Abstract
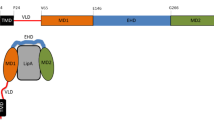
Secretion via the type II secretion pathway in Gram-negative bacteria often relies crucially on steric chaperones in the periplasm. Here, we report the crystal structure of the soluble form of a lipase-specific foldase (Lif) from Burkholderia glumae in complex with its cognate lipase. The structure reveals how Lif uses a novel α-helical scaffold to embrace lipase, thereby creating an unusually extensive folding platform.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Ellis, R.J. Trends Biochem. Sci. 23, 43–45 (1998).
Shinde, U.P., Liu, J.J. & Inouye, M. Nature 389, 520–522 (1997).
Baker, D., Sohl, J.L. & Agard, D.A. Nature 356, 263–265 (1992).
Filloux, A. Biochim. Biophys. Acta 1694, 163–179 (2004).
Frenken, L.G.J., de Groot, A., Tommassen, J. & Verrips, C.T. Mol. Microbiol. 9, 591–599 (1993).
El Khattabi, M., Van Gelder, P., Bitter, W. & Tommassen, J. J. Biol. Chem. 275, 26885–26891 (2000).
Pauwels, K. et al. Acta Crystallogr. F 61, 791–795 (2005).
Rosenau, F., Tommassen, J. & Jaeger, K.-E. ChemBioChem. 5, 152–161 (2004).
El Khattabi, M. et al. Mol. Gen. Genet. 261, 770–776 (1999).
Shibata, H., Kato, H. & Oda, J. Biosci. Biotechnol. Biochem. 62, 354–357 (1998).
Noble, M.E.M. et al. FEBS Lett. 331, 123–128 (1993).
Wodak, S.J. & Janin, J. Adv. Protein Chem. 61, 9–73 (2002).
Acknowledgements
We acknowledge the use of synchrotron beam time at the European Molecular Biology Laboratory beamlines (Deutsches Elektronen-Synchrotron and European Synchrotron Radiation Facility). We thank G. Machaidze, U. Aebi and E. De Genst for help with analytical ultracentrifugation and surface plasmon resonance experiments, respectively, and L. Buts, K. Decanniere and R. Loris for useful discussions. K. Pauwels is supported by the Institute for the Promotion of Innovation by Science and Technology in Flanders. This research was sponsored by the European Commission-FP5 (QLRT-2001-02086).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Analytical gel filtration (PDF 65 kb)
Supplementary Fig. 2
Sequence alignment of selected members of the Lif superfamily and comparison with secondary structure elements in Lif from B. glumae (PDF 166 kb)
Supplementary Fig. 3
Near-UV CD spectra of Lif and native LipA (PDF 18 kb)
Supplementary Table 1
Sedimentation velocity and sedimentation equilibrium experiments of Lif and LipA (PDF 13 kb)
Supplementary Table 2
Data collection and crystallographic refinement statistics (PDF 42 kb)
Supplementary Table 3
Conserved residues of Lif interacting with conserved residues in LipA (PDF 16 kb)
Rights and permissions
About this article
Cite this article
Pauwels, K., Lustig, A., Wyns, L. et al. Structure of a membrane-based steric chaperone in complex with its lipase substrate. Nat Struct Mol Biol 13, 374–375 (2006). https://doi.org/10.1038/nsmb1065
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1065
This article is cited by
-
Enzymatic transesterification of urethane-bond containing ester
Colloid and Polymer Science (2021)
-
Structural and dynamic insights revealing how lipase binding domain MD1 of Pseudomonas aeruginosa foldase affects lipase activation
Scientific Reports (2020)
-
First co-expression of a lipase and its specific foldase obtained by metagenomics
Microbial Cell Factories (2014)
-
The type II secretion system: biogenesis, molecular architecture and mechanism
Nature Reviews Microbiology (2012)
-
Structural characterization of a ribose-5-phosphate isomerase B from the pathogenic fungus Coccidioides immitis
BMC Structural Biology (2011)