ghosh, P. Process of protein transport by the type III secretion system. Microbiol. Mol. Biol. Rev. 68, 771–795 (2004).
Akeda, Y. & Galan, J.E. Genetic analysis of the Salmonella enterica type III secretion-associated ATPase InvC defines discrete functional domains. J. Bacteriol. 186, 2402–2412 (2004).
Akeda, Y. & Galan, J.E. Chaperone release and unfolding of substrates in type III secretion. Nature 437, 911–915 (2005).
Jenks, P.J. et al. A flagellar-specific ATPase (FliI) is necessary for flagellar export in Helicobacter pylori. FEMS Microbiol. Lett. 152, 205–211 (1997).
Pozidis, C. et al. Type III protein translocase: HrcN is a peripheral ATPase that is activated by oligomerization. J. Biol. Chem. 278, 25816–25824 (2003).
Woestyn, S., Allaoui, A., Wattiau, P. & Cornelis, G.R. YscN, the putative energizer of the Yersinia Yop secretion machinery. J. Bacteriol. 176, 1561–1569 (1994).
Schubot, F.D. & Waugh, D.S. A pivotal role for reductive methylation in the de novo crystallization of a ternary complex composed of Yersinia pestis virulence factors YopN, SycN and YscB. Acta Crystallogr. D Biol. Crystallogr. 60, 1981–1986 (2004).
Minamino, T., Gonzalez-Pedrajo, B., Kihara, M., Namba, K. & Macnab, R.M. The ATPase FliI can interact with the type III flagellar protein export apparatus in the absence of its regulator, FliH. J. Bacteriol. 185, 3983–3988 (2003).
Pallen, M.J., Bailey, C.M. & Beatson, S.A. Evolutionary links between FliH/YscL-like proteins from bacterial type III secretion systems and second-stalk components of the FoF1 and vacuolar ATPases. Protein Sci. 15, 935–941 (2006).
Thomas, J., Stafford, G.P. & Hughes, C. Docking of cytosolic chaperone-substrate complexes at the membrane ATPase during flagellar type III protein export. Proc. Natl. Acad. Sci. USA 101, 3945–3950 (2004).
Zhu, K., Gonzalez-Pedrajo, B. & Macnab, R.M. Interactions among membrane and soluble components of the flagellar export apparatus of Salmonella. Biochemistry 41, 9516–9524 (2002).
Walker, J.E., Saraste, M., Runswick, M.J. & Gay, N.J. Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J. 1, 945–951 (1982).
Abrahams, J.P., Leslie, A.G., Lutter, R. & Walker, J.E. Structure at 2.8 A resolution of F1-ATPase from bovine heart mitochondria. Nature 370, 621–628 (1994).
Groth, G. Structure of spinach chloroplast F1-ATPase complexed with the phytopathogenic inhibitor tentoxin. Proc. Natl. Acad. Sci. USA 99, 3464–3468 (2002).
Shirakihara, Y. et al. The crystal structure of the nucleotide-free alpha 3 beta 3 subcomplex of F1-ATPase from the thermophilic Bacillus PS3 is a symmetric trimer. Structure 5, 825–836 (1997).
Claret, L., Calder, S.R., Higgins, M. & Hughes, C. Oligomerization and activation of the FliI ATPase central to bacterial flagellum assembly. Mol. Microbiol. 48, 1349–1355 (2003).
Nadanaciva, S., Weber, J., Wilke-Mounts, S. & Senior, A.E. Importance of F1-ATPase residue alpha-Arg-376 for catalytic transition state stabilization. Biochemistry 38, 15493–15499 (1999).
Ogura, T. & Wilkinson, A.J. AAA+ superfamily ATPases: common structure–diverse function. Genes Cells 6, 575–597 (2001).
Gibbons, C., Montgomery, M.G., Leslie, A.G. & Walker, J.E. The structure of the central stalk in bovine F(1)-ATPase at 2.4 A resolution. Nat. Struct. Biol. 7, 1055–1061 (2000).
Abe, A. Passage of bacterial effectors into the host cell. Tanpakushitsu Kakusan Koso 49, 967–970 (2004).
Gauthier, A. & Finlay, B.B. Translocated intimin receptor and its chaperone interact with ATPase of the type III secretion apparatus of enteropathogenic Escherichia coli. J. Bacteriol. 185, 6747–6755 (2003).
Gauthier, A., Puente, J.L. & Finlay, B.B. Secretin of the enteropathogenic Escherichia coli type III secretion system requires components of the type III apparatus for assembly and localization. Infect. Immun. 71, 3310–3319 (2003).
Minamino, T. & MacNab, R.M. Interactions among components of the Salmonella flagellar export apparatus and its substrates. Mol. Microbiol. 35, 1052–1064 (2000).
Ramamurthi, K.S. & Schneewind, O. Type III protein secretion in yersinia species. Annu. Rev. Cell Dev. Biol. 18, 107–133 (2002).
Auvray, F., Ozin, A.J., Claret, L. & Hughes, C. Intrinsic membrane targeting of the flagellar export ATPase FliI: interaction with acidic phospholipids and FliH. J. Mol. Biol. 318, 941–950 (2002).
Yonekura, K., Maki-Yonekura, S. & Namba, K. Complete atomic model of the bacterial flagellar filament by electron cryomicroscopy. Nature 424, 643–650 (2003).
Minamino, T. & MacNab, R.M. FliH, a soluble component of the type III flagellar export apparatus of Salmonella, forms a complex with FliI and inhibits its ATPase activity. Mol. Microbiol. 37, 1494–1503 (2000).
Guex, N., Diemand, A. & Peitsch, M.C. Protein modelling for all. Trends Biochem. Sci. 24, 364–367 (1999).
DeLano, W.L. The PyMOL Molecular Graphics System 0.99 edn. (DeLano Scientific, San Carlos, 2002).
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Schneider, T.R. & Sheldrick, G. Substructure solution with SHELXD. Acta Crystallogr. D Biol. Crystallogr. 58, 1772–1779 (2002).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Deng, W. et al. Regulation of type III secretion hierarchy of translocators and effectors in attaching and effacing bacterial pathogens. Infect. Immun. 73, 2135–2146 (2005).
Schubot, F.D. et al. Three-dimensional structure of a macromolecular assembly that regulates type III secretion in Yersinia pestis. J. Mol. Biol. 346, 1147–1161 (2005).
Acknowledgements
We thank A.L. Lovering, C.K. Yip and P.I. Lario for discussions, C.K. Yip for performing the static light-scattering analysis, N.A. Thomas for reagents and the staff at Advanced Light Source beamline 8.2.2 for data collection time and assistance. R.Z. is supported by postdoctoral fellowships from Izaak Walton Killam Research, the Michael Smith Foundation for Health Research (MSFHR) and the Canadian Institutes of Health Research (CIHR). N.C.J.S. and B.B.F. thank the Howard Hughes International Scholar program and the CIHR for funding. N.C.J.S. also thanks the MSFHR and the Canada Foundation for Innovation for infrastructure funding support. N.C.J.S. is an MSFHR Senior Scholar and CIHR Investigator.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
EscN structure. (PDF 1305 kb)
Supplementary Fig. 2
Multiple sequence alignment. (PDF 1766 kb)
Supplementary Table 1
Structural comparison table. (PDF 2042 kb)
Rights and permissions
About this article
Cite this article
Zarivach, R., Vuckovic, M., Deng, W. et al. Structural analysis of a prototypical ATPase from the type III secretion system. Nat Struct Mol Biol 14, 131–137 (2007). https://doi.org/10.1038/nsmb1196
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1196
This article is cited by
-
Enhanced protein translocation to mammalian cells by expression of EtgA transglycosylase in a synthetic injector E. coli strain
Microbial Cell Factories (2022)
-
Cryo-EM analysis of the T3S injectisome reveals the structure of the needle and open secretin
Nature Communications (2018)
-
Assembly, structure, function and regulation of type III secretion systems
Nature Reviews Microbiology (2017)
-
Disease-Homologous Mutation in the Cation Diffusion Facilitator Protein MamM Causes Single-Domain Structural Loss and Signifies Its Importance
Scientific Reports (2016)
-
Secretion systems in Gram-negative bacteria: structural and mechanistic insights
Nature Reviews Microbiology (2015)