Abstract
Type A γ-aminobutyric acid receptors (GABAARs) are the principal mediators of inhibitory neurotransmission in the human brain. Endogenous neurosteroids interact with GABAARs to regulate acute and chronic anxiety and are potent sedative, analgesic, anticonvulsant and anesthetic agents. Their mode of binding and mechanism of receptor potentiation, however, remain unknown. Here we report crystal structures of a chimeric GABAAR construct in apo and pregnanolone-bound states. The neurosteroid-binding site is mechanically coupled to the helices lining the ion channel pore and modulates the desensitization-gate conformation. We demonstrate that the equivalent site is responsible for physiological, heteromeric GABAAR potentiation and explain the contrasting modulatory properties of 3a versus 3b neurosteroid epimers. These results illustrate how peripheral lipid ligands can regulate the desensitization gate of GABAARs, a process of broad relevance to pentameric ligand-gated ion channels.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Nemecz, Á., Prevost, M.S., Menny, A. & Corringer, P.J. Emerging molecular mechanisms of signal transduction in pentameric ligand-gated ion channels. Neuron 90, 452–470 (2016).
Miller, P.S. & Smart, T.G. Binding, activation and modulation of Cys-loop receptors. Trends Pharmacol. Sci. 31, 161–174 (2010).
Hosie, A.M., Wilkins, M.E., da Silva, H.M. & Smart, T.G. Endogenous neurosteroids regulate GABAA receptors through two discrete transmembrane sites. Nature 444, 486–489 (2006).
Majewska, M.D., Harrison, N.L., Schwartz, R.D., Barker, J.L. & Paul, S.M. Steroid hormone metabolites are barbiturate-like modulators of the GABA receptor. Science 232, 1004–1007 (1986).
Sarkar, J., Wakefield, S., MacKenzie, G., Moss, S.J. & Maguire, J. Neurosteroidogenesis is required for the physiological response to stress: role of neurosteroid-sensitive GABAA receptors. J. Neurosci. 31, 18198–18210 (2011).
Gunn, B.G., Brown, A.R., Lambert, J.J. & Belelli, D. Neurosteroids and GABA(A) Receptor Interactions: A Focus on Stress. Front. Neurosci. 5, 131 (2011).
Maguire, J.L., Stell, B.M., Rafizadeh, M. & Mody, I. Ovarian cycle-linked changes in GABA(A) receptors mediating tonic inhibition alter seizure susceptibility and anxiety. Nat. Neurosci. 8, 797–804 (2005).
Maguire, J. & Mody, I. GABA(A)R plasticity during pregnancy: relevance to postpartum depression. Neuron 59, 207–213 (2008).
Sigel, E. & Steinmann, M.E. Structure, function, and modulation of GABA(A) receptors. J. Biol. Chem. 287, 40224–40231 (2012).
Miller, P.S. & Aricescu, A.R. Crystal structure of a human GABAA receptor. Nature 512, 270–275 (2014).
Akk, G. et al. Mutations of the GABA-A receptor alpha1 subunit M1 domain reveal unexpected complexity for modulation by neuroactive steroids. Mol. Pharmacol. 74, 614–627 (2008).
Saras, A. et al. Histamine action on vertebrate GABAA receptors: direct channel gating and potentiation of GABA responses. J. Biol. Chem. 283, 10470–10475 (2008).
Bianchi, M.T. & Macdonald, R.L. Neurosteroids shift partial agonist activation of GABA(A) receptor channels from low- to high-efficacy gating patterns. J. Neurosci. 23, 10934–10943 (2003).
Hassaine, G. et al. X-ray structure of the mouse serotonin 5-HT3 receptor. Nature 512, 276–281 (2014).
Miyazawa, A., Fujiyoshi, Y. & Unwin, N. Structure and gating mechanism of the acetylcholine receptor pore. Nature 423, 949–955 (2003).
Du, J., Lü, W., Wu, S., Cheng, Y. & Gouaux, E. Glycine receptor mechanism elucidated by electron cryo-microscopy. Nature 526, 224–229 (2015).
Huang, X., Chen, H., Michelsen, K., Schneider, S. & Shaffer, P.L. Crystal structure of human glycine receptor-α3 bound to antagonist strychnine. Nature 526, 277–280 (2015).
Althoff, T., Hibbs, R.E., Banerjee, S. & Gouaux, E. X-ray structures of GluCl in apo states reveal a gating mechanism of Cys-loop receptors. Nature 512, 333–337 (2014).
Hibbs, R.E. & Gouaux, E. Principles of activation and permeation in an anion-selective Cys-loop receptor. Nature 474, 54–60 (2011).
Unwin, N. Refined structure of the nicotinic acetylcholine receptor at 4A resolution. J. Mol. Biol. 346, 967–989 (2005).
Morales-Perez, C.L., Noviello, C.M. & Hibbs, R.E. X-ray structure of the human α4β2 nicotinic receptor. Nature 538, 411–415 (2016).
Hori-Tanaka, Y., Yura, K., Takai-Igarashi, T. & Tanaka, H. Structural classification of steroid-binding sites on proteins by coarse-grained atomic environment and its correlation with their biological function. Steroids 96, 81–88 (2015).
Bracamontes, J.R., Li, P., Akk, G. & Steinbach, J.H. A neurosteroid potentiation site can be moved among GABAA receptor subunits. J. Physiol. (Lond.) 590, 5739–5747 (2012).
Chen, Z.W. et al. Neurosteroid analog photolabeling of a site in the third transmembrane domain of the β3 subunit of the GABA(A) receptor. Mol. Pharmacol. 82, 408–419 (2012).
Belelli, D., Casula, A., Ling, A. & Lambert, J.J. The influence of subunit composition on the interaction of neurosteroids with GABA(A) receptors. Neuropharmacology 43, 651–661 (2002).
Cho, A.E., Guallar, V., Berne, B.J. & Friesner, R. Importance of accurate charges in molecular docking: quantum mechanical/molecular mechanical (QM/MM) approach. J. Comput. Chem. 26, 915–931 (2005).
Eick, G.N. & Thornton, J.W. Evolution of steroid receptors from an estrogen-sensitive ancestral receptor. Mol. Cell. Endocrinol. 334, 31–38 (2011).
Wang, M. et al. 3beta-hydroxypregnane steroids are pregnenolone sulfate-like GABA(A) receptor antagonists. J. Neurosci. 22, 3366–3375 (2002).
Aryal, P., Sansom, M.S. & Tucker, S.J. Hydrophobic gating in ion channels. J. Mol. Biol. 427, 121–130 (2015).
Unwin, N. & Fujiyoshi, Y. Gating movement of acetylcholine receptor caught by plunge-freezing. J. Mol. Biol. 422, 617–634 (2012).
Gielen, M., Thomas, P. & Smart, T.G. The desensitization gate of inhibitory Cys-loop receptors. Nat. Commun. 6, 6829 (2015).
Brickley, S.G. & Mody, I. Extrasynaptic GABA(A) receptors: their function in the CNS and implications for disease. Neuron 73, 23–34 (2012).
Jansen, M., Bali, M. & Akabas, M.H. Modular design of Cys-loop ligand-gated ion channels: functional 5-HT3 and GABA rho1 receptors lacking the large cytoplasmic M3M4 loop. J. Gen. Physiol. 131, 137–146 (2008).
Aricescu, A.R., Lu, W. & Jones, E.Y. A time- and cost-efficient system for high-level protein production in mammalian cells. Acta Crystallogr. D Biol. Crystallogr. 62, 1243–1250 (2006).
Molday, R.S. & MacKenzie, D. Monoclonal antibodies to rhodopsin: characterization, cross-reactivity, and application as structural probes. Biochemistry 22, 653–660 (1983).
Oprian, D.D., Molday, R.S., Kaufman, R.J. & Khorana, H.G. Expression of a synthetic bovine rhodopsin gene in monkey kidney cells. Proc. Natl. Acad. Sci. USA 84, 8874–8878 (1987).
Reeves, P.J., Callewaert, N., Contreras, R. & Khorana, H.G. Structure and function in rhodopsin: high-level expression of rhodopsin with restricted and homogeneous N-glycosylation by a tetracycline-inducible N-acetylglucosaminyltransferase I-negative HEK293S stable mammalian cell line. Proc. Natl. Acad. Sci. USA 99, 13419–13424 (2002).
Aricescu, A.R. & Owens, R.J. Expression of recombinant glycoproteins in mammalian cells: towards an integrative approach to structural biology. Curr. Opin. Struct. Biol. 23, 345–356 (2013).
Zacharias, D.A., Violin, J.D., Newton, A.C. & Tsien, R.Y. Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells. Science 296, 913–916 (2002).
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 20, 87–90 (2002).
Pardon, E. et al. A general protocol for the generation of Nanobodies for structural biology. Nat. Protoc. 9, 674–693 (2014).
Chang, V.T. et al. Glycoprotein structural genomics: solving the glycosylation problem. Structure 15, 267–273 (2007).
Walter, T.S. et al. A procedure for setting up high-throughput nanolitre crystallization experiments. Crystallization workflow for initial screening, automated storage, imaging and optimization. Acta Crystallogr. D Biol. Crystallogr. 61, 651–657 (2005).
Parker, J.L. & Newstead, S. Current trends in α-helical membrane protein crystallization: an update. Protein Sci. 21, 1358–1365 (2012).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Evans, P.R. & Murshudov, G.N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
Van den Abbeele, A. et al. A llama-derived gelsolin single-domain antibody blocks gelsolin-G-actin interaction. Cell. Mol. Life Sci. 67, 1519–1535 (2010).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Murshudov, G.N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Afonine, P.V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Strong, M. et al. Toward the structural genomics of complexes: crystal structure of a PE/PPE protein complex from Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 103, 8060–8065 (2006).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Stuart, D.I., Levine, M., Muirhead, H. & Stammers, D.K. Crystal structure of cat muscle pyruvate kinase at a resolution of 2.6 A. J. Mol. Biol. 134, 109–142 (1979).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Smart, O.S., Goodfellow, J.M. & Wallace, B.A. The pore dimensions of gramicidin A. Biophys. J. 65, 2455–2460 (1993).
Pettersen, E.F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Hosie, A.M., Dunne, E.L., Harvey, R.J. & Smart, T.G. Zinc-mediated inhibition of GABA(A) receptors: discrete binding sites underlie subtype specificity. Nat. Neurosci. 6, 362–369 (2003).
Banks, J.L. et al. Integrated modeling program, applied chemical theory (IMPACT). J. Comput. Chem. 26, 1752–1780 (2005).
Friesner, R.A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Halgren, T.A. et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J. Med. Chem. 47, 1750–1759 (2004).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Hattori, M., Hibbs, R.E. & Gouaux, E. A fluorescence-detection size-exclusion chromatography-based thermostability assay for membrane protein precrystallization screening. Structure 20, 1293–1299 (2012).
Acknowledgements
We thank staff at Diamond Light Source beamlines I03 and I04 for assistance at the synchrotron, K. Harlos and T. Walter for technical support with crystallization, J. Kammonen and staff at Pfizer for very kind time and assistance with electrophysiology, E. Beke for technical assistance during nanobody discovery, Y. Zhao for technical support with tissue culture, members of STRUBI for helpful discussions and G. Sutton and T. Malinauskas for feedback regarding the manuscript. This work was supported by the UK Biotechnology and Biological Sciences Research Council grant BB/M024709/1 (A.R.A. and P.S.M.), the UK Medical Research Council grants MR/L009609/1 and MC_UP_1201/15 (A.R.A.), the Human Frontier Science Program grant RGP0065/2014 (A.R.A.) and the Wellcome Trust studentships 105247/Z/14/Z (S.S.) and 084655/Z/08/Z (S.M.). We also thank INSTRUCT, part of the European Strategy Forum on Research Infrastructures and the Research Foundation-Flanders (FWO) for their nanobody discovery support. Further support from the Wellcome Trust Core Award 090532/Z/09/Z is acknowledged.
Author information
Authors and Affiliations
Contributions
Experimental work was performed by P.S.M. and S.S. (protein expression, purification, crystallization, electrophysiology), S.M. (protein expression, purification), L.D.C. (docking experiments), E.P. and J.S. (nanobody generation) and A.R.A. (crystallography). The manuscript was written by P.S.M. and A.R.A. with input from all coauthors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Detergent solubilised α5TMD binds pregnanolone at the Q245 neurosteroid potentiation site.
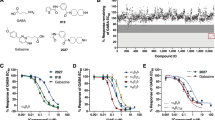
(a) Gel filtration profiles of purified α5TMD samples at 100 nM, heated beforehand for 1 hr at a range of temperatures to determine the decay of the pentameric peak protein species (a variable peak of protein aggregate is observed at 0.8 ml). (b) Thermostability plots of the pentameric peak decay with increasing temperature for α5TMD and its Q245L and Q245W mutants. Data points are from single melting experiments. (c) and (d) Gel filtration profiles for α5TMD and its Q245W mutant, respectively, showing the pentamer peak after heating to a temperature sufficient to cause 70 % peak decay (73 °C for α5TMD; 77 °C for Q245W), and subsequent dose-dependent stabilisation of the protein by increasing concentrations of pregnanolone. Remarkably, increasing pregnanolone concentrations drive α5TMD stability very close to its no heating (4 °C) level. (e) Pregnanolone dose-response curves for thermostabilisation of 70 % decayed α5TMD and its Q245L and Q245W mutants. NOTE: The solubility limit of pregnanolone in this buffer is 30 μM. Each data point represents mean ± s.e.m. of n = 3 experiments, each data point being from separate aliquots of purified protein.
Supplementary Figure 2 Architecture of the α5TMD-Nb25 complex.
(a) Side-on view of pregnanolone-bound (blue stick representation, in dashed box) α5TMD structure. Nb25 molecules are shown in green spacefill representation. Principal (P) and complementary (C) sides of one inter-subunit interface are labelled. (b) Close-up of Nb25 bound across the P/C interface near the neurotransmitter site. Dashed lines denote subunit/nanobody boundaries. Nb25 is shown in green, with a yellow CDR3 loop. The α5TMD P face is coloured in blue, with a cyan loop-C. (c) Close-up of interactions between the α5TMD neurotransmitter-binding loop-C (cyan) and the antigen recognising CDR3 loop of Nb25 (yellow). The closed conformation of the loop-C mimics that of the agonist-bound GABAAR-β3 (beige overlay, PDB ID: 4COF) rather than the open loop-C conformation of the antagonist-bound glycine receptor (grey overlay, PDB ID: 5CFB). (d) Binding footprint of Nb25 peeled 180° away from α5TMD. The hypervariable Nb25 regions, CDR1 (dashed circle), CDR2 (double-bordered ellipse) and CDR3 (single bordered ellipse), map broadly over the α5TMD β6-7 loop, loop-F and loop-C, respectively. Salt-bridge contacts are shown in red (Asp/Glu) and purple (Arg/Lys), polar contacts in cyan and van der Waals contacts in orange. Each Nb25 molecule forms ~1080 Å2 interfaces with α5TMD, roughly equally distributed between P/C faces. (e) and (f) Structural alignment of α5TMD pregnanolone-bound (blue) with apo (grey) or GABAAR-β3 (beige) ECDs, respectively. RMSD values over 204 equivalent ECD Cα positions are 0.19 Å (pregnanolone-bound vs apo α5TMD) and 0.45 Å (pregnanolone-bound α5TMD vs GABAAR-β3). (g) Bar chart showing that the amount of potentiation of EC10 histamine (his) by 10 μM pregnanolone (preg) is the same in the presence or absence of 50 μM Nb25. Each data point represents mean ± s.e.m. of n = 5 experiments, each measurement being from different cells.
Supplementary Figure 3 Crystallographic quality control for pregnanolone binding to α5TMD.
(a) SigmaA-weighted 2Fo-Fc (blue, contoured at 2σ) and Fo-Fc (green/red contoured at +3σ/-3σ) electron density maps following refinement in the absence of pregnanolone. (b) The same electron density maps and contour levels following refinement in the presence of pregnanolone. (c) Final model, showing pregnanolone (cyan) bound between the principal (P) and complementary faces (C) of adjacent subunits (PDB chains C and B respectively). Thick dotted lines indicate contacts within hydrogen-bonding distance. Thin dotted lines mark the inter-subunit boundary. (d) Overlay of the five pregnanolone molecules in α5TMD (cyan as in (c), the other four, at equivalent inter-subunit sites, in green). Pregnanolone molecules were overlaid by superposing the complementary chains from each site. Putative hydrogen bond distances between the pregnanolone 3α-hydroxyl and the M1 Gln245 εO range from 2.7-2.8 Å. Putative hydrogen bond distances between the pregnanolone C20 ketone and the Thr309 hydroxyl range from 3.0-3.4 Å.
Supplementary Figure 4 Impact of residue mutations lining the pregnanolone binding site on potentiation, and location of α-subunit M4 residues Asn410 and Tyr413 outside the site.
(a) Dose response curves for pregnanolone potentiation of EC10 histamine responses for α5TMDT287K and with mutations to single residues lining the pregnanolone binding site. (b) and (c) Pregnanolone bound to the inter-subunit site between M3 residues on the principal face (indicated by P in box) and M1 residues on the complementary face (indicated by C in box). The complementary face is shown in (a) cartoon or (b) cartoon with overlaying space-fill to highlight the top border of the pocket (dashed line). Putative hydrogen bonds between pregnanolone and α5TMD residues are indicated by dotted lines. Two M4 residues, Asn410 and Tyr413 (green labels) that have previously been implicated in pregnanolone binding by mutagenesis studies are shown to reside outside the site. M1 Ile238 and M4 Phe406 (blue labels) close the top of the site and bisect between M1 Gln245 and M4 Asn410 and Tyr413. Each data point represents mean ± s.e.m. Pregnanolone EC50 n number: WT n = 5, Q245S n = 8, V246A n = 4, W249L n = 7, I305 n = 3, T309A n = 3. Each n EC50 measurement is from different cells.
Supplementary Figure 5 Pregnanolone binding mode and α5TMD potentiation by neurosteroids.
(a) Top panel: structural formula of pregnanolone. Lower panel: the neurosteroid binding site in the α5TMD structure with pregnanolone bound (blue sticks, shown within SigmaA-weighted 2Fo-Fc electron density map contoured at 1.3σ, grey mesh) overlaid with computationally docked pregnanolone (yellow sticks). (b) Dose response curves for pregnanolone (3α5β), allopregnanolone (3α5α) and epipregnanolone (3β5β) potentiation of EC10 histamine responses for α5TMDT287K. Each data point represents mean ± s.e.m. Steroid EC50 n number: WT n = 5, allopregnanolone n = 3, epipregnanolone n = 10. Each n EC50 measurement is from different cells.
Supplementary Figure 6 Comparison between pregnanolone-bound α5TMD and GABAAR-β3 channel pore profiles and hydrophobicity.
Cross-sections showing the pore-lining M2 helices and the solvent-accessible surface mapped and colored according to the Eisenberg hydrophobicity scale (Eisenberg, D., Schwarz, E., Komaromy, M. & Wall, R., J. Mol. Biol. 179, 125–142, 1984) (pale green is hydrophobic; bright green is polar). Panels a-c show α5TMD and panels d-f show GABAAR-β3 (PDB ID: 4COF). (a) and (d) Sagittal ‘slices’ along the longitudinal pore axis. The dashed lines mark the beginning of the views shown in c and f. (b-c) and (e-f) Top-down views of the pore. View points are from the dashed lines shown a and d for b and e, respectively, and from further down the pore, just above the 2’ ring, for c and f. The desensitisation gate of α5TMD has two constrictions, the hydrophobic 2’ Val260 ring (pale green) and the -2’ Pro256 ring. In GABAAR-β3, the desensitisation gate has a single constriction, formed by the -2’ Ala248 ring.
Supplementary Figure 7 Crystal packing of pregnanolone-bound and apo α5TMD.
(a) Pregnanolone-bound α5TMD (space group I23) and (b) apo α5TMD (space group C2). Ligands are not shown, for clarity. Coloring scheme: ECD – blue, TMD – red, Nb25 – green. Dashed boxed areas in the left-hand panels are enlarged as close-ups in the right-hand panels. The M1-M2 linker (yellow) is not involved in crystal contacts.
Supplementary Figure 8 Comparison between the α5TMD pregnanolone-binding site and GluCl TMD modulator site locations.
Top-down (top panel) and side-on (lower panel) views of two neighbouring subunit TMDs. (a) Pregnanolone-bound α5TMD. (b) Ivermectin-bound GluCl (PDB ID: 3RIF). Boxed “P” and “C” letters indicate principal and complementary faces, respectively. Whilst pregnanolone occupies a location in the lower half, and on the outer surface of the TMD, the ivermectin site is in the top half of the GluCl TMD and ivermectin inserts deeper in the intersubunit interfaces.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1803 kb)
Structure of the α5TMD pentamer and its pregnanolone binding sites.
Cartoon representation of the α5TMD pentamer, complexed with Nb25 (green) and pregnanolone (ball-and-stick and spacefilling models, carbon atoms in blue, oxygen atoms in red), followed by zoom-in on one neurosteroid potentiation site. Subunits each comprise the human GABAA β3 ECD (light/dark blue) and α5 TMD (M1–M4 -helices, red/pink). N-linked glycans are shown as spheres coloured by atom type (carbon atoms in orange, oxygen atoms in red). The overview structure and the inter-subunit location of pregnanolone molecules are highlighted by oscillations (0–16 s). The zoom-in on the neurosteroid potentiation site (17–27 s) shows pregnanolone bound to an inter-subunit site between M3 residues on the principal face of one subunit (red) and M1 residues on the complementary face of the next (pink). (MP4 22055 kb)
Structural impact of pregnanolone binding to α5TMD.
Overview of the structural transition between the TMDs of globally superposed apo (orange/light orange) and pregnanolonebound(red/pink) α5TMD. A close-up view of one neurosteroid site (0–8 s) reveals the reshaping taking place upon pregnanolone (balland-stick representation, carbon atoms in blue, oxygen atoms in red) binding. The upper portion of the site contracts between principal face Ile305 and complementary face Gln245, whereas the lower portion of the site expands between Ile305 and complementary face Trp249. This creates torque on the lower part of the M1 helix twisting it sideways and outwards. A top-down view of the TMDs (10–16 s) shows a sideways and outward flexing of the lower (intracellular) halves of the five subunit helical bundles through the transition from apo to pregnanolone-bound states. A top-down zoom-in view of the bottom of the pore (18–24 s) shows the flexing of the transmembrane helices, the swing of the M1–M2 linker, and the dilation of the lower ring, defined by Pro256, of the desensitisation gate (Val260 surrounds the upper ring). The M2 motions are further emphasised by a side-on view between two opposing M2 helices lining the pore (29–34 s), where residues forming the desensitisation gate are highlighted. (MP4 15779 kb)
Impact of pregnanolone binding on the intra-subunit ECD-TMD relative orientation.
Neurosteroid binding induces changes in the relative positions of ECDs vs TMDs. Cartoons are coloured in light/dark blue (ECD, both α5TMD structures) and orange/light orange (apo α5TMD TMD) or ruby/pink (pregnanolone-bound α5TMD TMD); one subunit is highlighted. Pregnanolone molecules are shown in ball-and-stick and space-filling representations, carbon atoms in blue, oxygen atoms in red. The movie oscillates between global superpositions of apo and pregnanolone-bound α5TMD pentamers viewed side-on (0–6s), followed by focus on a single subunit viewed from the outside of the pentamer (7–12 s) and subsequently rotated to look into the inter-subunit principal face (14–20 s). The change in shape of the helical bundle in response to preganolone binding is visible from both orientations and when viewed looking into the inter-subunit principal face (14–20 s) reveals the flexure along M2 and the outward swing of the lower half of the TMD helical bundle. These motions induce an overall straightening of individual subunits due to a rocking motion of the ECD and TMD about their shared interface. (MP4 5143 kb)
Source data
Rights and permissions
About this article
Cite this article
Miller, P., Scott, S., Masiulis, S. et al. Structural basis for GABAA receptor potentiation by neurosteroids. Nat Struct Mol Biol 24, 986–992 (2017). https://doi.org/10.1038/nsmb.3484
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3484
This article is cited by
-
Neurosteroids and their potential as a safer class of general anesthetics
Journal of Anesthesia (2024)
-
Cryo-EM structures reveal native GABAA receptor assemblies and pharmacology
Nature (2023)
-
Conformational transitions and allosteric modulation in a heteromeric glycine receptor
Nature Communications (2023)
-
Structural insights into opposing actions of neurosteroids on GABAA receptors
Nature Communications (2023)
-
The molecular basis of drug selectivity for α5 subunit-containing GABAA receptors
Nature Structural & Molecular Biology (2023)