Abstract
Histone variants are structural components of eukaryotic chromatin that can replace replication-coupled histones in the nucleosome. The histone variant macroH2A1.1 contains a macrodomain capable of binding NAD+-derived metabolites. Here we report that macroH2A1.1 is rapidly induced during myogenic differentiation through a switch in alternative splicing, and that myotubes that lack macroH2A1.1 have a defect in mitochondrial respiratory capacity. We found that the metabolite-binding macrodomain was essential for sustained optimal mitochondrial function but dispensable for gene regulation. Through direct binding, macroH2A1.1 inhibits basal poly-ADP ribose polymerase 1 (PARP-1) activity and thus reduces nuclear NAD+ consumption. The resultant accumulation of the NAD+ precursor NMN allows for maintenance of mitochondrial NAD+ pools that are critical for respiration. Our data indicate that macroH2A1.1-containing chromatin regulates mitochondrial respiration by limiting nuclear NAD+ consumption and establishing a buffer of NAD+ precursors in differentiated cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Belenky, P., Bogan, K.L. & Brenner, C. NAD+ metabolism in health and disease. Trends Biochem. Sci. 32, 12–19 (2007).
Stein, L.R. & Imai, S. The dynamic regulation of NAD metabolism in mitochondria. Trends Endocrinol. Metab. 23, 420–428 (2012).
Altmeyer, M. & Hottiger, M.O. Poly(ADP-ribose) polymerase 1 at the crossroad of metabolic stress and inflammation in aging. Aging (Albany NY) 1, 458–469 (2009).
Bai, P. Biology of poly(ADP-ribose) polymerases: the factotums of cell maintenance. Mol. Cell 58, 947–958 (2015).
Gibson, B.A. & Kraus, W.L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat. Rev. Mol. Cell Biol. 13, 411–424 (2012).
Cantó, C., Menzies, K.J. & Auwerx, J. NAD+ metabolism and the control of energy homeostasis: a balancing act between mitochondria and the nucleus. Cell Metab. 22, 31–53 (2015).
Douet, J. et al. MacroH2A histone variants maintain nuclear organization and heterochromatin architecture. J. Cell Sci. 130, 1570–1582 (2017).
Buschbeck, M. & Di Croce, L. Approaching the molecular and physiological function of macroH2A variants. Epigenetics 5, 118–123 (2010).
Posavec, M., Timinszky, G. & Buschbeck, M. Macro domains as metabolite sensors on chromatin. Cell. Mol. Life Sci. 70, 1509–1524 (2013).
Rivera-Casas, C., Gonzalez-Romero, R., Cheema, M.S., Ausió, J. & Eirin-López, J.M. The characterization of macroH2A beyond vertebrates supports an ancestral origin and conserved role for histone variants in chromatin. Epigenetics 11, 415–425 (2016).
Chakravarthy, S. et al. Structural characterization of the histone variant macroH2A. Mol. Cell. Biol. 25, 7616–7624 (2005).
Kustatscher, G., Hothorn, M., Pugieux, C., Scheffzek, K. & Ladurner, A.G. Splicing regulates NAD metabolite binding to histone macroH2A. Nat. Struct. Mol. Biol. 12, 624–625 (2005).
Timinszky, G. et al. A macrodomain-containing histone rearranges chromatin upon sensing PARP1 activation. Nat. Struct. Mol. Biol. 16, 923–929 (2009).
Mehrotra, P.V. et al. DNA repair factor APLF is a histone chaperone. Mol. Cell 41, 46–55 (2011).
Ramamoorthy, M. & Smith, S. Loss of ATRX suppresses resolution of telomere cohesion to control recombination in ALT cancer cells. Cancer Cell 28, 357–369 (2015).
Jankevicius, G. et al. A family of macrodomain proteins reverses cellular mono-ADP-ribosylation. Nat. Struct. Mol. Biol. 20, 508–514 (2013).
Rosenthal, F. et al. Macrodomain-containing proteins are new mono-ADP-ribosylhydrolases. Nat. Struct. Mol. Biol. 20, 502–507 (2013).
Buschbeck, M. et al. The histone variant macroH2A is an epigenetic regulator of key developmental genes. Nat. Struct. Mol. Biol. 16, 1074–1079 (2009).
Kapoor, A. et al. The histone variant macroH2A suppresses melanoma progression through regulation of CDK8. Nature 468, 1105–1109 (2010).
Pasque, V., Gillich, A., Garrett, N. & Gurdon, J.B. Histone variant macroH2A confers resistance to nuclear reprogramming. EMBO J. 30, 2373–2387 (2011).
Changolkar, L.N. et al. Developmental changes in histone macroH2A1-mediated gene regulation. Mol. Cell. Biol. 27, 2758–2764 (2007).
Boulard, M. et al. Histone variant macroH2A1 deletion in mice causes female-specific steatosis. Epigenetics Chromatin 3, 8 (2010).
Sheedfar, F. et al. Genetic ablation of macrohistone H2A1 leads to increased leanness, glucose tolerance and energy expenditure in mice fed a high-fat diet. Int. J. Obes. (Lond.) 39, 331–338 (2015).
Pehrson, J.R., Changolkar, L.N., Costanzi, C. & Leu, N.A. Mice without macroH2A histone variants. Mol. Cell. Biol. 34, 4523–4533 (2014).
Sporn, J.C. et al. Histone macroH2A isoforms predict the risk of lung cancer recurrence. Oncogene 28, 3423–3428 (2009).
Dell'Orso, S. et al. The histone variant MacroH2A1.2 is necessary for the activation of muscle enhancers and recruitment of the transcription factor Pbx1. Cell Rep. 14, 1156–1168 (2016).
Houtkooper, R.H. et al. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 497, 451–457 (2013).
Creppe, C., Posavec, M., Douet, J. & Buschbeck, M. MacroH2A in stem cells: a story beyond gene repression. Epigenomics 4, 221–227 (2012).
Pirinen, E. et al. Pharmacological inhibition of poly(ADP-ribose) polymerases improves fitness and mitochondrial function in skeletal muscle. Cell Metab. 19, 1034–1041 (2014).
Bai, P. et al. PARP-1 inhibition increases mitochondrial metabolism through SIRT1 activation. Cell Metab. 13, 461–468 (2011).
Ouararhni, K. et al. The histone variant mH2A1.1 interferes with transcription by down-regulating PARP-1 enzymatic activity. Genes Dev. 20, 3324–3336 (2006).
Berger, F., Lau, C., Dahlmann, M. & Ziegler, M. Subcellular compartmentation and differential catalytic properties of the three human nicotinamide mononucleotide adenylyltransferase isoforms. J. Biol. Chem. 280, 36334–36341 (2005).
Chen, H. et al. MacroH2A1.1 and PARP-1 cooperate to regulate transcription by promoting CBP-mediated H2B acetylation. Nat. Struct. Mol. Biol. 21, 981–989 (2014).
Revollo, J.R., Grimm, A.A. & Imai, S. The NAD biosynthesis pathway mediated by nicotinamide phosphoribosyltransferase regulates Sir2 activity in mammalian cells. J. Biol. Chem. 279, 50754–50763 (2004).
Yang, H. et al. Nutrient-sensitive mitochondrial NAD+ levels dictate cell survival. Cell 130, 1095–1107 (2007).
Pittelli, M. et al. Inhibition of nicotinamide phosphoribosyltransferase: cellular bioenergetics reveals a mitochondrial insensitive NAD pool. J. Biol. Chem. 285, 34106–34114 (2010).
Nikiforov, A., Dölle, C., Niere, M. & Ziegler, M. Pathways and subcellular compartmentation of NAD biosynthesis in human cells: from entry of extracellular precursors to mitochondrial NAD generation. J. Biol. Chem. 286, 21767–21778 (2011).
Cambronne, X.A. et al. Biosensor reveals multiple sources for mitochondrial NAD+. Science 352, 1474–1477 (2016).
Creppe, C. et al. MacroH2A1 regulates the balance between self-renewal and differentiation commitment in embryonic and adult stem cells. Mol. Cell. Biol. 32, 1442–1452 (2012).
Sporn, J.C. & Jung, B. Differential regulation and predictive potential of MacroH2A1 isoforms in colon cancer. Am. J. Pathol. 180, 2516–2526 (2012).
Podrini, C. et al. MacroH2A1 isoforms are associated with epigenetic markers for activation of lipogenic genes in fat-induced steatosis. FASEB J. 29, 1676–1687 (2015).
Pazienza, V. et al. SIRT1-metabolite binding histone macroH2A1.1 protects hepatocytes against lipid accumulation. Aging (Albany NY) 6, 35–47 (2014).
Perdiguero, E. et al. Genetic analysis of p38 MAP kinases in myogenesis: fundamental role of p38alpha in abrogating myoblast proliferation. EMBO J. 26, 1245–1256 (2007).
Cantariño, N. et al. A cellular model reflecting the phenotypic heterogeneity of mutant HRAS driven squamous cell carcinoma. Int. J. Cancer 139, 1106–1116 (2016).
Dardenne, E. et al. Splicing switch of an epigenetic regulator by RNA helicases promotes tumor-cell invasiveness. Nat. Struct. Mol. Biol. 19, 1139–1146 (2012).
Buschbeck, M. et al. PML4 induces differentiation by Myc destabilization. Oncogene 26, 3415–3422 (2007).
Teperino, R. et al. Hedgehog partial agonism drives Warburg-like metabolism in muscle and brown fat. Cell 151, 414–426 (2012).
Wettenhall, J.M. & Smyth, G.K. limmaGUI: a graphical user interface for linear modeling of microarray data. Bioinformatics 20, 3705–3706 (2004).
Zhu, L.J. et al. ChIPpeakAnno: a Bioconductor package to annotate ChIP-seq and ChIP-chip data. BMC Bioinformatics 11, 237 (2010).
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS One 6, e21800 (2011).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zang, C. et al. A clustering approach for identification of enriched domains from histone modification ChIP-Seq data. Bioinformatics 25, 1952–1958 (2009).
Acknowledgements
We thank P. Muñoz Canoves for tools, training and advice; E. Gallardo (Institut de Recerca Hospital de la Santa Creu i Sant Pau, Barcelona, Spain) for primary human myoblasts; S. Samino for help with metabolomics analyses; S.-V. Forcales, A. Mikoč and the Ahel lab for helpful discussions; and the departments of pathology and biochemistry of the Hospital Universitari Germans Trias I Pujol (HGTP) for excellent support. This project was supported by MINECO (grants SAF2012-39749 and BFU2015-66559-P to M.B.; grant SAF2012-37427 to M.S.), AFM-Téléthon (grant 18738 to M.B.), the Marie Skłodowska Curie training network 'ChroMe' (grant H2020-MSCA-ITN-2015-675610 to M.B., A.G.L., O.Y. and J.A.P.), the Minerva Foundation (ARCHES award to T.P.), DZD (T.P.), the ERC (grants 281641 and 682679 to J.A.P.), DFG (grants SFB 646 and SFB 1064 to A.G.L.), the Wellcome Trust (grant 101794 to I.A.), Cancer Research UK (grant C35050/A22284 to I.A.), the Unity through Knowledge Fund (grant UKF 1B 2/13 to I.A.), ISCIII (grant PI15/00701 to P.M.G.-R.), MECD (FPU14/06542 to D.C.), AGAUR (FI fellowship to M.P.M.), and a Juan de la Cierva fellowship (JCI-2011-10831 to J.D.). Work in the Buschbeck lab is further supported by the Deutsche José Carreras Leukaemie Stiftung (DJCLS R 14/16), MINECO–ISCIII (PIE16/00011) and AGAUR (2014-SGR-35). Research at the IJC is supported by the 'La Caixa' Foundation, the Fundació Internacional Josep Carreras, Celgene Spain and the CERCA Programme/Generalitat de Catalunya.
Author information
Authors and Affiliations
Contributions
M.P.M., R.T. and M.B. conceived the project; M.P.M., S.H.-B., M.S., P.B., I.A., A.G.L., P.M.G.-R., O.Y., J.A.P., R.T. and M.B. designed experiments and interpreted data; O.Y. contributed methods; M.P.M., S.H.-B., M.L., V.V., H.D., M.N., D.C., I.G., J.D. and P.G.-P. performed experiments; R.M. analyzed high-content data; and M.P.M., J.A.P., R.T. and M.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Analysis of macroH2A1 splice variants during differentiation of primary myoblasts.
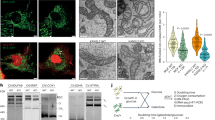
(a) Scheme of the mouse macroH2A1 transcript containing the region of the two alternative exons that give rise to macroH2A1.1 (exon 7) and macroH2A1.2 (exon 6). The macro domain of macroH2A1.1 variant was shown to bind ADPR, o-acetyl-ADPR and poly-ADPR in vitro (REF 12,13).
(b) Schematic representation of the RFLP analysis of mouse and human spliced transcripts encoding macroH2A1 isoforms. Exon numbers and the length of digested fragments (in bp) are indicated.
(c) mRNA levels of macroH2A variants were measured by RT-qPCR during mouse primary myoblasts differentiation and normalized to equimolar reference samples to allow direct comparison. Data is represented as mean + s.e.m. of three independent cell culture experiments. Mouse Myogenin (Myog) and muscle-type creatine kinase (Ckm) genes are used as markers of differentiation. Time point 0 refers to subconfluent cells grown in GM medium (same for d). Asterisks indicate significantly higher expression compared to the other isoform at the same time point (p<0.05).
(d) Same as (c) analyzing human primary myoblasts. Cells from two different donors were used. Data is the mean + s.e.m. of two independent cell cultures.
(e) Immunoblot analysis of mouse primary myoblasts using indicated antibodies.
(f) Comparative immunoblot of differentiated C2C12 cells and FLAG-tagged reference samples. All three macroH2A proteins, histone H3 and FLAG were analyzed. A sample of C2C12 cells treated with macroH2A1.1-specific siRNA was also included. Uncropped blot images of (e) and (f) are shown in Supplementary Data Set 1.
Supplementary Figure 2 Nuclear localization and chromatin association of macroH2A1.1.
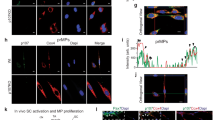
(a) Cross-sections of snap-frozen human muscle were fixed in cold 100% aceton and immunostained with anti-macroH2A1.1 antibody. Arrowheads indicate the positive signal of laterally located myonuclei in the muscle fibers. Scale bar is 50 μm.
(b) C2C12 cells were fixed at day 1.5 of differentiation and analyzed by immunofluorescence with anti-macroH2A1.1 antibody. DNA was stained with DAPI. Scale bar is 20 μm.
(c) Immunoblot analysis after cellular fractionation of differentiated C2C12 cells. Uncropped blot images are shown in Supplementary Data Set 1.
(d) Overview table of peaks called by SICER in proliferating myoblasts and after 5 days of differentiation.
(e) UCSC genome browser window of a representative 3 Mb window of chromosome 8 (8:121.500.000-124.500.000). Peaks called by SICER are indicated below the profiles.
(f) Overlap of SICER peaks at bp resolution.
Supplementary Figure 3 MacroH2A1.1 depletion has little influence on key metabolic genes.
(a) Overview of up- and downregulated probes comparing si macroH2A1.1 (mH2A1.1) to si ctrl myotubes after 4 days of differentiation. The numbers above and below brackets indicate the number of genes corresponding to these probes. For full detail please see Supplementary Data File 1.
(b) The expression of known regulatory genes of glucose uptake (Glut1, Glut4), glycolysis (Hk2, Pdk4, Pdh1Ea) and fatty acid uptake and metabolism were analyzed by semi-quantitative in control and si macroH2A1.1 knockdown cells at day 4 of differentiation. Data is represented as mean + s.d. of three independent experiments (*p<0.05).
Supplementary Figure 4 MacroH2A1.1-sensitive genes are not rescued by NMN or by PARP inhibition.
(a) The purity of isolated mitochondria was checked by western blot. (b) The mRNA levels of Nmnat3 in siRNA-treated C2C12 cells was analyzed by RT-qPCR. Data is the mean + s.d. of three independent experiments (*p<0.05).
(c) No rescue of gene expression. The mRNA levels of si macroH2A1.1-sensitive genes were analyzed at day 4 of differentiation under control and rescue conditions with PARP inhibitor (100 nM PARP inhibitor ABT-888 for 16 hours as in Figure 6g) and NMN (500 nM for 24 hours as in Figure 7e) by RT-qPCR. Data is represented as mean + s.d. of three independent experiments. Differences between si macroH2A1.1 alone and co-treatments were not significant.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Table 1. (PDF 814 kb)
Supplementary Data Set 1
Uncropped images and blots. (PDF 4648 kb)
Supplementary Data Set 2
Expression array data. (XLS 296 kb)
Rights and permissions
About this article
Cite this article
Posavec Marjanović, M., Hurtado-Bagès, S., Lassi, M. et al. MacroH2A1.1 regulates mitochondrial respiration by limiting nuclear NAD+ consumption. Nat Struct Mol Biol 24, 902–910 (2017). https://doi.org/10.1038/nsmb.3481
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3481
This article is cited by
-
Deficiency of histone variant macroH2A1.1 is associated with sexually dimorphic obesity in mice
Scientific Reports (2023)
-
MacroH2A restricts inflammatory gene expression in melanoma cancer-associated fibroblasts by coordinating chromatin looping
Nature Cell Biology (2023)
-
MacroH2A1.1 as a crossroad between epigenetics, inflammation and metabolism of mesenchymal stromal cells in myelodysplastic syndromes
Cell Death & Disease (2023)
-
Solid tumours hijack the histone variant network
Nature Reviews Cancer (2021)
-
MacroH2A1.1 has evolved to let PARP1 do more by loosening its grip on PAR
Nature Structural & Molecular Biology (2021)