Abstract
Hsp90 couples ATP hydrolysis to large conformational changes essential for activation of client proteins. The structural transitions involve dimerization of the N-terminal domains and formation of 'closed states' involving the N-terminal and middle domains. Here, we used Hsp90 mutants that modulate ATPase activity and biological function as probes to address the importance of conformational cycling for Hsp90 activity. We found no correlation between the speed of ATP turnover and the in vivo activity of Hsp90: some mutants with almost normal ATPase activity were lethal, and some mutants with lower or undetectable ATPase activity were viable. Our analysis showed that it is crucial for Hsp90 to attain and spend time in certain conformational states: a certain dwell time in open states is required for optimal processing of client proteins, whereas a prolonged population of closed states has negative effects. Thus, the timing of conformational transitions is crucial for Hsp90 function and not cycle speed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Zhao, R. et al. Navigating the chaperone network: an integrative map of physical and genetic interactions mediated by the hsp90 chaperone. Cell 120, 715–727 (2005).
Young, J.C., Moarefi, I. & Hartl, F.U. Hsp90: a specialized but essential protein-folding tool. J. Cell Biol. 154, 267–273 (2001).
Picard, D. Heat-shock protein 90, a chaperone for folding and regulation. Cell. Mol. Life Sci. 59, 1640–1648 (2002).
McClellan, A.J. et al. Diverse cellular functions of the Hsp90 molecular chaperone uncovered using systems approaches. Cell 131, 121–135 (2007).
Röhl, A., Rohrberg, J. & Buchner, J. The chaperone Hsp90: changing partners for demanding clients. Trends Biochem. Sci. 38, 253–262 (2013).
Mayer, M.P. & Le Breton, L. Hsp90: breaking the symmetry. Mol. Cell 58, 8–20 (2015).
Taipale, M., Jarosz, D.F. & Lindquist, S. HSP90 at the hub of protein homeostasis: emerging mechanistic insights. Nat. Rev. Mol. Cell Biol. 11, 515–528 (2010).
Prodromou, C. et al. Identification and structural characterization of the ATP/ADP-binding site in the Hsp90 molecular chaperone. Cell 90, 65–75 (1997).
Nemoto, T., Ohara-Nemoto, Y., Ota, M., Takagi, T. & Yokoyama, K. Mechanism of dimer formation of the 90-kDa heat-shock protein. Eur. J. Biochem. 233, 1–8 (1995).
Bose, S., Weikl, T., Bügl, H. & Buchner, J. Chaperone function of Hsp90-associated proteins. Science 274, 1715–1717 (1996).
Wegele, H., Muschler, P., Bunck, M., Reinstein, J. & Buchner, J. Dissection of the contribution of individual domains to the ATPase mechanism of Hsp90. J. Biol. Chem. 278, 39303–39310 (2003).
Prodromou, C. et al. The ATPase cycle of Hsp90 drives a molecular 'clamp' via transient dimerization of the N-terminal domains. EMBO J. 19, 4383–4392 (2000).
Smith, D.F. Tetratricopeptide repeat cochaperones in steroid receptor complexes. Cell Stress Chaperones 9, 109–121 (2004).
Panaretou, B. et al. ATP binding and hydrolysis are essential to the function of the Hsp90 molecular chaperone in vivo. EMBO J. 17, 4829–4836 (1998).
Obermann, W.M., Sondermann, H., Russo, A.A., Pavletich, N.P. & Hartl, F.U. In vivo function of Hsp90 is dependent on ATP binding and ATP hydrolysis. J. Cell Biol. 143, 901–910 (1998).
Mishra, P. & Bolon, D.N. Designed Hsp90 heterodimers reveal an asymmetric ATPase-driven mechanism in vivo. Mol. Cell 53, 344–350 (2014).
Shiau, A.K., Harris, S.F., Southworth, D.R. & Agard, D.A. Structural analysis of E. coli hsp90 reveals dramatic nucleotide-dependent conformational rearrangements. Cell 127, 329–340 (2006).
Ali, M.M. et al. Crystal structure of an Hsp90-nucleotide-p23/Sba1 closed chaperone complex. Nature 440, 1013–1017 (2006).
Hessling, M., Richter, K. & Buchner, J. Dissection of the ATP-induced conformational cycle of the molecular chaperone Hsp90. Nat. Struct. Mol. Biol. 16, 287–293 (2009).
Weikl, T. et al. C-terminal regions of Hsp90 are important for trapping the nucleotide during the ATPase cycle. J. Mol. Biol. 303, 583–592 (2000).
Mickler, M., Hessling, M., Ratzke, C., Buchner, J. & Hugel, T. The large conformational changes of Hsp90 are only weakly coupled to ATP hydrolysis. Nat. Struct. Mol. Biol. 16, 281–286 (2009).
Krukenberg, K.A., Förster, F., Rice, L.M., Sali, A. & Agard, D.A. Multiple conformations of E. coli Hsp90 in solution: insights into the conformational dynamics of Hsp90. Structure 16, 755–765 (2008).
Richter, K., Reinstein, J. & Buchner, J. N-terminal residues regulate the catalytic efficiency of the Hsp90 ATPase cycle. J. Biol. Chem. 277, 44905–44910 (2002).
Nathan, D.F. & Lindquist, S. Mutational analysis of Hsp90 function: interactions with a steroid receptor and a protein kinase. Mol. Cell. Biol. 15, 3917–3925 (1995).
Vaughan, C.K., Piper, P.W., Pearl, L.H. & Prodromou, C. A common conformationally coupled ATPase mechanism for yeast and human cytoplasmic HSP90s. FEBS J. 276, 199–209 (2009).
Hubert, D.A., He, Y., McNulty, B.C., Tornero, P. & Dangl, J.L. Specific Arabidopsis HSP90.2 alleles recapitulate RAR1 cochaperone function in plant NB-LRR disease resistance protein regulation. Proc. Natl. Acad. Sci. USA 106, 9556–9563 (2009).
Meyer, P. et al. Structural and functional analysis of the middle segment of hsp90: implications for ATP hydrolysis and client protein and cochaperone interactions. Mol. Cell 11, 647–658 (2003).
Retzlaff, M. et al. Asymmetric activation of the hsp90 dimer by its cochaperone aha1. Mol. Cell 37, 344–354 (2010).
Koulov, A.V. et al. Biological and structural basis for Aha1 regulation of Hsp90 ATPase activity in maintaining proteostasis in the human disease cystic fibrosis. Mol. Biol. Cell 21, 871–884 (2010).
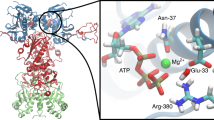
Cunningham, C.N., Southworth, D.R., Krukenberg, K.A. & Agard, D.A. The conserved arginine 380 of Hsp90 is not a catalytic residue, but stabilizes the closed conformation required for ATP hydrolysis. Protein Sci. 21, 1162–1171 (2012).
Lorenz, O.R. et al. Modulation of the Hsp90 chaperone cycle by a stringent client protein. Mol. Cell 53, 941–953 (2014).
Li, J., Richter, K., Reinstein, J. & Buchner, J. Integration of the accelerator Aha1 in the Hsp90 co-chaperone cycle. Nat. Struct. Mol. Biol. 20, 326–331 (2013).
Li, J., Richter, K. & Buchner, J. Mixed Hsp90–cochaperone complexes are important for the progression of the reaction cycle. Nat. Struct. Mol. Biol. 18, 61–66 (2011).
Richter, K., Walter, S. & Buchner, J. The Co-chaperone Sba1 connects the ATPase reaction of Hsp90 to the progression of the chaperone cycle. J. Mol. Biol. 342, 1403–1413 (2004).
Toogun, O.A., Dezwaan, D.C. & Freeman, B.C. The hsp90 molecular chaperone modulates multiple telomerase activities. Mol. Cell. Biol. 28, 457–467 (2008).
Echtenkamp, F.J. et al. Global functional map of the p23 molecular chaperone reveals an extensive cellular network. Mol. Cell 43, 229–241 (2011).
Mimnaugh, E.G., Worland, P.J., Whitesell, L. & Neckers, L.M. Possible role for serine/threonine phosphorylation in the regulation of the heteroprotein complex between the hsp90 stress protein and the pp60v-src tyrosine kinase. J. Biol. Chem. 270, 28654–28659 (1995).
Xu, Y. & Lindquist, S. Heat-shock protein hsp90 governs the activity of pp60v-src kinase. Proc. Natl. Acad. Sci. USA 90, 7074–7078 (1993).
Johnson, J.L., Halas, A. & Flom, G. Nucleotide-dependent interaction of Saccharomyces cerevisiae Hsp90 with the cochaperone proteins Sti1, Cpr6, and Sba1. Mol. Cell. Biol. 27, 768–776 (2007).
Hawle, P. et al. The middle domain of Hsp90 acts as a discriminator between different types of client proteins. Mol. Cell. Biol. 26, 8385–8395 (2006).
Richter, K. et al. Intrinsic inhibition of the Hsp90 ATPase activity. J. Biol. Chem. 281, 11301–11311 (2006).
Zurawska, A. et al. Mutations that increase both Hsp90 ATPase activity in vitro and Hsp90 drug resistance in vivo. Biochim. Biophys. Acta 1803, 575–583 (2010).
Tsutsumi, S. et al. Charged linker sequence modulates eukaryotic heat shock protein 90 (Hsp90) chaperone activity. Proc. Natl. Acad. Sci. USA 109, 2937–2942 (2012).
Soroka, J. et al. Conformational switching of the molecular chaperone Hsp90 via regulated phosphorylation. Mol. Cell 45, 517–528 (2012).
Mollapour, M. & Neckers, L. Post-translational modifications of Hsp90 and their contributions to chaperone regulation. Biochim. Biophys. Acta- 1823, 648–655 (2012).
Buchner, J., Weikl, T., Bügl, H., Pirkl, F. & Bose, S. Purification of Hsp90 partner proteins Hop/p60, p23, and FKBP52. Methods Enzymol. 290, 418–429 (1998).
Richter, K., Muschler, P., Hainzl, O. & Buchner, J. Coordinated ATP hydrolysis by the Hsp90 dimer. J. Biol. Chem. 276, 33689–33696 (2001).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Vranken, W.F. et al. The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59, 687–696 (2005).
Svergun, D.I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992).
Stafford, W.F. III. Boundary analysis in sedimentation transport experiments: a procedure for obtaining sedimentation coefficient distributions using the time derivative of the concentration profile. Anal. Biochem. 203, 295–301 (1992).
Hayes, D.B. & Stafford, W.F. SEDVIEW, real-time sedimentation analysis. Macromol. Biosci. 10, 731–735 (2010).
Acknowledgements
We acknowledge C. Göbl and C. Hartlmüller for help with the SAXS measurements, B. Tremmel for help with protein expression and purification, J. Soroka and J. Reinstein for inspiring discussions and comments on the manuscript and S. Lindquist (Whitehead Institute) for providing reagents. F.T. acknowledges a scholarship from the Studienstiftung des deutschen Volkes. This work was supported by the Bavarian Ministry of Sciences, Research and the Arts (Bavarian Molecular Biosystems Research Network, to T.M.), the Austrian Academy of Sciences (APART-fellowship to T.M.), the Austrian Science Fund (grant no. FWF: P28854 to T.M.), the Deutsche Forschungsgemeinschaft (grant no. SFB1035 to J.B. and M.S.) and the Emmy Noether program (grant no. MA 5703/1-1 to T.M.).
Author information
Authors and Affiliations
Contributions
B.K.Z. and F.T. performed experiments and data analysis. M.R. recorded and analyzed NMR data. T.M. performed and analyzed SAXS measurements. F.H.S. generated Hsp90-knockout strains and performed yeast tetrad analysis. K.R. performed the AUC experiments and analyzed data, J.B. designed experiments. B.K.Z., K.R., M.S. and J.B. wrote the manuscript. D.A.R. provided ATTO488-labeled GR and performed AUC runs.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 In vivo control experiments.
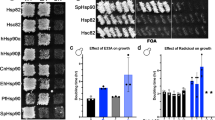
(a) Test for plasmid loss. Yeast cells were streaked out on media lacking uracil after 5’FOA shuffling. As control non shuffled cells expressing Hsp90 wt (pKAT6) were used. (b) Tetrad analysis was performed with p415-GPD-HSP82E33A or the empty vector (ev) as control. The resulted Hsp90 double knock-out strain is complemented by Hsp90 variant E33A (left panel) whereas the empty vector p415GPD was not able to support growth of the double knock-out as always one spore is inviable (right panel). The distributions of the kanMX cassette show tetra type (TT) distribution in the E33A complemented strain, always one out of four spores must carry a genomic double knock-out of Hsp90. In the vector control only two spores are G418 resistant confirming that the deletion of both Hsp90 alleles is lethal when Hsp90 is not provided by a plasmid. (c) 5’FOA Shuffling approach was carried out with different yeast shuffling strains and under GPD- and endogenous promotor. In all strains Hsp90 variant E33A supports yeast cell growth.
Supplementary Figure 2 SAXS scattering curves.
Experimental X-ray scattering data of Hsp90 versions recorded at different sample concentrations. Both the s, and I(s) axes are shown in a logarithmic representation. The angular ranges from 0.0012 - 0.4 nm-1 are compared.
Supplementary Figure 3 1H,15N-HSQC spectra of the Hsp90 N-domain mutants.
1H,15N-HSQC spectra of the indicated mutant (red) are superposed with spectra of the wildtype (black) of the N-terminal domain of yeast Hsp90. Negative peaks are plotted in orange and grey respectively. Examples of strongly shifting peaks are highlighted in boxes.
Supplementary Figure 4 Chemical-shift-perturbation plots of the Hsp90 N-domain mutants.
Chemical shift perturbation of the 1H,15N-HSQC spectra from Supplementary Figure 3 between the indicated mutant and the wildtype of the N-terminal domain of yeast Hsp90. Negative chemical shifts indicate shifting residues that could not be assigned in the complex. Residues which are not assigned in the wildtype are indicated by gaps. Chemical shift changes larger than 0.15 ppm are indicated on top of the bars. The red lines indicate the cut-off values used in Figure 2.
Supplementary Figure 5 1H,15N-HSQC spectra of the Hsp90 N-domain mutants in complex with nucleotide.
1H,15N-HSQC spectra of the indicated variant of the N-terminal domain of yeast Hsp90 with (red) and without (black) the indicated nucleotide are superposed. Negative peaks are plotted in orange and grey respectively. Examples of strongly shifting peaks are highlighted in boxes.
Supplementary Figure 6 Fluorescence spectra of the N- and M-domain labeled Hsp90 heterocomplex in the absence or presence of nucleotides.
The inset indicates the subunit exchange of the Hsp90 variants. Decrease in donor channel and increase in acceptor channel fluorescence were monitored and indicate FRET (inset). Following fluorescence spectra were recorded: donor only (grey), Hsp90 hetero-complex without nucleotide (black), in the presence of ATP (red) and ATPγS (blue).
Supplementary Figure 7 Hsp90-Aha1 interaction in vitro.
(a) The binding of the labeled co-chaperone Aha1* to different Hsp90 mutants in the presence of ATP monitored by analytical ultracentrifugation with fluorescence detection and derived from dc/dt plots (left panel). Following color code is used: Aha1* in brown, Aha1* in complex with: wt in black, A107N in yellow, Δ8 in blue, T22I in green, R346S in purple, R380A in grey, E33A in red and D79N in light blue. The areas of the Hsp90-Aha1* complex peaks in presence of 2 mM ATP (right panel). Error bars indicate standard error of the fit. (b) N-terminal dimerization stability of the Hsp90 variants in presence of ATP and Aha1. FRET chase experiments were performed with preformed Hsp90 FRET-complexes in the presence of ATP and the co-chaperone Aha1. The chase was induced by addition of 20-fold excess of the unlabeled Hsp90 D79N and the disruption of complex was monitored by following the decrease in acceptor fluorescence. Apparent half-lives (t1/2) were derived from a non-linear fit of the acceptor signal changes. Means of technical replicates are shown. Error bars indicate s.d. (n value=3).
Supplementary Figure 8 Nucleotide-dependent Hsp90-p23 and Hsp90-GR interactions in vitro.
Hsp90 variants were titrated with increasing amounts of fluorescein-labeled p23*. The binding of p23 to different Hsp90 mutants in the presence of (a) 2 mM ATPγS, (b) 2 mM ATP and (c) without nucleotide were monitored by analytical ultracentrifugation with a fluorescence detection unit and derived from dc/dt plots. (d) The binding of labeled GR* to different Hsp90 mutants in the absence of nucleotide were monitored by analytical ultracentrifugation and derived from dc/dt plots. Following color code is used: GR* in brown, GR* in complex with: wt in black, A107N in yellow, Δ8 in blue, T22I in green, R346S in purple, R380A in grey, E33A in red and D79N in light blue.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–4 (PDF 1828 kb)
Supplementary Data Set 1
Uncropped Western Blot data (PDF 149 kb)
Rights and permissions
About this article
Cite this article
Zierer, B., Rübbelke, M., Tippel, F. et al. Importance of cycle timing for the function of the molecular chaperone Hsp90. Nat Struct Mol Biol 23, 1020–1028 (2016). https://doi.org/10.1038/nsmb.3305
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3305
This article is cited by
-
Nucleotide exchange is sufficient for Hsp90 functions in vivo
Nature Communications (2023)
-
Hsp90 modulates human sperm capacitation via the Erk1/2 and p38 MAPK signaling pathways
Reproductive Biology and Endocrinology (2021)
-
A methylated lysine is a switch point for conformational communication in the chaperone Hsp90
Nature Communications (2020)
-
Conformational dynamics modulate the catalytic activity of the molecular chaperone Hsp90
Nature Communications (2020)
-
Glucocorticoid receptor complexes form cooperatively with the Hsp90 co-chaperones Pp5 and FKBPs
Scientific Reports (2020)