Abstract
Repair of DNA double-strand breaks (DSBs) in mammals is coordinated by the ubiquitin-dependent accumulation of 53BP1 at DSB-flanking chromatin. Owing to its ability to limit DNA-end processing, 53BP1 is thought to promote nonhomologous end-joining (NHEJ) and to suppress homology-directed repair (HDR). Here, we show that silencing 53BP1 or exhausting its capacity to bind damaged chromatin changes limited DSB resection to hyper-resection and results in a switch from error-free gene conversion by RAD51 to mutagenic single-strand annealing by RAD52. Thus, rather than suppressing HDR, 53BP1 fosters its fidelity. These findings illuminate causes and consequences of synthetic viability acquired through 53BP1 silencing in cells lacking the BRCA1 tumor suppressor. We show that such cells survive DSB assaults at the cost of increasing reliance on RAD52-mediated HDR, which may fuel genome instability. However, our findings suggest that when challenged by DSBs, BRCA1- and 53BP1-deficient cells may become hypersensitive to, and be eliminated by, RAD52 inhibition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Jackson, S.P. & Bartek, J. The DNA-damage response in human biology and disease. Nature 461, 1071–1078 (2009).
Ceccaldi, R., Rondinelli, B. & D'Andrea, A.D. Repair pathway choices and consequences at the double-strand break. Trends Cell Biol. 26, 52–64 (2016).
Chapman, J.R., Taylor, M.R. & Boulton, S.J. Playing the end game: DNA double-strand break repair pathway choice. Mol. Cell 47, 497–510 (2012).
Mehta, A. & Haber, J.E. Sources of DNA double-strand breaks and models of recombinational DNA repair. Cold Spring Harb. Perspect. Biol. 6, a016428 (2014).
Bunting, S.F. & Nussenzweig, A. End-joining, translocations and cancer. Nat. Rev. Cancer 13, 443–454 (2013).
Symington, L.S. End resection at double-strand breaks: mechanism and regulation. Cold Spring Harb. Perspect. Biol. 6, a016436 (2014).
Prakash, R., Zhang, Y., Feng, W. & Jasin, M. Homologous recombination and human health: the roles of BRCA1, BRCA2, and associated proteins. Cold Spring Harb. Perspect. Biol. 7, a016600 (2015).
Zhao, W. et al. Promotion of BRCA2-dependent homologous recombination by DSS1 via RPA targeting and DNA mimicry. Mol. Cell 59, 176–187 (2015).
Lukas, J., Lukas, C. & Bartek, J. More than just a focus: the chromatin response to DNA damage and its role in genome integrity maintenance. Nat. Cell Biol. 13, 1161–1169 (2011).
Panier, S. & Durocher, D. Push back to respond better: regulatory inhibition of the DNA double-strand break response. Nat. Rev. Mol. Cell Biol. 14, 661–672 (2013).
Boersma, V. et al. MAD2L2 controls DNA repair at telomeres and DNA breaks by inhibiting 5′ end resection. Nature 521, 537–540 (2015).
Bunting, S.F. et al. 53BP1 inhibits homologous recombination in Brca1-deficient cells by blocking resection of DNA breaks. Cell 141, 243–254 (2010).
Callen, E. et al. 53BP1 mediates productive and mutagenic DNA repair through distinct phosphoprotein interactions. Cell 153, 1266–1280 (2013).
Chapman, J.R. et al. RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Mol. Cell 49, 858–871 (2013).
Escribano-Díaz, C. et al. A cell cycle-dependent regulatory circuit composed of 53BP1-RIF1 and BRCA1-CtIP controls DNA repair pathway choice. Mol. Cell 49, 872–883 (2013).
Feng, L., Fong, K.W., Wang, J., Wang, W. & Chen, J. RIF1 counteracts BRCA1-mediated end resection during DNA repair. J. Biol. Chem. 288, 11135–11143 (2013).
Wang, J. et al. PTIP associates with Artemis to dictate DNA repair pathway choice. Genes Dev. 28, 2693–2698 (2014).
Xu, G. et al. REV7 counteracts DNA double-strand break resection and affects PARP inhibition. Nature 521, 541–544 (2015).
Zimmermann, M., Lottersberger, F., Buonomo, S.B., Sfeir, A. & de Lange, T. 53BP1 regulates DSB repair using Rif1 to control 5′ end resection. Science 339, 700–704 (2013).
Zimmermann, M. & de Lange, T. 53BP1: pro choice in DNA repair. Trends Cell Biol. 24, 108–117 (2014).
Altmeyer, M. & Lukas, J. To spread or not to spread: chromatin modifications in response to DNA damage. Curr. Opin. Genet. Dev. 23, 156–165 (2013).
Gudjonsson, T. et al. TRIP12 and UBR5 suppress spreading of chromatin ubiquitylation at damaged chromosomes. Cell 150, 697–709 (2012).
Zong, D. et al. Ectopic expression of RNF168 and 53BP1 increases mutagenic but not physiological non-homologous end joining. Nucleic Acids Res. 43, 4950–4961 (2015).
Toledo, L.I. et al. ATR prohibits replication catastrophe by preventing global exhaustion of RPA. Cell 155, 1088–1103 (2013).
Sartori, A.A. et al. Human CtIP promotes DNA end resection. Nature 450, 509–514 (2007).
Li, X. et al. Rad51 and Rad54 ATPase activities are both required to modulate Rad51-dsDNA filament dynamics. Nucleic Acids Res. 35, 4124–4140 (2007).
Somyajit, K., Basavaraju, S., Scully, R. & Nagaraju, G. ATM- and ATR-mediated phosphorylation of XRCC3 regulates DNA double-strand break-induced checkpoint activation and repair. Mol. Cell. Biol. 33, 1830–1844 (2013).
Nakanishi, K. et al. Human Fanconi anemia monoubiquitination pathway promotes homologous DNA repair. Proc. Natl. Acad. Sci. USA 102, 1110–1115 (2005).
Pierce, A.J., Johnson, R.D., Thompson, L.H. & Jasin, M. XRCC3 promotes homology-directed repair of DNA damage in mammalian cells. Genes Dev. 13, 2633–2638 (1999).
Cao, L. et al. A selective requirement for 53BP1 in the biological response to genomic instability induced by Brca1 deficiency. Mol. Cell 35, 534–541 (2009).
Chapman, J.R., Sossick, A.J., Boulton, S.J. & Jackson, S.P. BRCA1-associated exclusion of 53BP1 from DNA damage sites underlies temporal control of DNA repair. J. Cell Sci. 125, 3529–3534 (2012).
Kakarougkas, A. et al. Opposing roles for 53BP1 during homologous recombination. Nucleic Acids Res. 41, 9719–9731 (2013).
Muñoz, M.C. et al. RING finger nuclear factor RNF168 is important for defects in homologous recombination caused by loss of the breast cancer susceptibility factor BRCA1. J. Biol. Chem. 287, 40618–40628 (2012).
Xie, A. et al. Distinct roles of chromatin-associated proteins MDC1 and 53BP1 in mammalian double-strand break repair. Mol. Cell 28, 1045–1057 (2007).
Cruz-García, A., López-Saavedra, A. & Huertas, P. BRCA1 accelerates CtIP-mediated DNA-end resection. Cell Rep. 9, 451–459 (2014).
Lok, B.H. & Powell, S.N. Molecular pathways: understanding the role of Rad52 in homologous recombination for therapeutic advancement. Clin. Cancer Res. 18, 6400–6406 (2012).
O'Connor, M.J. Targeting the DNA damage response in cancer. Mol. Cell 60, 547–560 (2015).
Davoli, T. et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 155, 948–962 (2013).
Stephens, P.J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Delahaye-Sourdeix, M. et al. The 12p13.33/RAD52 locus and genetic susceptibility to squamous cell cancers of upper aerodigestive tract. PLoS One 10, e0117639 (2015).
Li, Z. et al. Association of a functional RAD52 genetic variant locating in a miRNA binding site with risk of HBV-related hepatocellular carcinoma. Mol. Carcinog. 54, 853–858 (2015).
Halazonetis, T.D., Gorgoulis, V.G. & Bartek, J. An oncogene-induced DNA damage model for cancer development. Science 319, 1352–1355 (2008).
Acknowledgements
We are grateful to L. Toledo and K.J. Neelsen for useful suggestions and critical comments on the manuscript. Work in the laboratory of J.L. was supported by grants from the Novo Nordisk Foundation (NNF14CC0001) and the Danish Cancer Society (R72-A4436).
Author information
Authors and Affiliations
Contributions
F.O., C.L., and J.L. conceived the project. C.L. and J.L. supervised the study. F.O. carried out high-content microscopy, quantitative image analysis and survival assays. K.S. performed reporter assays, biochemical analyses and survival assays. C.L. performed confocal microscopy. M.A. helped to develop high-content imaging assays. M.-B.R. performed western blots, generated cell lines, and contributed to characterization of the anti-RAD52 antibody. J.L. wrote the manuscript. All authors contributed to conceptual development of the project and manuscript editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 QIBC defines an intrinsic threshold for 53BP1 accumulation at DSBs.
(a) Schematic depiction of the QIBC workflow to quantitatively interrogate nuclear repair reactions during the cell cycle. A typical experiment consists of three steps. Step 1, exponentially growing cells are exposed to IR and stained for DAPI, cyclin A and a specific genome caretaker. Step 2, images are acquired using automated wide field microscopy and the cells are plotted according to their cell cycle distribution based on DAPI and cyclin A signals (every dot represents a single cell). Step 3, fluorescent signals associated with the genome caretaker are quantified and expressed in a color range (heat map), whose intensity reflects the frequency and magnitude of repair events for every single cell. Typically 1000 cells are analyzed for each condition. Scale bar, 10 μm. Additional technical specification can be found in Online Methods and ref. 24. (b) QIBC of 53BP1 accumulation at DSB sites in U-2-OS cells exposed to increasing dose of IR. The sum of the intensities of 53BP1 foci per nucleus is shown (n=1000 cells for each IR dose). Median levels are depicted in green. A.U., arbitrary units. Green arrow marks the IR dose where 53BP1 accumulation at DSBs reaches its threshold. (c) QIBC of γ-H2AX accumulation at DSB sites in U-2-OS cells exposed to increasing dose of IR. Mean γ-H2AX intensity per nucleus is shown (n=1000 cells for each IR dose). Median levels in red. A.U., arbitrary units.
Supplementary Figure 2 Impaired accumulation of RAD51 at IRIF after 53BP1 exhaustion is evident after preextraction of soluble nuclear protein.
(a) QIBC of RAD51 accumulation at IRIF in U-2-OS cells pulse-labeled with EdU, exposed to the indicated doses of IR, and stained after pre-extraction for DAPI and RAD51 (n=1000 cells for each IR dose). Heat map indicates the sum of the intensities of RAD51 foci per nucleus. A.U., arbitrary units. (b) Schematic depiction of cell cycle gating over DAPI-EdU profile obtained by QIBC (left). Quantification of cell cycle distribution of RAD51 IRIF generated by 1 and 10 Gy respectively (right) under QIBC conditions shown in (a). Median values show the sum of the intensities of RAD51 foci per nucleus (n=300 cells for each IR dose and cell cycle stage). A.U., arbitrary units.
Supplementary Figure 3 Suppression of RAD51 after 53BP1 exhaustion is specific and proceeds with similar kinetics across different cell types.
(a) QIBC of RAD51 accumulation at IRIF in U-2-OS cells treated with control siRNA, exposed to the indicated doses of IR and stained for DAPI, cyclin A and RAD51 (n=1000 cells for each IR dose). The heat map indicates the sum of the intensities of RAD51 foci per nucleus. A.U., arbitrary units. (b) QIBC of RAD51 accumulation at IRIF in U-2-OS cells treated with BRCA1 siRNA and analyzed as in (a). A.U., arbitrary units. (c) QIBC of RAD51 accumulation at IRIF in U-2-OS cells treated with BRCA2 siRNA and analyzed as in (a). A.U., arbitrary units. (d) QIBC of RAD51 accumulation at IRIF in Tert-immortalized RPE cells exposed to the increasing dose range of IR and stained for DAPI, cyclin A and RAD51 (n=1000 cells for each IR dose). The heat map indicates the sum of the intensities of RAD51 foci per nucleus. A.U., arbitrary units.
Supplementary Figure 4 Slowdown of resection by partial knockdown of CtIP restores RAD51 focus formation after 53BP1 exhaustion.
(a) Western blots of total cell lysates of U-2-OS cells treated with increasing doses of CtIP siRNA for 24 h. (b) QIBC of RPA accumulation in repair foci in U-2-OS cells treated with the indicated siRNAs, exposed to the indicated doses of IR and stained after pre-extraction for DAPI and RPA1. Mean RPA intensity per nucleus is shown (n=1000 cells for each IR dose). Median levels are depicted in blue. A.U., arbitrary units. (c) QIBC of RAD51 accumulation at IRIF in U-2-OS cells treated with the indicated siRNAs, exposed to IR (10 Gy), and stained for DAPI, cyclin A and RAD51 (n=1000 cells for each condition). The heat map indicates the sum of the intensities of RAD51 foci per nucleus. The bar chart shows the average sum of RAD51 foci intensity late S and G2 cells depicted in insets (n=300 cells for each condition). A.U., arbitrary units.
Supplementary Figure 5 Characterization of the YFP-RAD52 cell line.
(a) Western blots of total cell lysates of naïve and (YFP-RAD52) variants of U-2-OS cells treated with RAD51 siRNA as indicated. (b) QIBC plot of YFP-RAD52 protein levels in U-2-OS (YFP-RAD52) cells stained with DAPI. Mean intensities of the YFP signal per nucleus are shown. The plot is divided into four intensity layers (left) with representative cells (right). The highlighted intensity layer marks the range of intensities that are similar to the level of endogenous RAD52 and thus delineates cells selected for QIBC analyses throughout this study. (c) QIBC of YFP-RAD52 accumulation at IRIF in U-2-OS (YFP-RAD52) cells treated with control siRNA (top) and CtIP siRNA (bottom), exposed to the indicated doses of IR and stained for DAPI and cyclin A (n=1000 cells for each IR dose). The heat map indicates the sum of the intensities of YFP-RAD52 foci per nucleus. A.U., arbitrary units.
Supplementary Figure 6 Increasing load of DSBs triggers DSB hyper-resection.
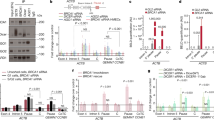
(a) Quantification of RPA QIBC shown in Figure 3d. Mean RPA1 intensity per nucleus is shown (n=1000 cells for each IR dose). Median levels are depicted in blue. A.U., arbitrary units. (b) Dot blots of genomic DNA from U-2-OS cells treated with the indicated doses of IR and stained for ssDNA. Primary data (left) and their quantification (right; single data points with mean, n=2 technical replicates) are shown.
Supplementary Figure 7 53BP1 promotes GC and suppresses SSA.
(a) Quantification of the GC reporter assay in U-2-OS cells treated with the indicated siRNAs and transfected with the I-SceI expression plasmid (average ± s.d.; n=3 independent knockdown experiments). (b) The SSA reporter assay in U-2-OS cells treated as in (a). Top panel shows PCR products amplified with the indicated primers (see Fig. 6b for schematic depiction). Bottom panel is quantification of the SSA-specific PCR products (average ± s.d.; n=3 independent knockdown experiments). A.U., arbitrary units. (c) Quantification of the GC reporter assay in U-2-OS cells treated with the indicated siRNAs and transfected with the I-SceI expression plasmid (average ± s.d.; n=3 independent knockdown experiments). (d) The SSA reporter assay in U-2-OS cells treated as in (c). Top panel shows PCR products amplified with the indicated primers as in (b). Bottom panel is quantification of the 0.8 kb SSA-specific PCR products (average ± s.d.; n=3 independent knockdown experiments). A.U., arbitrary units. (e) Clonogenic survival of U-2-OS cells treated with the indicated siRNAs and exposed to the indicated doses of IR (mean values, n=2 technical replicates). (f) Clonogenic survival of U-2-OS cells treated with the indicated siRNAs and exposed to the indicated doses of IR (mean values, n=2 technical replicates). (g) Clonogenic survival of U-2-OS cells treated with the indicated siRNAs and exposed to the indicated doses of IR (mean values, n=2 technical replicates).
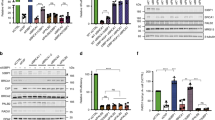
Supplementary Figure 8 Specificity of siRNA knockdowns.
Western blots of total cell lysates from U-2-OS cells treated with the indicated siRNAs.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1565 kb)
Supplementary Data Set 1
Uncropped western blots (PDF 8068 kb)
Rights and permissions
About this article
Cite this article
Ochs, F., Somyajit, K., Altmeyer, M. et al. 53BP1 fosters fidelity of homology-directed DNA repair. Nat Struct Mol Biol 23, 714–721 (2016). https://doi.org/10.1038/nsmb.3251
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3251
This article is cited by
-
Fanconi anemia associated protein 20 (FAAP20) plays an essential role in homology-directed repair of DNA double-strand breaks
Communications Biology (2023)
-
ABRAXAS1 orchestrates BRCA1 activities to counter genome destabilizing repair pathways—lessons from breast cancer patients
Cell Death & Disease (2023)
-
Genetic separation of Brca1 functions reveal mutation-dependent Polθ vulnerabilities
Nature Communications (2023)
-
Depletion of NK6 Homeobox 3 (NKX6.3) causes gastric carcinogenesis through copy number alterations by inducing impairment of DNA replication and repair regulation
Oncogenesis (2021)
-
Fanconi anemia pathway as a prospective target for cancer intervention
Cell & Bioscience (2020)